All published articles of this journal are available on ScienceDirect.
Analysis of a Pool of Small Plasmids from Soil Heterotrophic Cultivable Bacterial Communities
Abstract
In this work the analysis of the plasmid presence on soil aerobic cultivable heterotrophic bacterial communities was carried out checking a panel of 1,200 isolates, in order to establish the frequency of plasmid presence as well as the degree of plasmid flow between strains affiliated to the same or different taxon. Bacterial communities were isolated from two different sites of a 13-year experimental field with a clay-silt texture. Plasmid molecules were detected at low frequency (27 isolates, 2%) with a size ranging between 2 Kb and 40 Kb. The RAPD analysis performed on the plasmid-harboring isolates and the phylogenetic analysis of the whole community using the 16S rRNA gene sequences revealed the existence of transfer of the same plasmids between strains belonging to the same species and, in some cases, to different species of the same genus. As it might be expected, even though the viable cells title did not differ significantly between the two samplings, the overall data disclosed an uneven distribution of both species and plasmid-harboring strains.
INTRODUCTION
Soil is a complex and dynamic ecosystem whose functionality is related to the equilibrium existing between chemical, physical, biological parameters and the resident microbial communities. The biodiversity of these communities may undergo fluctuations as a consequence of environmental changes. It has been recognized that one of the key factors responsible for the biodiversity of soil microbial communities and, in general, for microbial evolution is represented by mobile genetic elements (MGE: bacteriophages, transposons and plasmids) that are involved in the horizontal transfer of genetic information (HGT) [1]. In fact, “while point mutations contribute to microbial adaptation, horizontal dissemination of genes has proven to be critical in promoting rapid genomic flexibility and microbial evolution” [2]. Particularly interesting from this viewpoint are plasmids, for the essential role they play in the ecological adaptation of Bacteria and Archaea; indeed, they can contribute to shape prokaryotic genomes, “promoting intra- and inter-species variability and distributing functional genetic modules” [3].
Genomics approaches allowed to disclose a large and untapped diversity of plasmids inhabiting plant-associated or soil bacteria. “Surveys on the presence of plasmids from soil and plant-associated bacteria have been performed and revealed that a considerable portion of bacteria from different environments carried plasmids; for examples, approximately 18% of bacterial isolates from the phytosphere of sugar beets were found to contain plasmids” [4]. Furthermore, although the function of most of plasmid-borne genes is still unknown [5], there is a general agreement that MGE add some, often small or even not measurable, metabolic burden to their host. “Traits conferring an improved fitness or ability to colonize environmental niches are often located on conjugative MGE; consequently, the prevalence of plasmids indicates that they can benefit bacteria in the environment” [6].
Rhizosphere, together with soil, is one of the main “hot” spots for gene transfer activity performed by bacteria. This is due to different factors, including the enhanced nutrient input and water fluxes that might stimulate bacterial metabolic activities. The enhanced conjugative transfer of chromosomal genes between Pseudomonas spp. in the wheat rhizosphere in respect to bulk soil was previously reported [7]. Another example of bacterial metabolic abilities, which very likely have been rather recently evolved and spread through HGT is the capability to perform biodegradation of man-made xenobiotic compounds [8].
The plasmids ecology is still poorly understood and we know little of their distribution and diversity. In spite of the importance of plasmid molecules, an extensive analysis of the presence and frequency of plasmids in large natural cultivable microbial communities has not been performed up to now. Therefore, the aim of this work was to analyze the presence of plasmids in a large heterotrophic cultivable bacterial community isolated from soil and to check the degree of genetic flow between strains belonging to the same or different species/genus. To this purpose we used the experimental strategy schematically represented in Fig. (1).
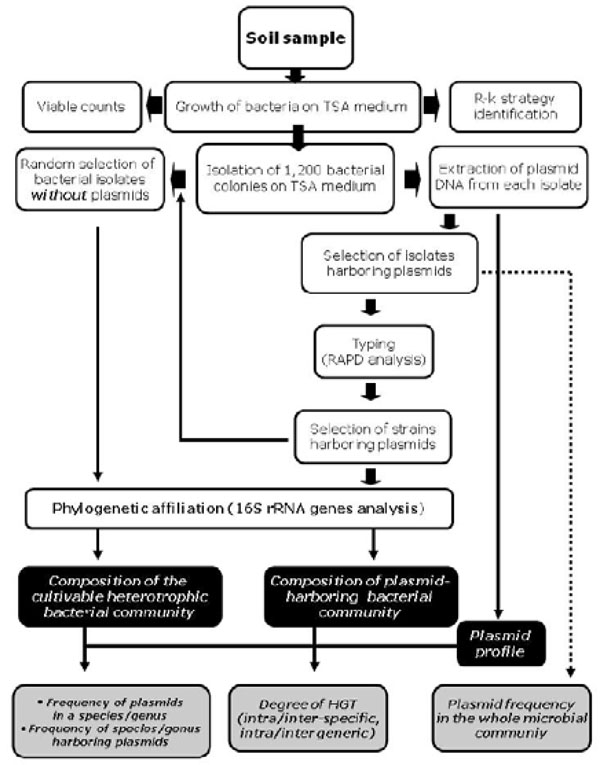
Schematic representation of the overall experimental strategy used in this work.
