All published articles of this journal are available on ScienceDirect.
Study of Various Virulence Genes, Biofilm Formation and Extended-Spectrum β-lactamase Resistance in Klebsiella pneumoniae Isolated from Urinary Tract Infections
Abstract
Objective:
The aims of the current study were to evaluate the capacity of K. pneumoniae isolated from hospital-acquired urinary tract infection to form biofilm, the relation of this capacity to various virulence genes and the prevalence of Extended Spectrum β-lactamases (ESBL) among these isolates by phenotypic and genotypic methods.
Material and Methods:
The study included 100 non-duplicate strains of K. pneumoniae isolated from 100 different urine samples from patients with hospital-acquired urinary tract infection. The isolated strains were studied for biofilm formation, ESBL production by phenotypic methods. Molecular studies were applied for the detection of ESβLs genes blaTEM, blaSHV, blaCTX-M and for detection of virulence genes fimH, uge, rmpA, mag A, wzy, kfa and aerobactin genes.
Result:
The majority of the isolates had the capacity to form a biofilm (81%), with ESBL prevalence rate 41%. The most prevalent gene among ESBL producing K. pneumoniae was blaCTX-M (73.2%) followed by blaSHV (53.6%) and blaTEM (51.2%). Among the virulence genes studied in K. pneumoniae isolates, the most prevalent gene was fimH (76%), uge (70%). There was significant association between ESBL production, and resistance to amikacin, cefepime, ceftazidime, gentamicin, imipenem and meropenem and biofilm production in K. pneumoniae isolates. There was significant association between blaCTX-M, blaSHV, fimH, mag, kfa, wzy, rmpA and aerobactin and biofilm production in K. pneumoniae.
Conclusion:
The present study highlights the prevalence of virulence genes among biofilm-forming strains of K. pneumoniae isolated from hospital-acquired urinary tract infection. Moreover, there was association between biofilm formation and ESBL production. Further studies are required to elucidate the clinical impact of the association of these different mechanisms.
1. INTRODUCTION
Klebsiella pneumoniae (K. pneumoniae), as a gram-negative bacillus, is known to be associated with various infections such as Urinary Tract Infection (UTI) especially hospital-acquired Catheter-Associated Urinary Tract Infection (CAUTI), sepsis and pneumonia [1, 2].
There are many virulence factors in K. pneumoniae associated with its pathogenicity. Among these factors is the capacity of K. pneumoniae to form a biofilm that protects it from the host immune response as well as from the antibiotics [3, 4]. The formation of biofilm leads to the prolonged persistence of K. pneumoniae on the epithelium and on the medical devices with difficulty in its eradication. There are genes related to biofilm formation in K. pneumoniae such as capsular genes like rmpA gene associated with K1/K2 virulent type, fimbriae genes such as fimH and mrkD that regulate fimbrial types I and 3, respectively and genes encoding the enzymes used for the biosynthesis of lipopolysaccharide such as uge gene that encodes for the uridine diphosphate galacturonate 4-epimerase [5, 6] and genes that regulate iron uptake by K. peumoniae including aerobactin and kfu genes [7]. Moreover, quorum sensing system especially type 2 and pga ABCD operon that regulates the synthesis and translocation of poly-b-1,6-N-acetyl-D-glucosamine adhesin are involved in the biofilm formation through enhancement of cell to cell communication and intercellular adhesion process [8].
Beside biofilm formation, clinical isolates of K. pneumoniae are known to possess resistance to third generation of cephalosporines through the production of Extended Spectrum β-Lactamase (ESBL).
Extended-spectrum beta-lactamase is associated with the mutation of genes that regulate β-lactamases mainly blaSHV, blaTEM, and blaCTX-M genes. More than 300 different ESBLs variants have been described [9].
There are few reports about the association of biofilm and ESBL within isolates of K. pneumoniae from hospital-acquired urinary tract infection.
Therefore, the present study aimed to study the capacity of K. pneumoniae isolated from hospital-acquired urinary tract infection to form biofilm and to correlate this capacity to various virulence genes and finally detect the prevalence of ESBL among these isolates by phenotypic and genotypic methods.
2. MATERIALS AND METHODS
The study included 100 non-duplicate strains of K. pneumoniae isolated from 100 patients with hospital-acquired urinary tract infections from Mansoura University Hospital, Egypt as defined by CDC in a period from December 2016 till January 2018. The study was approved by Mansoura Faculty of Medicine ethical committee and approval was obtained from each participant.
