All published articles of this journal are available on ScienceDirect.
Characterization of Vibrio cholerae O139 of an Aquatic Isolate in Northern Vietnam
Abstract
During the cholera survey in Namdinh province (northern Vietnam) in July, 2010, one strain of Vibrio cholerae O139 was isolated from 7 environmental water samples positive for ctxA, toxR,VCO139 genes and named as V. cholerae O139, ND1 strain. This strain was lysogenic harbouring a genome similar to the filamentous phage fs1. The replicative form DNA of this phage (named as ND1-fs1, 6856 bp) was sequenced and compared with the other filamentous phages. The filamentous phage ND1-fs1 integrates into the region between ctxB and rtxA genes. The genetic organization of the CTXϕ of V. cholerae O139, strain ND1 was determined and the schematic representation of the genetic organization was shown together with the ND1-fs1 prophage.
INTRODUCTION
Vibrio cholerae O139 Bengal has caused large explosive outbreaks in India, Bangladesh and neighbouring countries since it first appeared in late 1992. In 1993, nine cases were reported from Bangkok, Thailand [1]. In spite of rapid spread of O139 in Southeast Asia, however, little information on cholera in Vietnam due to O139 is available in the national and international scientific literature.
Therefore, we conducted a survey on cholera due to V. cholerae O139 between 2006 and 2010 in Vietnam to shed light on the present situation of V. cholerae O139 in the aquatic environment in Vietnam. During the survey in Namdinh in 2010, we isolated one strain of V. cholerae O139 from an aquatic environment. Strains of V. cholerae O139 are subtyped according to the organization of CTXϕ [2].
The 6.9 kb CTXϕ genome contains genes of core and RS2 regions.The core region encodes ctxA,B and contains genes involved in phage morphogenesis. The RS2 region contains genes required for replication (rstA), integration (rstB) and regulation (rstR) of CTXϕ [3]. The CTXϕ is often flanked by the satellite phage RS1 that contains the rstC gene in addition to genes present in the RS2 element [4]. Based on the variations in the repressor gene rstR, CTXϕ has been classified into three major types, CTXClass ϕ (Classical type), CTXET ϕ (El Tor type), and CTXCalc ϕ (Calcutta type) [5]. In addition to the CTXϕ, another two types of filamentous phage group are reported in V. cholerae O139,i.e. fs1 group (fs1, VGJ ϕ, VSK, VEJ ϕ) and fs2 group (fs2, phage 493).
We report the characterization of V. cholerae O139, strain ND1, first isolated in Vietnam with special attention to the CTXϕ organization and the other filamentous phage integration.
MATERIALS AND METHODS
Sampling Sites
Water samples were collected in May 2010 at 14 sites in Namdinh province, including a deep well, a container for rain, and 1 kg of home made ice. The other 11 samples were collected from a large canal. The ice was left in a sterile plastic beaker to thaw, then transferred into a 1L-bottle. Each water sample was collected 500 ml in a sterile 1L glass bottle.
The sampling sites were located 120 km from the National Institute of Hygiene and Epidemiology (NIHE). The samples were sent to NIHE at room temperature by car as soon as collected.
Bacteriology
Each 450-ml water sample was mixed in a 1L flask with 50 ml of 10 X alkaline peptone water (Bactopeptone 10%, NaCl 10%; pH 8.5) used as an enrichment medium and incubated at 37°C for 18h. Thiosulfate-citrate bile salts-sucrose agar (TCBS; Eiken, Tokyo, Japan) was used as the selective medium for the isolation of V. cholerae. Presumptive identification was performed on the basis of standard biochemical reactions, including oxidase activity, Gram stain, growth in nutrient broth containing 0% NaCl, and acid production from glucose, inositol, mannitol, and sucrose. All suspected V. cholerae isolates were tested by agglutination tests with polyvalent O1 and O139 antisera. Ten ml of culture in alkaline peptone water was used for the extraction of genomic DNA.