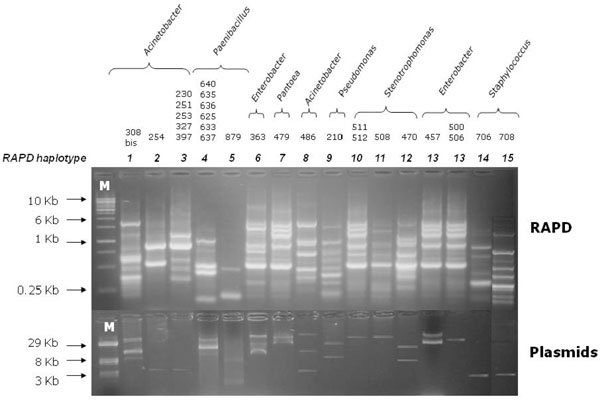
Agarose gel electrophoresis of RAPD profiles (upper) from 27 soil bacterial isolates harboring plasmids (lower). The last line of the right side was cut from another figure and pasted in Fig. (2), without changing neither the intensity nor the size of each band. Line M: (upper) DNA linear marker; (lower) reference plasmids of known size.
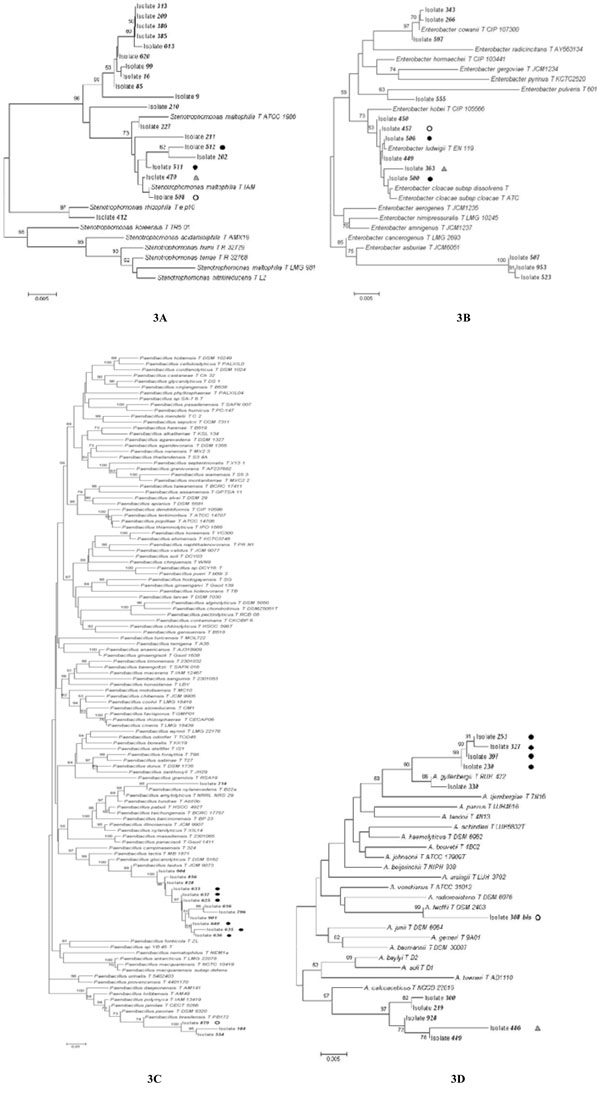
Phylogenetic trees constructed using the 16S rRNA sequences obtained from Stenotrophomonas (A), Enterobacter (B), Paenibacillus (C), and Acinetobacter (D) isolates analyzed in this work and the most similar sequences retrieved from databases. Bootstrap values > 50 are shown. The pairwise deletion option was used. Isolates harboring one or more plasmids are marked by a black or white dot or a grey triangle. Isolates harboring a plasmid with the same electrophoretic mobility are marked with the same symbol.
List of bacterial isolates analysed in this work; the phylogenetic affiliation of each isolate is also reported along with the 16S rRNA gene accession number.
