All published articles of this journal are available on ScienceDirect.
Iron Acquisition Mechanisms: Promising Target Against Mycobacterium tuberculosis
Abstract
Continuous deployment of antitubercular drugs in treating Tuberculosis (TB) caused by Mycobacterium tuberculosis (MTB) has led to the emergence of drug resistance resulting in cross-resistance to many unrelated drugs, a phenomenon termed as Multi-Drug Resistance (MDR-TB). Despite reasonable documentation of major factors which contribute to MDR mechanisms, it appears unavoidable to consider novel mechanisms combating MDR. The ability of pathogenic MTB, to sense and become accustomed to changes in the host environment is essential for its survival and confers the basis of their success as dreadful pathogen. One such significant environmental factor that MTB must surmount is iron limitation, since they encounter diverse anatomical sites during the establishment of infection within the host. Considering the importance of MTB, being the second most common cause of mortality, this review focuses on gaining insights of iron acquisition mechanisms in MTB and how it can be exploited as efficient anti-mycobacterial drug target.
INTRODUCTION
Tuberculosis (TB) continues to pose significant global health challenges that require immediate treatment regimens directed at new targets. According to World Health Organization (WHO) one third of the world's population is infected with Mycobacterium tuberculosis (MTB), the causative agent of TB. After HIV/AIDS, TB is the second leading cause of death despite its uneven distribution around the world [1]. Widespread and continuous usage of anti–TB drugs have led to the development of multidrug resistance (MDR) to various frontline anti-TB drugs [2, 3]. To counteract the spread of drug resistant TB, there is an urgent need for either designing new anti-TB drugs and drug targets or devise novel strategies to combat MDR.
In recent years, emerging evidences has demonstrated that there are various novel mechanisms that can be targeted to combat the ever increasing burden of MDR in various human pathogen such as Candida albicans [4] and improved knowledge of such molecular mechanisms controlling MDR in pathogenic MTB may also facilitate the development of novel therapies to combat this deadly infection. Iron deprivation represents one of the crucial environmental stress conditions that MTB encounters during infection process due to non-availability of free iron in human host [5]. This review focuses on the iron acquisition mechanisms of MTB that can be employed as novel strategy to combat MDR.
SIGNIFICANCE OF IRON
Iron is a critical micronutrient required by almost all the organisms baring few exceptions like Borrelia burgdorferi especially as a cofactor in enzymatic system of many important metabolic functions [6]. Despite the fact that iron is the fourth most abundant element in the earth crust and second most abundant metal, it is not freely available neither to humans nor the invading pathogens. Being the transition metal it can exists in many oxidation states, however, in the living cell the most relevant oxidation states in which iron exists are Fe(II) and Fe(III). The variation in redox states makes it available to indulge in the generation of hydroxyl radicals via Fenton reaction which can cause deleterious effects on the living cell [7]. Thus, availability of free iron in host cells is tightly regulated making it less available to both the host and invading pathogen like MTB.
Role of iron has already been established in other major human pathogen, Candida albicans [8, 9]. MTB colonize a variety of anatomical sites which are likely to be scarce in iron, therefore it needs to exploit limited host iron reservoirs to make their survival favorable. Similarly, macrophages also need to protect iron from intracellular pathogens to gratify their own need as iron is needed as cofactors of many enzymes including those generating reactive oxygen or nitrogen species as immediate defense against MTB [7]. Thus, protecting iron from invading organisms is immensely important for macrophages and this competition between MTB and host macrophages for iron represents a critical factor which governs the elimination of invading pathogen.
AVAILABLE IRON SOURCES FOR MTB
MTB needs iron, to survive within hostile environment offered inside the host, therefore sequestration of iron represents a formidable challenge for MTB. For this MTB have adequate mechanisms to cope up with this essential nutritional deficiency. MTB has both non-heme and heme iron as the available resources to deal with iron limiting conditions (Fig. (1)). The following section discusses the various strategies employed by MTB to capture iron.