DNA Extraction from Water Samples
Ten ml of alkaline peptone water (primary culture) was harvested by centrifugation at 5000g for 10 min at 4°C. The pellet was resuspended in 600 µl of TESS buffer (30 mM Tris-HCl, pH 7.5, 5 mM EDTA, 25% sucrose), then supplemented with 200 µl of lysozyme (5 mg/ml in 10 mM Tris HCl [pH 8.0]., 1 mM EDTA, 10 mM NaCl) and kept at 37 °C for 30 min.One hundred µl of proteinase K (10 mg/ml in autoclaved water)) was added to the suspension and further incubated for 30 min at 37°C, then 100 µl of 10% sodium dodecyl sulfate was added and mixed well. DNA was extracted by the phenol chloroform method and ethanol precipitation was performed as described by Maniatis et al., [6]. DNA was recovered using a toothpick, then washed with 70% ethanol once and with 100% ethanol once, dried by Micro Vac (TOMY,, Tokyo, Japan), then suspended with 200 µl of sterilized water and stored at -20°C until use.
DNA Extraction from a Pure Culture
In the case of pure culture, Vibrio cholerae O139 was grown overnight in 10 ml of LB (Luria-Bertani) broth [6]. Total cellular DNA was extracted and purified as described above.
Multiplex PCR
The primers used in this study are listed in Table 1. Amplification with the four primer pairs was performed simultaneously in 0.2 ml microcentrifuge tubes. Samples (1 µl) were added to the PCR mixture in a 30 µl volume of a final mixture of GoTaq Green Master Mix (Promega, Madison, WI, USA) containing 0.25 µM of each pair of primers for toxR, ctxA, O1, and O139. The amplification conditions were as follows: 5 min at 94°C for initial denaturation of DNA and 35 cycles, each consisting of 1 min at 94°C, 1 min at 55°C, 1 min at 72°C, with a final one cycle of extension for 7 min at 72°C in an iCycler (Bio-Rad, CA, USA).
List of Primer used in This Study
| Primers | Sequences (5` to 3`) | Amplicon Size (bp), Genes | Source or Reference |
|---|---|---|---|
| 101-F | CCT TCG ATC CCC TAA GCA ATA C | 779 (toxR) | [7] |
| 837-R | AGG GTT AGC AAC GAT GCG TAA G | ||
| CTX2 | CGG GCA GAT TCT AGA CCT CCT G | 564 (ctxA) | [8] |
| CTX3 | CGA TGA TCT TGG AGC ATT CCC AC | ||
| VCO1-F2-1, | GTT TCA CTG AAC AGA TGG G | 192 (VCO1-rfb) | [9] |
| VCO1-R2-2, | GGT CAT CTG TAA GTA CAA C | ||
| VCO139-F2, | AGC CTC TTT ATT ACG GGT GG | 449 (VCO139-rfb) | [9] |
| VCO139-R2 | GTC AAA CCC GAT CGT AAA GG | ||
| CORE-F | CTG TCT TTC CAC AGC GCC G | 1.5 kb (core) | [10] |
| CORE-R | CTT TAC TCT CGT GTT GCG GC | ||
| RS-F | CCG GTA CCA CTC ACC TTG TAT TCG | 900 (RS) | [11] |
| RS-R | CGG GTA CCT CGA CAT CAA ATG GCA TG | ||
| Zot-F | AAA CCT TGA ACG CAT AGC | 846(zot) | [11] |
| Zot-R | GCC CAT AGA CCA CGA TAA | ||
| RstRET-F | GCA CCA TGA TTT AAG ATG CTC | 320 (rstRET) | [12] |
| RstTRET-R | GGC AAT TAA GAC TCA GGC AC | ||
| RstRClass-F | TCT CAT CAG CAA AGC CTC CAT C | 243 (rstRClass) | [12] |
| RstTRClass-R | GTA GCA AAT GGT ATC GGC GTT GG | ||
| RstRCalc-F | GCT TTA TCT TAT GGA AGT CTT C | 160 (rstRCalcs) | [12] |
| RstTRCalc-R | GGC AAC AAA GCA CAT TAA AGA C | ||
| 3’zot-F | TCA CTG TTG GTG ATG AGC GT | This study | |
| ctxB-F2 | GCC GCA ATT AGT ATG GCA AA | [13] | |
| attP-L-R | CAA GAT AGC GAT AGC GGC TA | This study | |
| attP-R-F | CTT GAA CTT GTC GCC ACA GA | This study | |
| rtxA-Ri | GTT CAA GCG ATG AGT GGT TT | This study |
Purification of Replicative form DNA (RF DNA)
The RF DNA was extracted from a 10 ml culture in LB broth by using the QIAprep Spin Miniprep Kit (Qiagen).