| Isolate | Sampling | Plasmid presence |
16S rDNA accession number |
First Blast Hit | Phylogenetic affiliation |
|---|---|---|---|---|---|
| 1 | 2.2.A | - | GU808446 | AF131549 Streptomyces sp. IM-7082 | Streptomyces |
| 2 | - | GU646899 | JF700422 Stenotrophomonas sp. WR49 | Stenotrophomonas | |
| 9 | - | GU831905 | FR682931 Stenotrophomonas sp. R-41388 | Stenotrophomonas | |
| 16 | - | GU814021 | JF700422 Stenotrophomonas sp.WR49 | Stenotrophomonas | |
| 56 | - | GU814032 | FJ006929 Microbacterium sp. WPCB194 | Microbacterium | |
| 57 | - | GU808432 | EU366363 Bacillus pumilus strain I5 | Bacillus | |
| 85 | - | GU814025 | JF700422 Stenotrophomonas sp. WR49 | Stenotrophomonas | |
| 98 | - | GU808428 | JF521654 Rhodococcus sp. CS1 | Rhodococcus | |
| 99 | - | GU814026 | JF700422 Stenotrophomonas sp.WR49 | Stenotrophomonas | |
| 116 | - | GU831909 | JF825992 Bacillus sp. DP5 (2011) | Bacillus | |
| 120 | - | GU831884 | GU361112 Klebsiella oxytoca strain SHD-1 | Klebsiella | |
| 154 | - | GU831895 | FM162997 Microbacteriaceae bacterium ACEMC 25-3 | Microbacterium | |
| 171 | - | GU831910 | HQ257249 Bacillus sp. SG3 | Bacillus | |
| 184 | - | GU831890 | HQ317157 Paenibacillus polymyxa strain DYJL14 | Paenibacillus | |
| 202 | 2.2.B | - | GU814020 | HQ185398 Stenotrophomonas maltophilia strain 5517 | Stenotrophomonas |
| 209 | - | GU831908 | HQ877451 Stenotrophomonas sp. 33T | Stenotrophomonas | |
| 210 | + | GU646895 | GU646895 Pseudomonas geniculata | Pseudomonas | |
| 211 | - | GU814019 | GQ381282 Stenotrophomonas sp. TC7 | Stenotrophomonas | |
| 219 | - | GU646916 | HQ406755 Acinetobacter sp. TY14McD | Acinetobacter | |
| 220 | - | GU814034 | GQ369018 Microbacterium sp. T0-YC6750 | Microbacterium | |
| 227 | - | GU646917 | HQ647282 S. maltophilia strain TS51 | Stenotrophomonas | |
| 230 | + | GU646884 | FJ263916 Acinetobacter johnsonii strain BA2 | Acinetobacter | |
| 238 | - | GU831885 | EU445236 Agrobacterium tumefaciens isolate EFLRI 54 | Agrobacterium | |
| 253 | + | GU646885 | FJ263916 Acinetobacter johnsonii strain BA2 | Acinetobacter | |
| 266 | - | GU831891 | EF120473 Enterobacter cloacae | Enterobacter | |
| 300 | - | GU814027 | FM207522 Acinetobacter calcoaceticus strain CWS20 | Acinetobacter | |
| 308 bis | + | GU646904 | FN393790 Acinetobacter lwoffii strain ES-117-3 | Acinetobacter | |
| 313 | - | GU814024 | HQ877451 Stenotrophomonas sp. 33T | Stenotrophomonas | |
| 327 | + | GU646886 | FJ263916 Acinetobacter johnsonii strain BA2 | Acinetobacter | |
| 330 | - | GU646918 | AY366481 Acinetobacter sp. C1010 | Acinetobacter | |
| 343 | - | GU831896 | EU855207 Enterobacter sp. CTSP29 | Enterobacter | |
| 363 | + | GU646906 | AB461770 Enterobacter sp. M429 | Enterobacter | |
| 385 | - | GU831907 | HQ877451 Stenotrophomonas sp. 33T | Stenotrophomonas | |
| 386 | - | GU831906 | HQ877451 Stenotrophomonas sp. 33T | Stenotrophomonas | |
| 397 | + | HM046411 | FJ263916 Acinetobacter johnsonii strain BA2 | Acinetobacter | |
| 408 | 2.2.C | - | GU814036 | EU714376 Microbacterium foliorum strain 720 | Microbacterium |
| 412 | - | GU814023 | AY599705 Stenotrophomonas sp. TB4-3-II | Stenotrophomonas | |
| 440 | - | GU814035 | DQ530139 Microbacterium sp. RI48 | Microbacterium | |
| 449 | - | GU808436 | JN009619 Enterobacter sp. lb11 | Enterobacter | |
| 450 | - | GU808435 | EU999992 Enterobacter ludwigii strain T4384 | Enterobacter | |
| 457 | + | GU646898 | EU999992 Enterobacter ludwigii strain T4384 | Enterobacter | |
| 458 | - | GU808431 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 470 | + | GU646887 | HQ185399 Stenotrophomonas maltophilia strain 2681 | Stenotrophomonas | |
| 479 | + | GU646888 | HQ694446 Pantoea agglomerans strain AIMST 4.P1.4 | Pantoea | |
| 486 | + | GU646905 | FR774919 Acinetobacter sp. R-45867 | Acinetobacter | |
| 489 | - | GU814028 | AB619594 Acinetobacter sp. NCCP 233 | Acinetobacter | |
| 500 | + | GU646900 | EU999992 Enterobacter ludwigii strain T4384 | Enterobacter | |
| 505 | - | GU831899 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 506 | + | GU646907 | EU999992 Enterobacter ludwigii strain T4384 | Enterobacter | |
| 507 | - | GU808439 | EU430751 Enterobacter sp. ZJUPD5 | Enterobacter | |
