All published articles of this journal are available on ScienceDirect.
Study of Antifungal Susceptibility, Virulence Genes and Biofilm Formation in Candida albicans
Abstract
Background:
Candida albicans has emerged as an important nosocomial pathogen. The morbidity and mortality associated with this pathogen are related to the presence of virulence genes and antifungal resistance. The objective of the present study was to investigate the prevalence of antifungal resistance, biofilm formation and some virulence genes such as ALS1, PLB1, INT1, SAP1 and HWP1, among clinical isolates of Candida albicans recovered from immunocompromised patients.
Methods:
The study included one hundred C. albicans isolates identified phenotypically and by a molecular technique using Polymerase Chain Reaction (PCR). The identified C. albicans was further subjected to antifungal study by the microdilution method, biofilm study and molecular study for virulence genes by PCR.
Results:
The resistance to antifungal drugs, fluconazole, caspofungin and itraconazole was 8% for each of them and for amphotericin B, it was 9%. The prevalence of the studied virulence genes was HWP1 77%, INT1 72%, ALS1 65%, SAP1 65% and PLB1 52%. The biofilm capacity was identified by the microplate method in 58% of C. albicans. The OD was intense in 20 isolates, moderate in 21 isolates and mild in 17 isolates. There was a statistically significant increase in the prevalence of the studied virulence genes INT1, ALS1, HWP1, SAP1 and PLB1 among biofilm forming C. albicans as compared to non-biofilm forming isolates (P=0.0001). Additionally, the resistance to fluconazole, itraconazole and caspofungin was statistically, significantly higher in C. albicans with the capacity to form biofilm as compared to non-biofilm forming C. albicans.
Conclusion:
The present study highlights the prevalence of resistance to antifungal drugs among C. albicans which are not uncommon. Moreover, there was a high prevalence of INT1, ALS1, HWP1, SAP1 and PLB1 genes in C. albicans. The resistance to antifungal drugs was common among isolates with the capacity to form the biofilm. There was an association between the biofilm formation and virulence genes.
1. INTRODUCTION
Candida genus is a polymorphic diploid fungus and considered a part of human microbiota [1]. However, in certain pathogenic conditions, it can be a leading cause of infection either in superficial infections of skin and mucous membranes or systemic invasive infections. Candida species are the fourth cause of nosocomial bloodstream infections with a mortality rate of up to 40% [2, 3].
There are several Candida species such as Candida albicans (C. albicans), Candida glabrata, Candida tropicalis, and Candida parapsilosis, but C. albicans remains the most frequent species isolated from bloodstream infections [4].
There are several predisposing factors for invasive infections with Candida species as the presence of patient’s immunocompromised conditions [5], physical alteration of the normal barrier such as surgery and overgrowth of Candida species after broad-spectrum antibiotics intake with the reduction of normal bacterial microbiota [6]. Early laboratory diagnosis of Candida invasive infections reduces the mortality rate from 40% to 15% after the start of appropriate antifungal therapy [7].
Several pathogenic virulence factors are encoded by C. albicans genes and assist the fungus to invade the host tissues leading to infections. Among these virulence factors are the capacity of C. albicans to change from the budding yeast form to filamentous form, the expression of adhesion factors, the ability to form biofilm and the secretion of hydrolytic enzymes [8].
In C. albicans, the adhesion and biofilm formation are supposed to be mediated through the hyphal wall protein 1(Hwp1) located on the surface of the hyphae [9]. The gene encoding Hwp1 is part of a core of eight genes induced during filamentation of C. albicans [10]. The adhesion of C. albicans is also mediated by agglutinin-like sequence 1 gene (ALS1) that mediates attachment to endothelial cells. It is also important for hyphal formation as well as adherence [11]. There are two genes with similar sequences and functions, ALS1 and ALS3 [12].
Additionally, the phospholipases protein 1 (PLB1 p) facilitates the invasion of Candida albicans through gastrointestinal tract leading to systemic infection [13]. Hydrolysis of peptide bonds of the host proteins is mediated by the secreted aspartyl proteinases (Saps), suggested to be involved in the virulence of Candida albicans [14]. The INT1 gene is a virulence factor that contributes to the ability of Candida albicans to adhere to epithelial cells and form hyphae [15, 16].