2.1. Microbiological Culture
Urine samples from catheterized patients were collected as part of the routine microbiological investigation, after disinfection of the catheter collection part using 70% ethyl alcohol solution discarding the first few drops of urine and collecting 5-10 ml urine using a sterile syringe. Urine samples were immediately sent to the laboratory to be processed and examined within 2 hours. Quantitative urine culture was performed by the use of sterile calibrated loop with a culture of I micron of the urine onto MacConkey agar plate and incubation was performed at 37°C for 24-48 hours. The colonies were identified by gram staining and biochemical identification was performed by VITEK 2 system according to the instructions of the manufacturers. The isolated K. pneumoniae were subjected to antibiotics susceptibility testing by disc diffusion method, determination of ESBL done by combined disc diffusion method and study of biofilm formation by Microtiter plate method. Molecular studies were carried out for the detection of virulence genes and bla genes.
2.2. Antibiotic Susceptibility Testing
Antibiotic susceptibility was performed by discs diffusion method according to Clinical and Laboratory Standards Institute (CLSI) recommendations (CLSI 2018) [10]. The used discs were: amikacin, cefotaxime, ceftazidime, cefepime, ciprofloxacin, levofloxacin, gentamicin, piperacillin-tazobactam, trimethoprim/sulfamethoxazole, imipenem, and meropenem (Oxoid-UK). The interpretation of the results was done according to CLSI 2018 [10].
2.3. ESBL Production
Determination of ESBL by Combined disk diffusion confirmatory test.
Klebsiella pneumoniae strains resistant to ceftazidime (30 μg) and/ or cefotaxime (30 μg) were subjected to the screening of ESBL enzymes production by combined disk diffusion confirmatory test. The principle of the test depends upon the use of ceftazidime and cefotaxime discs alone and conjugated with clavulanic acid. The increase of the inhibition zone diameter around discs conjugated with clavulanic acid ≥5mm was interpreted as positive ESBL production.
The suspension of overnight growth of K. pneumoniae in Muller-Hinton broth was prepared with adjusted turbidity to 0.5 McFarland and was subculture on Muller-Hinton agar medium and antibiotic discs containing ceftazidime, ceftazidime (30 μg)+clavulanic acid (10 μg), cefotaxime, and cefotaxime (30 μg + clavulanic (10 μg) pairs were placed with 20 mm space between them. Plates were incubated at 37ºC for 24 hours. The increase of growth inhibition zone ≥ 5 mm around ceftazidime/clavulanic acid and cefotaxime/clavulanic compared to ceftazidime and cefotaxime respectively was regarded as an ESBL producing isolate [11]. The strain of Klebsiella pneumoniae (ATCC 700603) was used as a positive quality control strain for ESBL production.
2.4. Biofilm Assay
Biofilm formation by K. pneumoniae isolates was assessed by the semi-quantitative method [12].
Briefly, 0.5 McFarland suspensions of K. pneumoniae isolates were prepared using brain heart infusion broth (Oxoid-UK) and 100 microliters of the suspensions were inoculated onto wells of 96-well polystyrene plate and incubated at 37°C for 18 hours. Negative control well was inoculated with sterile brain heart infusion broth. Then, wells were stained with 200 microliters of crystal violet with concentration 0.5% for 20 minutes and the excess stain was washed by 200 microns sterile distilled water for three times. Then the stain was eluted with 200 microliters of 95% of ethanol and the optical density was measured at 570 nm by enzyme-linked immunoassay reader. The positive Optical Density (OD) was defined as three standard deviations above the mean OD of the negative control. The K. pneumoniae NTUH-K2044 strain was used as positive control.
2.5. Molecular Studies of K. pneumoniae
2.5.1. Extraction of DNA
DNA of the isolated K. pneumoniae was prepared by boiling method. Briefly, colonies were suspended in 100 microliters of sterile distilled water and boiled at 100ºC in the water bath for 15 minutes then rapidly cooled at -20°C for one hour, then centrifugation and the supernatant were preserved for the use in the amplification processes [13].
2.5.2. PCR for Virulence Genes of K. pneumoniae
PCR was performed for the detection of virulence genes for K. pneumoniae. The genes detected and the sequences of the used primers were summarized in Table 1. The conditions of PCR were performed according to Shah et al., 2017 [13] for each gene. Negative controls were included as sterile distilled water and K. pneumoniae ATCC13883 was used as positive control. The products of the amplifications were analyzed by the use of gel electrophoresis 1.5% stained with ethidium bromide and the bands were visualized by UV.