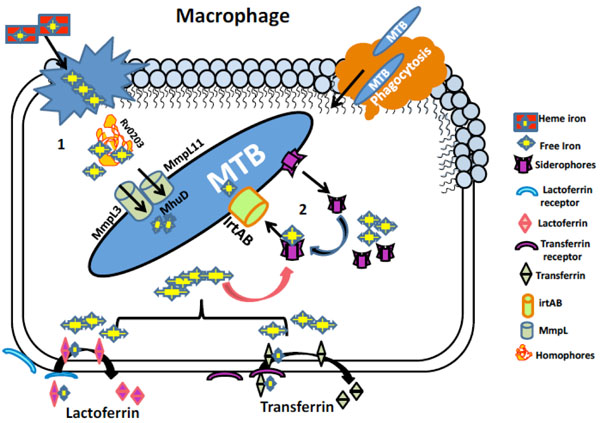
Iron acquisition mechanisms in MTB. (1) Iron uptake pathway utilizing heme source. Free heme from senescent RBCs engulfed by macrophages is bound to homophore Rv0203 and brings them to membrane proteins Mmpl3 and Mmpl11. These proteins transport heme to MTB cytoplasm where heme degrading protein MhuD degrades heme to release free iron. (2) Iron acquisition utilizing non-heme iron sources mediated by siderophores (Carboxymycobactin/ Mycobactin). Iron associated with host iron proteins i.e. transferrin and lactoferrin enters macrophage through their respective receptors where the siderophores synthesized by MTB chelates free iron and lactoferrin and transferrin are recycled. Siderophore bound iron is then transported to MTB cytoplasm via irtAB, an ABC family transporter protein.
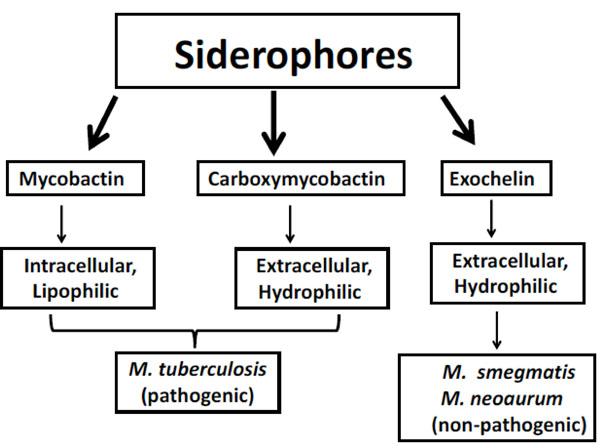
Outline of principle forms of siderophores synthesized by Mycobacteria.
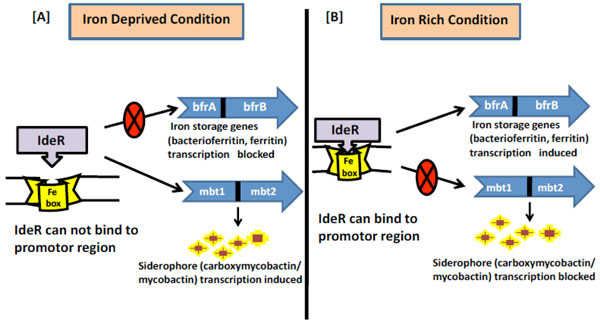
Regulation of iron metabolism in MTB (A). Under iron deprived condition the regulator IdeR cannot bind to the iron box in the promotor region as it requires iron for its activation. Thus the genes (iron storage) which are positively regulated by IdeR are not transcribed whereas the genes (siderophore) which are negatively regulated by IdeR are actively transcribed. (B). Under iron rich condition the regulator IdeR can bind to the iron box in the promotor region hence in active form. Thus the genes (iron storage) which are positively regulated by IdeR are actively transcribed whereas the genes (siderophore) which are negatively regulated by IdeR are not transcribed.
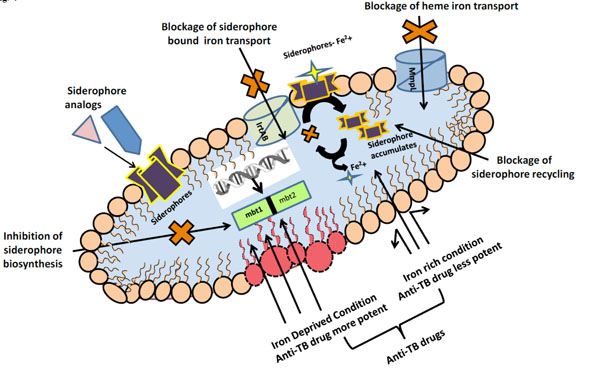
Promising Anti-TB drug targets exploiting iron acquisition mechanisms.