The presence of virulence genes and the increasing prevalence of resistance to antifungal therapy have contributed to the pathogenicity of Candida albicans. Nevertheless, the association between the resistance profiles and virulence genes of C. albicans has not been studied sufficiently [17], especially in invasive C. albicans infections.
The objective of the present study was to investigate the prevalence of antifungal resistance, biofilm formation and some virulence genes, such as ALS1, PLB1, INT1, SAP1 and HWP1 among clinical isolates of C. albicans.
2. MATERIALS AND METHODS
This study included one hundred Candida albicans isolates from patients with nosocomial infections under chemotherapy for the treatment of malignancies admitted to intensive care units of Mansoura Oncology Centre, Mansoura city, Egypt during March 2017 till January 2019. The healthcare-associated infections were actively identified by the National Surveillance System supported by Supreme Council of Egyptian Universities in cooperation with Center of Disease Control. The study was approved by the Mansoura Faculty of Medicine Ethical Committee and approvals were obtained from the participating patients.
2.1. Samples and their Processing
The study samples were ten-milliliter blood samples obtained from each patient under complete antiseptic conditions and inoculated to two blood cultures for fungal culture, Mycosis-IC/F blood culture bottles. Urine samples were obtained as mid-stream urine sample in sterile container or from the catheter by sterile syringe from the sampling port on the catheter bag.
The isolated C. albicans was recovered from 56 blood cultures using BACTEC 9240 system and Mycosis-IC/F blood culture (Becton Dickinson Diagnostics-Heidelberg Germany). In addition, 44 isolates were recovered from urine cultures on Sabouraud dextrose media (Oxoid, Hampshire RG24 8PW, United Kingdom) with colony counts > 105 colony forming units/ml.
Phenotypic identification of C. albicans was performed by API20C biochemical reaction, the ability to form germ tube, the use of positive growth on hypertonic Sabouraud broth and colony character on the culture of tobacco agar media. Further molecular identification was carried out through a molecular method by the use of Polymerase Chain Reaction (PCR). The identified C. albicans was further subjected to antifungal study by microdilution method, biofilm study and molecular study for virulence genes by PCR.
2.2. Phenotypic Identification of Candida albicans
2.2.1. Growth on Hypertonic Sabouraud Broth.
Hypertonic Sabouraud broth was inoculated with the isolated colonies from Sabouraud dextrose agar and incubated at 28˚C for 96 hours with a daily check for growth. Positive Candida growth indicated C. albicans species [18]. C. albicans (ATCC No. 90028) was used as a positive quality control strain.
2.2.2. Growth on Tobacco Agar
The subculture of the isolated Candida species was performed on Tobacco agar and the plates were incubated for 96 hours at 28˚C with a daily check for growth. White creamy smooth growth was identified as C. albicans [19].
2.2.3. Biofilm Study
Pure C. albicans colonies were cultured on Sabouraud broth at 28oC with a continuous shake for 18 hours. Then a suspension was prepared from C. albicans using RPMI 1640 at the concentration of 106/ml and 100 μl of the suspension was inoculated into a 96-well microtiter plate in triplicate. Then the plate was incubated at 37oC for 48 hours. Negative control wells were inoculated with RPMI 1640 without C. albicans. Each well was then washed with 200 μl phosphate buffer, air-dried and stained with 110 μl 0.4% crystal violet solution for 45 min. Subsequently, each well was washed four times with 350 μl sterile distilled water then 200 μl of 95% ethanol was added and left for 45 min. After that, 100 μl of the solution in each well was transferred to a new well and the optical density was measured with a microplate reader (Stat Fax 2100) at 595 nm. The absorbance values for the controls were subtracted from the values for the test wells [20].
Four biofilm density categories were used to grade the biofilms based on the established optical density cut-off values (ODc), which were derived from the mean values of negative controls (mean ODnc) summed with three standard deviations of the negative controls (3 × SDnc): ODc = mean ODnc + (3 × SDnc). The biofilm density categories used are as follows: OD ≤ ODC = biofilm negative, ODC < OD ≤ 2 x ODC = mildly positive for biofilm, 2 x ODC < OD ≤ 4 x ODC = moderately positive for biofilm, OD > 4 x ODC = intensely positive for biofilm [21].