2.5.3. Multiplex PCR for ESBL Genes
The extracted DNA was used for multiplex PCR for the detection of ESBL genes blaTEM, blaSHV and blaCTX-M. The sequences of the used primers were shown in Table 1. The amplification steps were primary denaturation at 94°C for 4 minutes, then 35 cycles of amplification (94°C for 45 seconds, 55°C for 45 seconds and 72°C for 1 minute) with final extension at 72°C for 7 minutes. K. pneumoniae ATCC 7881 was used as a positive control. The products of the amplifications were analyzed by the use of gel electrophoresis 1.5% stained with ethidium bromide and the bands were visualized by UV [13].
2.5.4. Statistical Analysis
Data were collected, revised, coded and entered to the Statistical Package for Social Science (SPSS) version 24. The quantitative data were presented as mean, standard deviations and ranges and qualitative data was presented by number and percentages. The comparison between qualitative data was done by the use of chi-square. P-value was considered significant if it was <0.05.
3. RESULTS
The study included non-duplicate 100 isolates of K. pneumoniae. The majority of the isolates had the capacity to form biofilm (81 isolates, 81%) with ESBL prevalence rate 41% (41 isolates). Forty-one isolates had the combined capacity to form biofilm and were ESBL, 40 isolates had the capacity to form biofilm and were non ESBL producers and 19 isolates were non ESBL and lack the capacity to form biofilm (Fig. 1).
Table 1.
| Gene | Sequence | bp | Positive Control |
|---|---|---|---|
| FimH-1 | F: 5/-ATG AAC GCC TGG TCC TTT GC-/3 R: 5/-GCT GAA CGC CTA TCC CCT GC-/3 |
688 | E. coli 700336 |
| mrkD | F: 5/-CCA CCA ACT ATT CCC TCG AA -/3 R: 5/-ATG GAA CCC ACA TCG ACA TT-/3 |
240 | K. pneumoniae ATCC BAA-2473 |
| magA (K1) | F: 5/-GGT GCT CTT TAC ATC ATT GC-/3 R: 5/-GCA ATG GCC ATT TGC GTT AG-/3 |
1282 | Klebsiella pneumoniae subsp. pneumoniae Strain K6 ATCC 700603 |
| wzy (K2) | F: 5/-GAC CCG ATA TTC ATA CTT GAC AGA G-/3 R: 5/-CCT GAA GTA AAA TCG TAA ATA GAT GGC-/3 |
641 | Klebsiella pneumoniae subsp. pneumoniae Strain K6 [ATCC® 700603™ |
| rmpA | F: 5/-ACT GGG CTA CCT CTG CTT CA -/3 R: 5/-CTT GCA TGA GCC ATC TTT CA-/3 |
535 | Klebsiella pneumoniae 1100770 |
| uge | F: 5/-TCT TCA CGC CTT CCT TCA CT-/3 R: 5/-GAT CAT CCG GTC TCC CTG TA-/3 |
534 | K. pneumoniae ATCC BAA2473 |