NON-HEME IRON
Transferrin & Ferritin
Pathogenic bacteria replicate inside the macrophage phagosome and needs iron for their growth and development. MTB fight for the available limited iron resources with host and one of the strategy employed is by extracting iron from iron storage molecules such as transferrin (TF)1, lactoferrin, and ferritin. Iron bonded with either transferrin or lactoferrin moves into the macrophages through binding with their respective receptors present on membranes. This is further internalized through clathrin-dependent endocytosis, also known as receptor - mediated endocytosis [10]. Within the endosome iron is discharged by reduction and acidification, after which iron gets transported to the cytoplasm through the divalent metal transporters, Dmt1 Nramp2 and DCT-1 [11].
Ferritin is another class of iron storage molecule available for MTB. In situ-Mossbauer spectroscopy revealed that siderophore-bound iron in intracellular pool is observed only upon short term labeling while long term labeling revealed a spectra coherent with bacterioferritin. Thus bacterioferritin like molecules are the principle iron storage molecule of MTB [12]. Two ORFs, bfrA (Rv1876) –a putative bacterioferritin and bfrB (Rv3841) – a ferritin like protein, have been identified in MTB genome. It is observed [13, 14] that the expressions of these iron storage molecules are induced under iron rich conditions and repressed under iron deprived conditions. Intriguingly, bfrA in MTB is under the influence of three promoters. Two of the promoters are repressed by high iron concentrations, while the third one is induced upon high iron concentrations [15, 16]. It is therefore, speculated that this mode of regulation allows maintenance of basal levels of bacterioferritin inside the pathogen so that any amount of excess iron can be immediately stored in a bound form.
HEME-IRON
Heme-iron contributes a substantial share of the human iron pool [17] and is a critical reservoir of iron for most microorganisms living inside human host. Refined heme uptake pathways have been found in both Gram-positive and Gram-negative microorganisms, which intercede host heme procurement [18-22]. Likewise, beside other pathways MTB also has a non-siderophore heme-uptake pathway which in comparison to non-heme pathway can draw large amount of iron which includes host hemoglobin [23]. Three MTB proteins have been proposed to be component of heme uptake pathway: the heme binding protein Rv0203, the transmembrane proteins MmpL3 and MmpL11and heme releasing protein MhuD.
Mmpl3 and MmpL11 are members of RND (resistance nodulation cell division) transmembrane transporter family involved in both heme uptake [24] and export of trehalose monomycolate. MTB encode gene for MmpL3 [23, 25] and is placed inside close genomic proximity of MmpL11, sharing 24% amino acid sequence identity and indicating a close functional relationship. Topologies of MmpL3 and MmpL11 are similar, consisting of 11 transmembrane domain and three soluble domains (E1, C1, and E2). It has already been shown that removal of MmpL11 experimentally causes abrogated heme uptake in MTB [23]. Notably, recombinant E1 and E2 domains from MmpL3 and MmpL11 were also shown to bind heme [23], suggesting that both MmpL3 and MmpL11 are transmembrane heme transporters. It has also been hypothesized that the extracellular heme-binding protein Rv0203 can directly transfer heme to MmpL3 and MmpL11 extracellular domains, thereby mirroring the mechanisms found in both Gram-positive and Gram-negative heme uptake systems [26, 27].
SIDEROPHORES AS FULCRUM OF IRON ACQUISITION IN MTB
Siderophores are low molecular weight compounds (generally 1 kDa) having high affinity for iron and synthesized by MTB in iron limiting conditions to chelate ferric iron found outside of MTB cell and then reinternalizing it through specific receptors. Mycobacteria synthesize different type of siderophore-like substances which are crucial for the endurance within macrophages [28, 29]. On the basis of presence or absence of a 2-hydroxyphenyloxazoline-ring system, they can be classified into two structural classes which consist of salicylate molecules known as mycobactins and carboxymycobactins [28]. On the basis of chain length in the alkyl substitution, these two types of siderophores differ in their polarity and solubility (Fig. 2). Alkyl chain of 2–9 carbons is found in carboxymycobactins whereas 10-21 carbons alkyl chain is found in mycobactins [30]. Carboxymycobactin directly competes with iron-binding molecules in the aqueous growth medium while the more lipophilic mycobactin is closely associated at the hydrophobic cell wall of MTB once iron is transferred from carboxymycobactins [31, 32].