2.2.4. Minimum Inhibitory Concentration (MIC) determination for C. albicans
The susceptibility of the isolated C. albicans to amphotericin B, fluconazole, itraconazole and caspofungin was determined by the microdilution method. The minimum inhibitory concentrations were determined by the broth microdilution method as described by the Clinical and Laboratory Standards Institute [22]. C. parapsilosis ATCC 22019 and C. krusei ATCC 6258 were included as quality controls for each test performed.
2.3. Molecular Study for the Isolated C. albicans
2.3.1. DNA Extraction for Fungi
DNA was extracted from the isolated C. albicans using Qiagen DNA extraction kit according to the recommendations of the manufacturer (Qiagen-Germany).
2.3.2. Identification of C. albicans
Extracted DNA was amplified by specific primers for C. albicans. The used primers are listed in Table 1. The amplification procedures were as described previously [23].
2.3.3. Detection of Virulence Gene Markers by PCR
Extracted DNA was used for amplification of HWP1, ALS1, αINT1, SAP1 and PLB1 virulence genes. The genes and the sequences of the used primers are listed in Table 1. The amplification procedures for each of the studied genes were as described previously and the used amplification mixtures were from Qiagen in addition to 2.5 μl of each primer and 5 μl of the extracted DNA [23]. The PCR amplification products were visualized by electrophoresis on 1% agarose gels for 45 min at 70v. The size of the amplicons was determined by comparison to the 100 bp DNA ladder (Promega, USA).
| Gene | Sequence of Primers | Size of Amplicon (bp) |
|---|---|---|
| CALB1 | FP/TTTATCAACTTGTTGTCACACCAGA RP/ATCCCGCCTTACCACTACCG |
273 |
| HWP1 | FP/ATGACTCCAGCTGGTTC RP/TAGAT CAAGAATGCAGC |
572 |
| ALS1 | FP/GACTAGGAACAACAAATACCAGA RP/CCAGAAGAAACAGCAGGTGA |
318 |
| INT1 | FP/AAGCTCTGATACCTACACTAGCGA RP/GTTAGGTCTAAAGTCGAAGTCATC |
239 |
| SAP1 | FP/GCTCTTGCTATTGCTTTATTA RP/CATCAGGAACCCATAAATCAG |
253 |
| PLB1 | FP/ATGATTTTGCATCATTTG RP/AGTATCTGGAGCTCTAC |
751 |
3. RESULTS
This study included 100 C. albicans isolates that were identified by phenotypic and molecular methods using a specific gene by PCR. The isolates were isolated from nosocomial infections of 56 patients with sepsis and 44 patients with urinary tract infections.
The resistance of the isolated C. albicans to antifungal drugs, fluconazole, caspofungin and itraconazole was 8% for each of them and 9% for amphotericin B (Table 2) using the microdilution method.
| Antifungal | No. | % |
|---|---|---|
| Fluconazole | 8 | 8% |
| Amphotericin B | 9 | 9% |
| Caspofungin | 8 | 8% |
| Itraconazole | 8 | 8% |
The prevalence of the studied virulence genes in C. albicans isolates was HWP1 77%, INT1 72%, ALS1 65%, SAP1 65% and PLB1 52% using PCR (Table 3, Fig. 1).
| Genes | Frequency | % |
|---|---|---|
| HWP1 | 77 | 77% |
| INT1 | 72 | 72% |
| ALS1 | 65 | 65% |
| SAP1 | 65 | 65% |
| PLB1 | 52 | 52% |
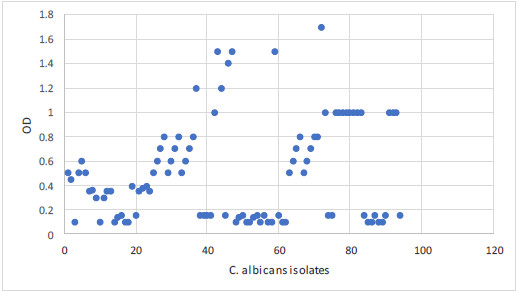
The biofilm capacity was identified by the microplate method in 58% of C. albicans isolates. The OD was intense in 20 isolates, moderate in 21 isolates and mild in 17 isolates. The optical density of all isolates is summarized in Figs. 2, 3.