Probes and Hybridization
Aliquots of DNA (4 µg) were digested with restriction enzymes AvaI,BglI, PstI, BglII and HindIII (NIPPON GENE, Tokyo, Japan), separated by electrophoresis in 1 % agarose gels and blotted on to nylon membrane (Hybond-N+; Amersham Pharmacia Biotec) by Southern blotting [14]. The gene probe used to detect ctxA was a 0.56 kb fragment amplified by PCR from V. cholerae A104, an El Tor strain isolated in Kenya in 1983; the CTXϕ core probe was 1.5 kb fragment amplified from the same strain; RS probe was a 0.9 kb amplified from the same strain; rstRET probe was a 320 bp PCR amplified product from the same strain, zot probe was a 0.84 kb fragment amplified from the same strain by PCR The probes used for hybridization were labeled with the DIG DNA Labeling and Detection Kit (Roche Molecular Biochemicals, Indianapolis, Ind). For Southern hybridization, the membranes were prehybridized at 42°C for 30 min in a solution containing 2x SSC (1x SSCis 0.15 M NaCl plus 0.015 sodium citrate). The membranes were then hybridized with the freshly denatured digoxigenin-labeled gene probes at 42°C for 15 h. The hybridized membranes were washed twice in 2x SSC-9.1% SDS for 5 min at room temperature and twice in 0.1x SSC-0.1% SDS for 15 min at 68°C. Non radioactive detection was based on digoxigenin-anti-digoxigenin enzyme–linked immunosorbent assay, according to the instructions for the DIG DNA Labeling and Detection Kit instructions. For typing of the filamentous phage, heat-denatured RF DNA of filamentous phage fs1 was used for Southern blotting after labeling with digoxigenin.
Nucleotide and Protein Sequence Analysis
Each PCR amplicon was gel purified by using QIAEX® (Qiagen). Nucleotide sequencing of all PCR products was carried out with the Taq Dye-terminator Sequencing Kit (Perkin-Elmer) and an ABI PRISM® 3100 Avant Genetic Analyzer (Applied Biosystems). The BLAST program was used to search for homologous DNA and protein sequences in databases.
Electron Microscopy (EM)
The Filamentous phage from a V. cholerae O139 was purified as reported before [15]. For negative staining, one drop of the phage solution was placed on a sheet of PARAFILM”M” (American Can Co., Greenwich, Conn., U.S.A.) and a Formvar-coated copper grid was floated on the drop for 2 min. The excess solution was removed with filter paper. The specimen was washed 3 times with distilled water, then stained with 1% uranyl acetate for 30 sec. The excess stain was removed by blotting with a filter paper. The specimen was examined with a JEOL JEM 1230 transmission electron microscope operated at 100 kV.
Nucleotide Sequence Accession Numbers
The nucleotide sequences of ND1-fs1, attR, attL regions and ctxB gene of V. cholerae O139, strain ND1 were deposited in the GenBank/EMBL/DDBJ database under accession numbers AB572858, AB646687, AB646688, and AB615200.
RESULTS AND DISCUSSION
Multiplex PCR
Seven of the 14 samples were positive for toxR, ctxA, VCO139 (Fig. 1).
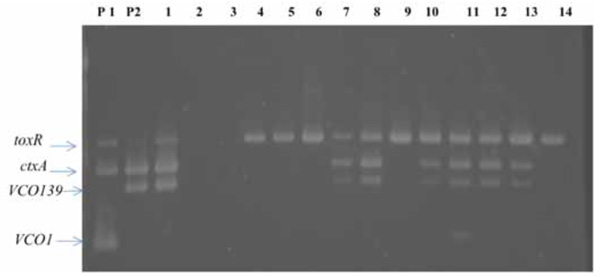
Multiplex PCR to detect O1, O139, ctxA, and toxR genes in environmental water samples in Nam Dinh province in June 2010. Lanes: P1, P2; positive control for VCO1 (strain HN1), VCO139 (strain AI4450) genes, 1; ice locally produced, 2; water from a deep well (150 m in depth), 3; water from a container for rain with a tight cover, 4-14; water samples from a canal.