Mycobactin and carboxymycobactin are synthesized in most extreme iron deprived conditions and are crucial for survival of MTB in vitro and in vivo [29]. It has been shown that synthesis of siderophore fails when Mbt-B gene was replaced with hygromycin-resistance cassette in a virulent strain of H37Rv and fail to survive in iron deprives condition [29]. Mbt-B gene is 46% similar to Ybt (HMWP2) gene responsible for the synthesis of yersiniabactin (siderophore) found in Yersinia pestis [33] and 48% similar to PchE, gene responsible in the synthesis of pyochelin found in Pseudomonas aeruginosa [34]. According to annotation of the MTB genome sequence, the assembly of mycobactin consist of cluster of 10 genes A to J encoding enzymes needed for its synthesis and coded by mbt1 [35]. When MTB faces low iron condition, IdeR regulator increases the gene expression for mycobactin biosynthesis i.e. genes mbtA-N. Genes MbA, MbtE and MbtF code for the peptide synthetases while genes MbtC and MbtD code for polyketide synthases, and Mbtl codes for an isochorismate synthase that gives salicylate triggered by MbtA. MbtB gene encodes an aryl carrier protein (ArCP) which is also the site for siderophore chain initiation. A progressive activity of MbtA (a salicyl-AMP ligase) and PptT (a phosphopantetheinyl transferase) appear in the MbtB (ArCp domain) activation as salicyl-S-ArCP. The other cluster encoded by mbt2 involves K-N four genes which also contain nonribosomal peptide synthetase (NRPS) and polyketide synthase (PKS) to help in assembling of mbtA-mbtf.
In case of non-pathogenic species of mycobacteria such as in M. smegmatis and M. neoaurum a peptidic siderophore is synthesized, known as exochelin. Structurally they are not related to carboxymycobactin, and are peptide-based hydrophilic siderophores. Carboxymycobactins and exochelins both are water soluble and are secreted to the extracellular environment [31]. Thus they are more suitably referred as dissolvable salicylate siderophore.
TRANSCRIPTIONAL REGULATION OF IRON HOMEOSTASIS
MTB needs to sense the iron excess or limiting conditions once inside the host cells to establish successful infection. This is achieved by tight regulation of gene expression such that iron dependent transcription machinery is turned on or off only when required to avoid any futile cycle. IdeR is a major regulatory protein which belongs to Diptheria Toxin repressor DxtR family of 230-amino acid protein and responsible for regulating iron homeostasis in MTB [36]. IdeR is active as a homodimer where each monomer consists of three functional domains having two metal-binding sites [37]. First two domains at N-terminal region are helix-turn-helix motif which includes the DNA binding domain, and the metal-binding domain. The third domain, found in the C-terminal region, folds like a SH3 (Src homology domain 3) domain, despite having no sequence homology to such domains [38]. The third domain combines with the first two domains and forms two ligand binding sites for metal [39], needed for stable dimer formation and full DNA binding activity. IdeR combines with the DNA as a double dimer with each dimer on opposite sides of the DNA [39]. Various metals can activate IdeR but iron (the natural cofactor) is the excellent metal for IdeR function because it is required in much lower concentration for dimerization and DNA binding in comparison to other metals [40]. The immediate impact of IdeR on the gene expression of iron sensitive genes associates with its capacity to bind to a 19bp transformed inverted-repeat of consensus sequence (TTAGGTTAGGCTAACCTAA). This sequence is available in the promoter region of IdeR regulated genes and most commonly referred as iron box [41, 42]. Interestingly, in response to iron availability, IdeR binds to the promoter region of siderophore and iron storage gene, however it influences their outflow in inverse ways Fig. (3). The iron box in the promoter region overlaps at -10 region alternately with the transcriptional start point (TSP) [40] of the genes that are repressed by IdeR, however binding of IdeR to tandem boxes located further upstream of TSP is needed for expression of bfrA and bfrB genes. [42]. Thus it has been hypothesized that IdeR binds at the –10 region blocking the access of RNA polymerase and represses transcription, but when IdeR binds between K35 to K100 region, it activates transcription by allowing the RNA polymerase access.