The resistance to the tested antifungal drugs increased in C. albicans with the capacity to form biofilm as compared to non-biofilm forming C. albican; this increase for fluconazole was 12.1% in biofilm forming C. albicans versus non biofilm forming isolates (2.4%), itraconazole (12.1%) in biofilm forming C. albicans versus non biofilm forming isolates (2.4%), amphotericin B (13.8%) in biofilm forming C. albicans versus non biofilm forming isolates (2.4%) and caspofungin (12.1%) in biofilm forming C. albicans versus non biofilm forming isolates (2.4%), (Table 4). Additionally, there was a statistically significantly high prevalence of the studied virulence genes, INT1, ALS1, HWP1, SAP1 and PLB1 (98.3%, 96.6%, 96.6%, 96.6%, and 81.03%, respectively) among biofilm forming C. albicans as compared to non-biofilm forming isolates (35.7%, 21.4%, 50%, 21.4%, and 11.9%, respectively), P=0.0001, (Table 4).
| – |
Biofilm forming C. albicans (No = 58) No. % |
Non-biofilm forming C. albicans (No = 42) No. % |
P |
|---|---|---|---|
|
Sepsis Urine |
33 56.9% 25 43.1% |
23 54.8% 19 45.2% |
P = 0.5 |
| Fluconazole resistance | 7 12.1% | 1 2.4% | P = 0.1 |
| Itraconazole resistance | 7 12.1% | 1 2.4% | P = 0.1 |
| Amphotericin B resistance | 8 13.8% | 1 2.4% | P = 0.05 |
| Caspofungin resistance | 7 12.1% | 1 2.4% | P = 0.1 |
| INT1 | 57 98.3% | 15 35.7% | P = 0.0001 |
| ALS1 | 56 96.6% | 9 21.4% | P = 0.0001 |
| HWP1 | 56 96.6% | 21 50% | P = 0.0001 |
| SAP1 | 56 96.6% | 9 21.4% | P = 0.0001 |
| PLB1 | 47 81.03% | 5 11.9% | P = 0.0001 |
4. DISCUSSION
In the last few years, Candida species particularly C. albicans has emerged as an important nosocomial pathogen [24]. The differentiation between different Candida species depends mainly upon the biochemical identification, germ tube formation and the specific culture appearance on specific media [25].

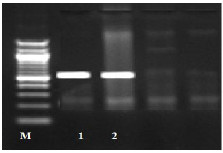
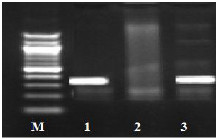
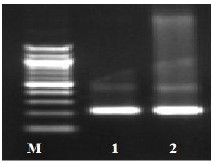
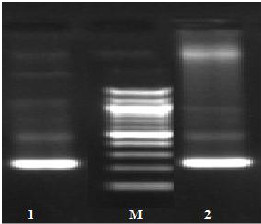
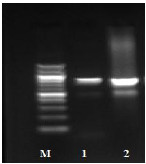
Fig. (1) Detection of different studied genes in the isolated C. albicans by PCR.
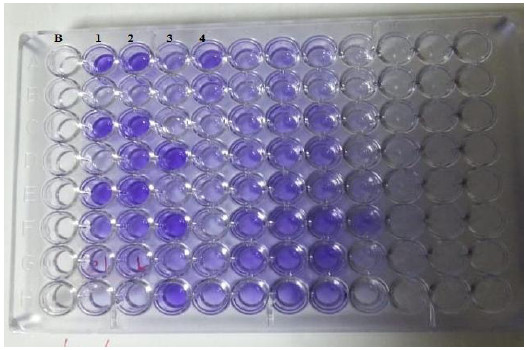
B: blank, Wells 1, 2: strong positive, Well 3: weak positive, Well 4: intermediates positive.
Similarly, in the present study, the identification of Candida albicans was performed by the biochemical reactions, germ tube and culture characters on specific media. Moreover, molecular identification was used to confirm the species of C. albicans. The primers used in the molecular identification were designed based on the sequence data for the Internal Transcribed Spacer (ITS) region-specific for C. albicans as shown previously [26].
The azoles are used for the treatment of fungal infections. These compounds exert their effects through the inhibition of ergosterol biosynthesis in the fatty cell membranes [27]. On the other hand, the antifungal effects of amphotericin B are directed through sequestration of ergosterol in the fungal cell membranes leading to its death and also through accumulation of reactive oxygen species which has been associated with cell damage, apoptosis induction and death [28].