| 508 | + | GU646889 | HQ185399 Stenotrophomonas maltophilia strain 2681 | Stenotrophomonas | |
| 511 | + | GU646897 | HQ647282 S. maltophilia strain TS51 | Stenotrophomonas | |
| 512 | + | GU646901 | HQ185399 Stenotrophomonas maltophilia strain 2681 | Stenotrophomonas | |
| 523 | - | GU808438 | EU430751 Enterobacter sp. ZJUPD5 | Enterobacter | |
| 524 | - | GU831901 | HM771092 Bacillus sp. INBio3686F | Bacillus | |
| 539 | - | GU831898 | JN006263 Bacillus sp. C-21 | Bacillus | |
| 546 | - | GU831892 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 554 | - | GU808442 | DQ298127 Paenibacillus polymyxa isolate U4D | Paenibacillus | |
| 555 | - | GU808437 | HM598440 Enterobacter sp. UFLA81 | Enterobacter | |
| 559 | - | GU814030 | HM355676 Microbacterium foliorum strain BAC3087 | Microbacterium | |
| 589 | - | GU831897 | JN006263 Bacillus sp. C-21 | Bacillus | |
| 597 | - | GU808434 | FN401343 Enterobacter cloacae isolate PHLTA-11 | Enterobacter | |
| 603 | 4.1.A | - | GU646913 | DQ530139 Microbacterium sp. RI48 | Microbacterium |
| 613 | - | GU814022 | HQ877451 Stenotrophomonas sp. 33T | Stenotrophomonas | |
| 616 | - | GU808444 | AM983496 Paenibacillus sp. AM27T2 | Paenibacillus | |
| 619 | - | GU808433 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 620 | - | GU831904 | FR682931 Stenotrophomonas sp. R-41388 | Stenotrophomonas | |
| 625 | + | GU646890 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 630 | - | GU814033 | DQ530139 Microbacterium sp. RI48 | Microbacterium | |
| 633 | + | GU646891 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 635 | + | GU646896 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 636 | + | GU646909 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 637 | + | GU646892 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 640 | + | GU646908 | EU880530 Paenibacillus sp. PRE17 | Paenibacillus | |
| 659 | - | GU646911 | FR823407 Bacillus sp. ITCr36 | Bacillus | |
| 675 | - | GU808430 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 706 | + | GU646893 | HM629374 Staphylococcus sp. B-G-R2A2 | Staphylococcus | |
| 708 | + | GU646894 | JF766691 Staphylococcus sp. BIHB 1375 | Staphylococcus | |
| 711 | - | GU831887 | DQ232617 Agromyces sp. VKM Ac-1802 | Agromyces | |
| 714 | - | GU808441 | AY337581 Paenibacillus sp. CC-SB818D1 | Paenibacillus | |
| 737 | - | GU831911 | FM992644 Bacillus safensis strain F5-77 | Bacillus | |
| 773 | - | GU831886 | DQ440827 Bosea sp. CRIB-12 | Bosea | |
| 776 | - | GU831888 | FN563149 Rhodococcus equi 103S | Rhodococcus | |
| 777 | - | GU831912 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 796 | - | GU808445 | JF798384 Paenibacillus lautus strain T1-11 | Paenibacillus | |
| 828 | 4.1.B | - | GU808443 | AJ746160 Paenibacillus sp. MG103 | Paenibacillus |
| 829 | - | GU831894 | JN006263 Bacillus sp. C-21 | Bacillus | |
| 836 | - | GU831903 | DQ026647 Streptomyces exfoliatus strain NRRL B-2494 | Streptomyces | |
| 856 | - | GU808429 | FR667174 Paenibacillus sp. ITP26 | Paenibacillus | |
| 860 | - | GU646912 | AM990746 Bacillus sp. MOLA 522 | Bacillus | |
| 874 | - | GU831900 | FR823407 Bacillus sp. ITCr36 | Bacillus | |
| 879 | + | GU646902 | EU362611 Paenibacillus polymyxa isolate TN99 | Paenibacillus | |
| 890 | - | GU831902 | JF772519 Sinorhizobium sp. bB42(2011) | Sinorhizobium | |
| 901 bis | - | GU831889 | GU097198 Paenibacillus sp. AT5 | Paenibacillus | |
| 904 | - | GU646910 | JF798384 Paenibacillus lautus strain T1-11 | Paenibacillus | |
| 917 | - | GU831893 | JN006263 Bacillus sp. C-21 | Bacillus | |
| 924 | - | GU814031 | DQ530139 Microbacterium sp. RI48 | Microbacterium | |
| 928 | - | GU814029 | HQ132733 Acinetobacter sp. Ld5 | Acinetobacter | |
| 953 | - | GU808440 | EU430751 Enterobacter sp. ZJUPD5 | Enterobacter | |
| 987 | - | GU831913 | JN036709 Bacillus thuringiensis strain AIMST KBT9-X | Bacillus | |
| 1003 | 4.1.C | - | GU831914 | GU252111 Bacillus sp. 2008724139 | Bacillus |
| 1005 | - | GU831915 | JF820106 Bacillus sp. PG-3-9 | Bacillus | |
| 1126 | - | GU646914 | EU729736 Arthrobacter aurescens strain MM10 | Arthrobacter | |
| 1128 | - | GU646915 | HQ597008 Arthrobacter aurescens strain ABRIINW 23 | Arthrobacter |
Symbols: +: presence of plasmid molecules; -: absence of plasmid molecules.