STRUGGLE FOR IRON AS KEY ATTRIBUTE FOR DRUG TARGETS
In clinical situations where disturbances of host, iron homeostasis are implicated in infectious disease pathogenesis, examining MTB iron acquisition mechanisms may allow better understanding of developing efficient therapeutics. The broad comprehensive detail on biosynthetic pathway and iron acquisition from different sources revealed that MTB require iron for their establishment of infection and in the pathogenicity. Hence it is pertinent to mention that iron uptake mechanism represents viable drug targets and there is a need to target crucial pathways of iron acquisition in MTB to stop MTB propagation Fig. (4).
MTB as already reviewed above synthesizes, the siderophores mycobactin and carboxymycobactin to empower productive iron utilization. One of the earlier anti-TB drug, p-aminosalicylic acid which is an analog of salicylic acid, is known to block mycobactin biosynthesis and subsequent release of iron inside MTB cell [14]. Moreover, interruption in the siderophore recycling causes the intracellular gathering of deferrated siderophores, which is harmful to MTB regardless of its capacity to utilize heme as an iron source [5]. This discovery recommends that focusing on siderophore recycling may prompt the advancement of required medications to compete world's most lethal bacterial pathogens. Another strategy that has been adopted is the use of synthetic analogs of siderophores which can act as their competitors and thereby preventing iron uptake in MTB. Scandium-siderophore has been used to compete with iron-siderophores and the same has shown promising results in Escherichiacoli and Klebsiella pneumonia [43]. Alternatively IrtAB, transporter of ABC family found in MTB can also be targeted which although does not takes part in the synthesis of siderophore but is mainly responsible in the transport of iron bound to carboxymycobactin [44]. Siderophore mediated drug delivery is another fascinating thought to be utilized as antibacterial drug. The concept of developing various siderophore conjugates such as conjugates with cephalosporin and penicillin have worked very actively against Gram negative bacteria [45]. Similarly, many research groups are in process to make synthetic siderophores for improved drug therapy system in MTB [46]. Targeting siderophore biosynthesis by employing bioinformatics tools has also emerged as a successful strategy. It has been reported that siderophore synthetic enzymes have been targeted by N-salicylsulfommyl adenosine (salicyl-AMS) to act as inhibitor on MbtA [44].
Similar to siderophore biosynthesis, even heme iron uptake pathway has also been used as drug target. It has been discovered that range of antibacterial compounds, including BM212 [47], SQ109 [48], and adamantyl urea-based compounds [49] were targeted on MmpL3. Although exploitation of heme pathway has not been studied and utilized in great extent so far [50], however, Rv0203, MmpL11 and MhuD which are MTB specific proteins and are involved in heme uptake pathway can be an excellent target for anti-TB drug.
Ascertaining the action of known anti-TB drugs in response to iron deprivation represents another important strategy that warrants immediate attention. Although limited research is done on this aspect, the approach is still in its infancy, given the fact that iron does have a role in enhancing the drug susceptibility of drugs like isoniazid [51], has provided positive hope. Recently, in-vitro effect of iron on growth and drug susceptibility of various drugs has been studied in MTB [52]. Transcriptional response and lipid composition has revealed that iron deprivation leads to differential gene expression of glycerolipid metabolism and isoprenoid quinone synthesis correlating with unique cell wall lipid profile of MTB [53]. Such studies will not be only helpful in revealing the specific iron dependent biomarkers in MTB that can be targeted but also provides insight on a global picture of MTB metabolome in response to iron deprivation.
CONCLUSION
Although, there exists only few reports, one can safely conclude that targeting MTB iron uptake could be an attractive therapeutic approach that needs to be exploited with deeper understanding. The significance of elucidating the iron regulation mechanisms in MTB will help to unravel how they are adapted to the mammalian host. Since iron availability is a crucial aspect, therefore, any iron depleting strategy that has an influence on changes in drug susceptibilities of MTB either alone or in combination with various anti-MTB drugs certainly merits a closer look.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
Financial assistances in the form of Young Scientist awards to Z.F from Science and Engineering Research Board (SERB), New Delhi (SR/FT/LS-173/2010) and to Z.F and S.H. from Board of Research in Nuclear Sciences (BRNS), Mumbai (2013/37B/45/BRNS/1903) is deeply acknowledged. We thank Prof. Rajendra Prasad, Director, Amity Institute of Biotechnology for encouragement.