In our study, the resistance to antifungal drugs, fluconazole, caspofungin and itraconazole was 8% for each of them and 9% for amphotericin B. The resistance of Candida species to different antifungal drugs either azoles or amphotericin B varied in different studies from 2.8% up to 80% [24, 29, 30]. Cross-resistance should be considered for some antifungal agents from the azole family [31]. The existing difference may be due to population or regional studies. Moreover, fluconazole in some immunocompromised patients was used as prophylaxis, which can explain the differences in resistance rates in some populations. Therefore, the present study highlights the importance to incorporate the antifungal susceptibility tests as a routine laboratory test for the management of systemic infection by C. albicans.
In the present work, the prevalence of the studied virulence genes was HWP1 77%, INT1 72%, ALS1 65%, SAP1 65% and PLB1 52%. A previous study revealed higher frequency for INT1, ALS1 (100%) and HWP1 (90.9%) with lower frequency of SAP1 (59.09%) and PLB1 (13.63%) [23]. Different prevalence rates for the virulence genes have been reported in the literature ranging from 5% to 100% [32, 33]. The difference in the prevalence rates of virulence genes may be attributed to several factors including the number of isolates studied and the difference of the isolation sites of C. albicans.
The biofilm capacity was identified by the microplate method in 58% of C. albicans. The OD was intense in 20 isolates, moderate in 21 isolates and mild in 17 isolates. The frequency of biofilm formation among isolated C. albicans from immunocompromised patients is similar to the previous study by Vijayalakshmi et al. [25], but higher than other work by Tumbarello et al. [34]. The difference in the frequency of biofilm formation may be attributed to the different in vitro study of biofilm formation. In the present work, the used method was the microplate method with the initial removal of non-adherent C. albicans as the standard method advised [35]. Considering the biofilm-forming capacity of these C. albicans isolates, their pathogenicity could result in severe co-infection with other diseases.
In the present study, there was a high prevalence of antifungal resistance to azoles, amphotericin B and caspofungin in biofilm forming C. albicans. This finding is in line with previous studies [25, 36]. This association between biofilm formation and antifungal resistance may be attributed to several mechanisms such as limited penetration of drug through biofilm matrix, decrease in the rate of C. albicans, need for nutrition and its persistence and increase in rate of the expression of resistance genes induced by contact with a surface and persistence of a small number of cells. The association between the capacity of C. albicans to form biofilm with the presence of resistance to antifungal drugs renders its treatment difficult and necessitates different lines of treatment [37].
This study revealed the statistically significant increase in the prevalence of the studied virulence genes, INT1, ALS1, HWP1, SAP1 and PLB1 among biofilm-forming C. albicans as compared to non-biofilm forming isolates. This was due to the hypothesis that adhesion of C. albicans at the initial step of the biofilm formation involves different virulence factors encoded by virulence genes such as Als proteins [35] and Hwp1 [38, 39], all of which are expressed at much higher levels in hyphae than in yeast cells [37].
Thus, the present study revealed significant association between the virulence genes, antifungal resistance and biofilm formation among C. albicans associated with nosocomial infections
CONCLUSION
The present study highlights the prevalence of resistance to antifungal drugs among C. albicans, which are not uncommon. Moreover, there is a high prevalence of INT1, ALS1, HWP1, SAP1 and PLB1 genes in C. albicans. The resistance to antifungal drugs is common among C. albicans isolates with a capacity to form a biofilm. Additionally, there is an association between biofilm formation and virulence genes.
LIST OF ABBREVIATIONS
| C. albicans | = Candida albicans |
| PCR | = Polymerase Chain Reaction |
| OD | = Optical Density |
| MIC | = Minimum Inhibitory Concentration |
| Bp | = Base Pair |
AUTHORS CONTRIBUTIONS
All authors have contributed to the manuscript.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by the Mansoura Faculty of Medicine Ethical Committee, Egypt.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Informed consent was obtained from all the participants.
AVAILABILITY OF DATA AND MATERIALS
All data generated or analyzed throughout this research are included in this published article.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors would like to thank all support staff and participating patients in this study.