Bacteria colony counts (Log cfu g-1 of soil) as they appeared on 0.1 TSA over a period of six days and total culturable bacteria.
| Soil samples | r/K class 1a | r/K class 2a | r/K class 3a | Total culturable |
|---|---|---|---|---|
| r-strategistsb | K-strategistsb | bacteria | ||
| Top | ||||
| 2.2 A | 5.90 | 5.31 | 5.85 | 6.23 |
| 2.2 B | 6.32 | 5.98 | 6.02 | 6.61 |
| 2.2 C | 5.61 | 5.42 | 5.59 | 6.03 |
| Mean ± SEM | 5.94 ± 0.21 c | 5.57 ± 0.21 c | 5.82 ± 0.12 c | 6.29 ± 0.17 c |
| Bottom | ||||
| 4.1 A | 5.91 | 5.14 | 5.59 | 6.13 |
| 4.1 B | 5.80 | 5.20 | 5.36 | 6.01 |
| 4.1 C | 5.80 | 5.03 | 5.24 | 5.96 |
| Mean ± SEM | 5.84 ± 0.04 c | 5.12 ± 0.05 c | 5.40 ± 0.10 c | 6.03 ± 0.05 c |
| Total Mean ± SEM | 5.89 + 0.10 | 5.35 + 0.14 | 5.61 + 0.12 | 6.16 ± 0.10 |
a r/K class 1 = bacteria producing visible colonies at day 1
r/K class 2 = bacteria producing visible colonies at day 2
r/K class 3 = bacteria producing visible colonies at day 6
b Bacteria recovered in day 1 and 2 are fast growers (r-like strategists) and those recovered on day 6 are slow growers (K-like strategists).
c Log data were analyzed for statistical significance by using t-test (Graph Pad Prism version 5 software). Values ± SEM followed by the same letter are not significantly different (P >0.05) within each vertical column.
Percentages of r/K bacterial strategists (classes 1, 2 and 3) on total culturable bacteria colony counts as they appeared on 0.1 TSA over a period of six days and EPI-index.
| Soil samples | r/K class 1 | r/K class 2 | r/K class 3 | EPI-index |
|---|---|---|---|---|
| r-strategistsa | K-strategistsa | |||
| Top | ||||
| 2.2 A | 46% | 12% | 42% | 0.42 |
| 2.2 B | 51% | 23% | 26% | 0.45 |
| 2.2 C | 39% | 25% | 37% | 0.47 |
| Mean ± SEM | 45.33 ± 3.48b | 20.00 ± 4.04a | 35.00 ± 4.73a | 0.47 ± 0.01b |
| Bottom | ||||
| 4.1 A | 61% | 10% | 29% | 0.39 |
| 4.1 B | 62% | 15% | 23% | 0.40 |
| 4.1 C | 69% | 12% | 19% | 0.36 |
| Mean ± SEM | 64.00 ± 2.52b | 12.33 ± 1.45a | 23.67 ± 2.91a | 0.41 ± 0.02b |
a Bacteria recovered in day 1 and 2 are fast growers (r-like strategists) and those recovered on day 6 are slow growers (K-like strategists).
b Percentage data were Logit transformed and analyzed for statistical significance by using t-test (Graph Pad Prism version 5 software). Values ± SEM followed by the same letter are not significantly different (P >0.05) within each vertical column. Significant data are also indicated in bold.
Percentages of r/K bacterial strategists (classes 1, 2 and 3) on total culturable bacteria colony counts as they appeared on 0.1 TSA over a period of six days and EPI-index.
| Organism | Taxonomy | Plasmid | Soil sample | Sub total |
Total | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Top | Bottom | ||||||||||
| 2.2.A | 2.2.B | 2.2.C | 4.1.A | 4.1.B | 4.1.C | ||||||
| Isolate numbering | |||||||||||
| 1-200 | 201-400 | 401-600 | 601-800 | 801-1000 | 1001-1200 | ||||||
| Stenotrophomonas |
g - proteobacteria |
+ | 0 | 0 | 4 | 0 | 0 | 0 | 4 | 19 | |
| - | 5 | 7 | 1 | 2 | 0 | 0 | 15 | ||||
| Acinetobacter | + | 0 | 7 | 1 | 0 | 0 | 0 | 8 | 13 | ||
| - | 0 | 3 | 1 | 0 | 1 | 0 | 5 | ||||
| Enterobacter | + | 0 | 1 | 3 | 0 | 0 | 0 | 4 | 13 | ||
| - | 0 | 2 | 6 | 0 | 1 | 0 | 9 | ||||
| Pseudomonas | + | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | ||
| Pantoea | + | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 1 | ||
| Klebsiella | - | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | ||
| Agrobacterium |
a - proteobacteria |
- | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | |
| Bosea | - | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | ||
| Sinorhizobium | - | 0 | 0 | 0 | 0 | 1 | 0 | 1 | 1 | ||
| Paenibacillus | Gram+ low G+C | + | 0 | 0 | 0 | 6 | 1 | 0 | 7 | 16 | |
| - | 1 | 0 | 1 | 3 | 4 | 0 | 9 | ||||
| Staphylococcus |
Gram + low GC |
+ | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 2 | |
| Bacillus | - | 3 | 0 | 6 | 5 | 5 | 2 | 21 | 21 | ||
| Agromyces |
Gram+ High GC |
- | 0 | 0 | 0 | 1 | 0 | 0 | 1 | 1 | |
| Streptomyces | - | 1 | 0 | 0 | 0 | 1 | 0 | 2 | 2 | ||
| Rhodococcus | - | 1 | 0 | 0 | 1 | 0 | 0 | 2 | 2 | ||
| Arthrobacter | - | 0 | 0 | 0 | 0 | 0 | 2 | 2 | 2 | ||
| Microbacterium | - | 2 | 1 | 3 | 2 | 1 | 0 | 9 | 9 | ||
| Total Isolates | + | 0 | 9 | 9 | 8 | 1 | 0 | 27 | |||
| - | 14 | 14 | 18 | 15 | 14 | 4 | 79 | ||||
| + | 18 | 9 | 27 | ||||||||
| - | 46 | 33 | 79 | ||||||||
Symbols: +/- represent the presence/absence of plasmids, respectively.