| kfu | F: 5/- GAA GTG ACG CTG TTT CTG GC -/3 R: 5/-TTT CGT GTG GCC AGT GAC TC-/3 |
797 | Klebsiella pneumoniae subsp. pneumoniae (Schroeter) Trevisan ATCC13882 |
| Aerobactin | F: 5/-GCA TAG GCG GAT ACG AAC AT-/3 R: 5/- CAC AGG GCA ATT GCT TAC C T-/3 |
556 | Klebsiella pneumoniae subsp. pneumoniae Strain K6 [ATCC 700603 |
| blaTEM | F: 5/-ATA AAA TTC TTG AAG ACG AAA -/3 R: 5/- GAC AGT TAC CAA TGC TTA ATC -/3 |
1080 | K. pneumoniae ATCC 7881 |
| blaSHV | F: 5/- GGG TTA TTC TTA TTT GTC GC -/3 R: 5/-TTA GCG TTG CCA GTG CTC-/3 |
930 | K. pneumoniae ATCC 7881 |
| blaCTX-M | F: 5/-GCT ATG TGC AGT ACC AGT AA-/3 R: 5/-ACC AGA ATG AGC GGC GC-/3 |
585 | K. pneumoniae ATCC 7881 |
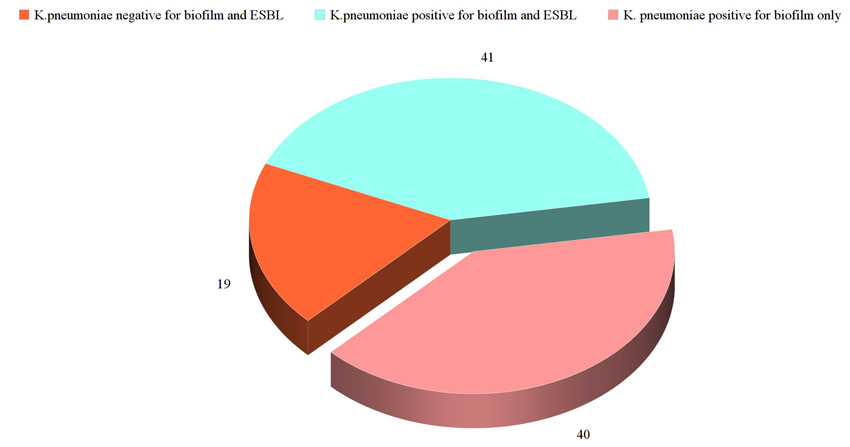
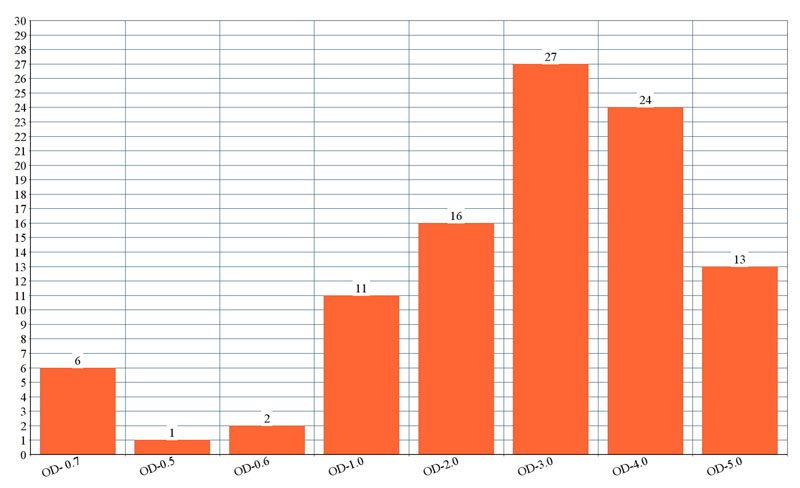
Fig (2). shows the optical density of biofilm formed by each isolate. The genes detected among ESBL producing K. pneumoniae was blaCTX-M in 30 isolates followed by blaSHV 22 isolates and blaTEM in 21 isolates. Multiple genes were present in 25 of the isolates. Among the virulent genes' studies in K. pneumoniae isolates, the most prevalent gene was fimH (76%), uge (70%), rmpA (62%), mag (59%), wzy (59%) and aerobactin gene (51%). Lower prevalence was detected for and kfa gene (33%).
There was significant association between ESBL production, and resistance to amikacin, cefepime, ceftazidime, gentamicin, imipenem and meropenem and biofilm-producing K. pneumoniae data not shown. There was a significant association between blaCTX-M, blaSHV, fimH,magA, kfa, wzy, rmpA and aerobact and biofilm-producing K .pneumoniae (Table 2).
| ESBLs and Other Virulence Genes |
Biofilm forming K. pneumoniae(n=81) No. (%) |
Non-Biofilm forming K. penumoniae (n=29) No. (%) |
P |
|---|---|---|---|
| blaTEM | 12 (14.8%) | 9 (31%) | P=0.007 |
| blaSHV | 11 (13.6%) | 11 (37. 9%) | P=0.005 |
| blaCTX-M | 30 (37%) | 0 (0%) | P=0.0002 |
| fimH | 72 (88.9%) | 4 (13.8%) | P=0.0001 |
| magA | 49 (60.5%) | 10 (34.4%) | P=0.0001 |
| wzy | 49(60.5%) | 10(34.4%) | P=0.0001 |
| rmpA | 58 (71.6%) | 4 (13.8%) | P=0.0001 |
| uge | 59 (72.8%) | 11 (37.9%) | P=0.2 |
| kfa | 33 (40.7%) | 0 (0%) | P=0.0001 |
| aerobact | 48 (59.3%) | 3 (10.3%) | P=0.0001 |
4. DISCUSSION
Klebsiella pneumoniae has emerged as an important pathogen causing around 20% of infections in the hospitals worldwide [14].