MATERIALS AND METHODS
Soil Sampling and Processing
Bacteria analyzed in this work (Table 1) were isolated from six soil samples collected from the top and the bottom of a 13-years experimental field located in the coastal hills of Marche (Agugliano, Ancona, Central Italy). The six soil samples were referred to as 2.2 A, 2.2 B, 2.2 C and 4.1 A, 4.1 B and 4.1 C (from the top and the bottom of the experimental field, respectively). The Agugliano soil is a Calcaric Gleyic Cambisol with 20% slope [9], which in the first 30 cm has an Ap horizon (that is the homogeneous layer due to plowing) and a clay-silt texture [10]. The soil is managed under a Triticum durum (in winter) and Zea mays (in summer) rotation. Six soil samples were collected on 11 June 2007, during the maize rotation, in no-tillage (NT) system (sod seeding with chemical desiccation and chopping) and unfertilized (UF) soil (0 Kg N ha-1) at 0-20 cm depth. Each soil sample consisted of five soil cores taken inside two NT-UF blocks (top and bottom) of experimental field free from roots and then pooled together. Soil samples were sieved immediately at 2-mm mesh size, kept at 4°C and processed for further analysis within 24 h from sampling.
Culturing of Fast- and Slow-growing Culturable Bacteria
About 1-g (wet weight) of each soil sample was suspended in 10 ml of sterile phosphate-buffered saline (PBS, pH 7.3), homogenized at low speed 3 (Ultra-Turrax Thyristor Regle 50, Janke & Kunkel IKA-Labortechnik) and vortexed for 30 seconds. Then, each sample was transferred into a sterile 100-ml Erlenmeyer flask containing 10 g of glass beads (average diameter, 2 mm previously autoclaved for 20 min at 121°C) and shaken for 1 h at 120 rpm and 28°C to disperse bacteria. The flasks and glass beads were autoclaved for 20 min at 121°C before use. The resulting soil suspension was removed and transferred to a sterile 15-ml Falcon tube. Serial dilutions of this suspension were performed with sterile saline solution (9 g l-1NaCl) from 10-1up to 10-7. Then, 100 μl aliquots of serially diluted soil suspensions were plated in triplicate on 0.1 tryptic soy broth (TSB, Difco) containing 15 g l-1 agar (0.1 TSA) and 100 ìg ml-1 cycloheximide (Sigma) to inhibit fungal growth. Plates were incubated at 28 °C for 6 days. Total culturable bacteria were enumerated on the basis of the r/K strategy concept [11] at day 1, 2 and 6; in this way, three counts per plate were performed, corresponding to three classes (1, 2, and 3) with different growth rate. Bacteria producing visible colonies at days 1 and 2 (classes 1 and 2) were defined as “fast growers” (copiotrophs or r-strategists), while bacteria that produced colonies later (class 3) were defined as “slow growers” (oligotrophs or K-strategists). The number of bacteria in each class was expressed as a percentage of the total count and gave insight into the distribution of r- and K-strategists in each sample. Characteristics of r-strategists include fast growth in response to medium enrichment, while K-strategists are characterized by slow growth in response to enrichment.
After six days of growth at 28°C, a set of 1,200 colonies (200 per each of the six soil samples) were randomly chosen from 0.1 TSA plates and isolated on the same medium for the further characterization.
Eco-physiological Index
To express the distribution of the fast- versus slow-growing bacteria (r-versusK- strategists) in soil samples, the Eco-Physiological (EP) index [12], was calculated using three classes (i.e. colonies grown after 1, 2, and 6 days) [11]. The EP index of each soil tested was calculated using the equation: H’= -∑(Pi x log10 Pi), where Pi represents the CFU at each day (1, 2 and 6 days of incubation) as a proportion of the total CFU in that sample after 6 days incubation i.e. the proportion of colonies appearing on counting day i (i = 1, 2, 6) with EPmin = 0. Higher values of EP index imply a more even distribution of proportions of bacteria developing on different days (i.e., different classes of bacteria).
Statistics
Bacterial population data (CFU g-1 of soil) were log transformed and subsequently analysed by using t-test (Graph Pad Prism version 5 software). Percentage data of r/K strategists and EP index value were logit-transformed, Logit (p) = log [p/(1-1-p)] for the proportion p, and compared using t-test (Graph Pad Prism version 5 software).
Agarose Gel Electrophoresis
Agarose gel electrophoresis in TAE buffer (0.04 M Tris-Acetate, 0.01 M EDTA) containing 0.5 μg/ml (w/v) of ethidium bromide [13] was used to check the presence of plasmids (0.8% w/v), and to analyze amplicons obtained either from PCR amplification of 16S rRNA genes (0.8% w/v) or RAPD fingerprinting (2.0 % w/v).
Analysis of Plasmids Content
Analytical amounts of plasmid DNA were obtained from 1.5 ml bacterial cultures using the commercial Kit Plasmid Miniprep (Qiagen) set up for Gram-negative bacteria with the use of a robotic workstation (QiaCube, Qiagen).
PCR Amplification and Sequencing of 16S rRNA Genes
PCR amplification of 16S rRNA genes was carried out according to Papaleo et al. [14] using a MJ Research PTC 100 Peltier Thermal Cycler (CELBIO). Amplicons were excised from agarose gel and purified using the “QIAquick” gel extraction kit (QiAgen). Direct sequencing was performed on both DNA strands using the chemical dye terminator [15].
RAPD Analysis
Random amplification of DNA fragments was carried out using primer 1253 (5’ GTTTCCGCCC 3’) and the amplification conditions described elsewhere [16].
Homologs Retrieval and Phylogenetic Analysis
Probing of the DNA databases was performed with the BLAST program [17], using default parameters. The ClustalW program [18] was used to align the 16S rRNA gene sequences obtained with the most similar ones retrieved from databases. Each alignment was analyzed using the neighbor-joining method [19] according to the model of Kimura 2-parameter distances [20]. Phylogenetic trees were constructed using the MEGA4 software [21]. The robustness of the inferred trees was evaluated by 1000 bootstrap resamplings.
RESULTS AND DISCUSSION
Characterization of Bacterial Communities Isolated from Soil Samples
Six soil samples were collected from two boxes of UF-NT soil, i.e. from the top (2.2 A, 2.2 B, and 2.2 C samples) and from the bottom (4.1 A, 4.1 B, and 4.1 C samples) of the hillside. The total microbial population density ranged from Log10 6.01 ± 0.05 (bottom) to Log10 6.61 ± 0.17 (top) cfu g-1 of soil (Table 2), and no significant difference (P>0.05) was observed between the soil samplings collected from the top and the bottom of the experimental field. The structure of the bacterial soil community at each sub-sampling was investigated using the concept of r/K strategy [11] and reported in Table 3. Results indicated that bacterial colonies visible after one day of incubation (r/K class 1) were more abundant in the bottom soil compared to those in the top soil (P<0.05), whereas no differences (P>0.05) were observed between the bottom and the top samplings colonies visible after two days of incubation (r/K classes 2 and 3) (Table 3). Significant differences in EPI-index between the top and the bottom soil were also found (P<0.05) (Table 3), suggesting changes in community structure of cultivable bacterial communities in the two blocks of experimental field.
Analysis of Plasmid Content
Two hundreds bacterial isolates from each of the six samples (2.2 A, 2.2 B, 2.2 C, 4.1 A, 4.1 B, and 4.1. C) were randomly selected for further characterization. The presence of plasmids was checked as described in Materials and Methods on each of the 1,200 bacterial isolates randomly selected and re-grown on 0.1 TSA medium (Tables 1 and 4). Data obtained are shown in Fig. (2) and revealed that only 27 out of the 1,200 bacterial isolates harbored plasmid molecules. Most of the strains exhibited only a single plasmid molecule, while in a few cases (i.e. isolates 308bis and 363) multiple plasmids were found in the same cell. Furthermore, some isolates showed plasmids with the same electrophoretic mobility. The size of plasmid molecules ranged between about 2 Kb and 40 Kb, as determined by comparing their eletrophoretic mobility with that of reference plasmids. However, we cannot a priori exclude the possibility that the genome of some of the bacterial isolates analyzed might contain large and/or low copy number plasmids, which might have not been revealed by the extraction methodology used in this work.
RAPD Fingerprinting
In order to type the 27 bacterial isolates harboring plasmids, a RAPD [16] analysis using the primer 1253 was carried out. The comparative analysis of RAPD profiles obtained allowed the bacterial strains to be clustered in groups embedding bacterial isolates exhibiting the very same amplification profile (hereinafter haplotype). Bacterial isolates with the same haplotype were considered as the same strain. Data obtained are reported in Fig. (2), which shows that the 27 isolates can be split into 15 RAPD groups. Indeed, some isolates exhibited the same RAPD profile, suggesting that they might correspond to the same bacterial strain. In most cases, isolates exhibiting the same RAPD profile share a plasmid with the same electrophoretic mobility (i.e. the same plasmid if we assume that plasmids with the same electrophoretic mobility correspond to the same molecule) (see, for instance, isolates 230, 251, 253, 327 and 398 – RAPD haplotype 3). Just in one case, isolates sharing the same RAPD profile (i.e. cells of the same strain) harbored different plasmids (isolates 457, 500, 506 - RAPD haplotype 13), suggesting that the same strain may host different plasmids.