There are several factors that increase the virulence of K. pneumoniae, of which the capacity to form biofilm is prevalent. The majority of the isolates in the present study had the capacity to form biofilm (81%). This finding was similar to the previous report concerning isolates from urinary tract infections [15]. The formation of biofilm protects K. pneumoniae species against the host immune responses, the action of antibiotics and enhance its persistence [16, 17].
One of the other factors associated with infection by K. pneumoniae is the emergence of resistance to antibiotics by different mechanisms. The common mechanism is ESBL production. In the present study, the prevalence of ESBL production was 41%. The prevalence of ESBL production among K. pneumoniae species varied from 30% up to 65% in different studies [18, 19]. The variations of the prevalence of ESBL depend upon the empirical use of antibiotics in local regions with positive pressure on K. pneumoniae leading to the selection of resistant strains.
The most prevalent genes among ESBL producing K. pneumoniae were blaCTX-M followed by blaSHV and blaTEM. Several reports had a similar finding with blaCTX-M was the most prevalent gene followed by blaTEM and blaSHV [20, 21]. The high prevalence of blaCTX-M among K. pneumoniae may represent selective pressure due to the wide prescriptions of cephalosporins, particularly cefotaxime and ceftriaxone in many geographical regions including Egypt [22]. However, the prevalence rates vary not only between different countries but even between different hospitals in the same country [23].
The ESBL enzymes are encoded by plasmid genes, therefore, it is expected for one isolate to carry different genes as the mixed genes were present in 65.8% of the isolates in our study. Similar findings were reported by [20]. Further studies are required to understand the mechanisms mediated by multiple bla genes on the same isolate.
The present study revealed that the most prevalent virulence genes among K. pneumoniae species were fimH (76%), uge (70%), rmpA (62%), mag (59%) andwzy (59%) and the lowest prevalence was detected for aerobactin gene (50%) and kfa gene (33%). Similar prevalence rates were reported for fimH, uge and rmpA genes [23, 24]. The prevalence of aerobactin gene varies in different studies from 3% up to 92.8% [25, 26]. The difference in the prevalence of virulence genes appears to be associated with the difference of the clinical sources of the isolated K. pneumoniae species.
Several studies demonstrated an association between the aerobactin gene and the regulatory genes of mucous in K. pneumoniae [26, 27]. There are cumulative data that the virulence of K. pneumoniae is enhanced by the presence of aerobactin due to the presence of other multiple virulence genes on the same plasmid carrying it [27].
There was a significant association between resistance to amikacin, cefepime, ceftazidime, gentamicin, imipenem and meropenem and biofilm production by K. pneumoniae. The finding was explained by several mechanisms such as the adherent bacterial cells are resistant to the actions of different antibiotics due to the limitation of the penetration of the antibiotics, beside the slow growth of the bacterial cells inside the biofilm [16]. Moreover, the exchange of plasmids that carry antibiotics resistance genes between contact microbial cells may play role [28, 29]. ESBL production by isolated K. pneumoniae species was significantly associated with biofilm formation. This finding was similar to previous reports [29, 30]. Again, this finding can be explained by the formation of protective environment to K. pneumoniae away from the antibiotics and the transmission of resistant genes. This was confirmed in the present study by significant association of blaSHV and blaCTX-M genes with biofilm formation.
Virulence genes of K. pneumoniae, fimH, magA, kfa, rmpA and aerobactin had a significant association with biofilm formation. Several studies correlated the association of genes of fimbriae, capsule and lipopolysaccharides with the biofilm formation by K. pneumoniae through initial adhesion by fimbriae and lipopolysaccharide followed by coverage of substratum and construction of mature biofilm architecture by the capsule [31, 32].
CONCLUSION
The present study highlights the prevalence of virulence genes among biofilm-forming strains of K. pneumoniae isolated from hospital-acquired urinary tract infections. Moreover, there was an association between biofilm formation and ESBL production. Further studies are required to elucidate the clinical impact of the association of these different mechanisms.
ETHICAL APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by Mansoura Faculty of Medicine ethical committee, Egypt.
HUMAN AND ANIMAL RIGHTS
No Animals were used in this research. All human research procedures followed were in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national), and with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Informed consent was obtained from all the participants prior to publication.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
UGC-BSR Start-up grant by University Grant Commission, New Delhi, India.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none