Phylogenetic Affiliation of Bacterial Isolates Harboring Plasmid Molecules
To affiliate each bacterial isolate to a given taxon, the nucleotide sequence of the 16S rRNA gene from at least one isolate per each RAPD group was determined. The 16S rRNA genes were PCR-amplified and sequenced from 25 isolates and their analysis revealed that:
i) The 15 RAPD haplotypes were representative of seven bacterial genera, two Gram positive (Staphylococcus and Paenibacillus) and five Gram negative (Acinetobacter, Enterobacter, Pantoea, Stenotrophomonas, and Klebsiella, all belonging to (-proteobacteria), with Acinetobacter and Paenibacillus representing half of the bacteria-harboring plasmids community (Fig. 2 and Table 4).
ii) The 16S rRNA gene sequences from three Enterobacter isolates (457, 500, 506) were identical in agreement with the finding that they also share the same RAPD profile. The fourth Enterobacter isolate (363), exhibiting a different RAPD profile (Fig. 2), also possesses a 16S rRNA gene sequence differing in one position in respect to the other three ones.
iii) The two Staphylococcus strains (706 and 708 exhibiting different RAPD haplotypes) shared the same 16S rRNA gene sequence, suggesting that they belong to the same species.
iv) The Acinetobacter sequences were placed in three distant branches of the trees, suggesting that they very likely belong to (at least) three different species, a finding that is in agreement with their very different RAPD profile.
Analysis of the Composition of the Aerobic Heterotrophic Cultivable Bacterial Communities Lacking Small Plasmids
In order to get some information also on the composition of the heterotrophic cultivable bacterial community, which did not exhibit plasmids under the experimental conditions used in this work and to compare it with the taxonomical position of bacteria harboring small plasmids, the 16S rRNA genes were amplified and sequenced as described in Materials and Methods from a panel of 79 randomly chosen bacterial isolates (Table 1). The analysis of the 79 nucleotide sequences obtained revealed that they were affiliated to 14 bacterial genera, with Bacillus (21 isolates) and Stenotrophomonas (15 isolates) being the most represented ones. Half of the isolates belong to Gram- bacteria represented only by members of the α- and γ-proteobacteria; the other isolates belong mainly to Bacillus and Paenibacillus, even though representatives of different genera (Agromyces, Arthrobacter, Microbacterium, Rhodococcus, and Streptomyces) of high GC Gram+ bacteria were disclosed.
Data reported in Table 4 revealed that bacteria belonging to Stenotrophomonas, Acinetobacter, Enterobacter and Paenibacillus included both isolates harboring or lacking plasmid molecules. To check the existence of a possible correlation between the presence of plasmids and the phylogenetic position of bacterial isolates, a phylogenetic tree for each of these four genera was constructed (Fig. 3), whose analysis revealed that some isolates embedded in the same genus very likely belong to different species, since the sequences joined different clusters of a phylogenetic tree. This is particularly true for Enterobacter and Acinetobacter isolates, whereas Stenotrophomonas and Paenibacillus exhibited a more homogeneous distribution within the respective tree.
CONCLUSION
In this work we have analyzed the aerobic soil heterotrophic cultivable bacterial community consisting of 1,200 bacterial isolates from NT and UF soil, with high organic input and increased enzyme activities, which favour functional diversity of the microbial community [10]. Total plate counts did not reveal any differences between the two sub-samplings (i.e. top and the bottom of the experimental field), while significant differences were found in the structure of the soil bacterial community that showed an uneven distribution of r- and K- strategists in the two sub-samples. Concerning the presence and the frequency of plasmids in these communities, data obtained suggest that they were harbored only by a low percentage (2.0 %) of bacterial isolates, that their size ranged between 2 and 40 kb, and that multiple plasmids were present only in a very limited number of isolates. The lack of plasmids of higher size did not per se imply their absence in the bacterial cells analyzed in this work; indeed, the procedure for plasmid extraction utilized in this work did not allow the isolation of plasmid molecules with a size higher than 40 kb.
We are completely aware that the size of the bacterial communities (1,200 isolates) that we have analyzed is much smaller of the extant total soil microbial communities and thus it represent a small sub-fraction of it. In spite of this limitation, the overall phylogenetic analysis revealed an uneven distribution of both species and plasmid-harboring strains. Interestingly, in most cases, isolates harboring plasmids of the same or different length clustered together in the same branch of the tree, which however also includes strains lacking plasmids. This might suggest that plasmids preferentially flow (vertically and/or horizontally) between (closely) related strains rather than between bacteria belonging to different species of the same genus. This is the case of Enterobacter, where at least three different strains of the same species show the same plasmid profile. Data reported in Table 4 also revealed that there was not a uniform distribution of plasmid-harboring isolates in the two samples collected from the top and the bottom of the experimental field. Indeed, Stenotrophomonas, Acinetobacter and Enterobacter isolates were detected only in the first sub-sample, whereas the Paenibacillus isolates were disclosed in the second one. Furthermore, some bacterial genera (such as Bacillus spp.) did not exhibit any plasmid molecule, even though they were “over”-represented in the bacterial community.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
This work was financially supported by the Italian Ministry for Education, University and Research through the National Project funded by National Programmes (FISR) SOILSINK: Climate change and agro-forestry systems, impacts on SOIL carbon sink and microbial diversity.