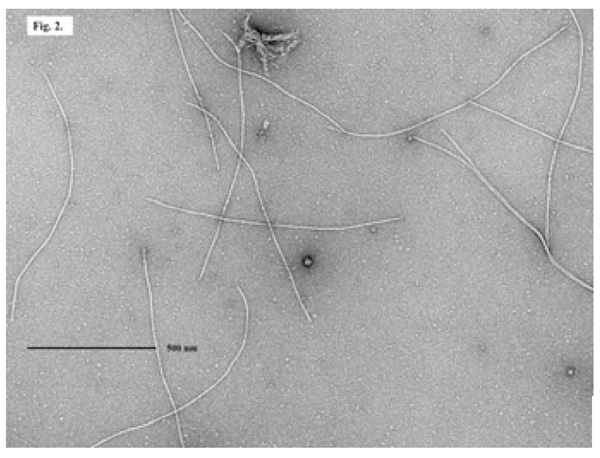
An EM picture of filamentous phage ND1-fs1 isolated from V. cholerae O139, strain ND1.
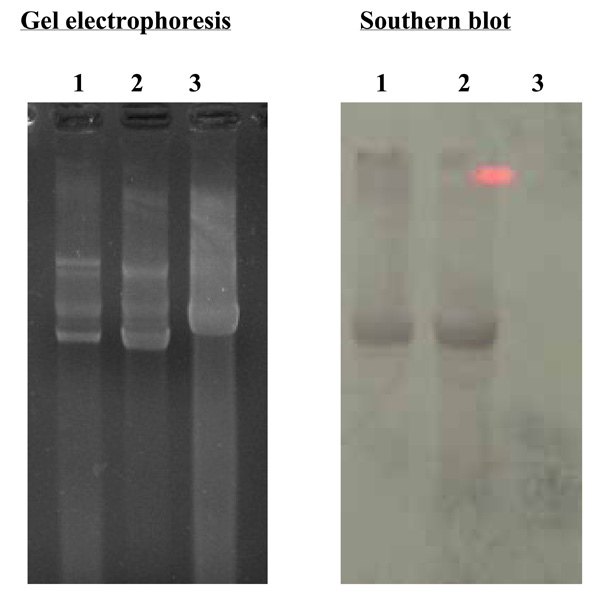
Southern blot analysis of RF-DNA from V. cholerae O139, strain ND1 probed with RF-DNA of fs1 labeled with digoxigenin. Lane1; RF-DNA extracted from V. cholerae O139, strain ND1, lames 2, 3; RF-DNA of filamentous phages, fs1 and fs2.
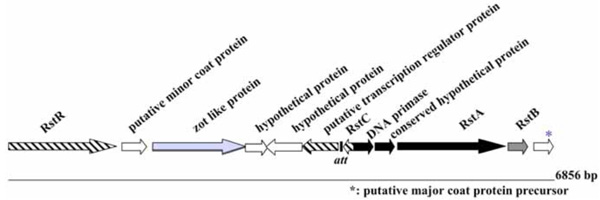
Modular structure of the genome of ND1-fs1. ORFs are shown with arrows pointing in the direction of transcription. The replication module is represented in black, the structural module in white, and the assembly and secretion module in light grey. The ORFs belonging to the regulation module are hatched. The integration module is represented in dark grey. Genome size is shown on the right. “att” recombination site, similar to XerCD, dif1 recombination site of V. cholerae is also shown.
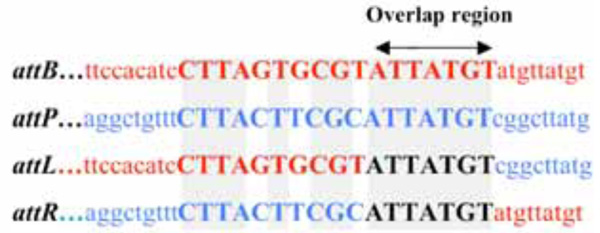
DNA region with attP, attB, attL, and attR sites as determined by sequencing. The core homologous sequences are indicated by uppercase letters, and identical nucleotides are indicated by a light grey background. DNA sequences of bacterial origin and DNA sequences of phage origin are indicated by red and blue letters, respectively. Black letters indicate sequences of unknown origin at the moment of recombination. The arrow indicates the putative overlap region, where the cutting and rejoining of the recombination partners took place.
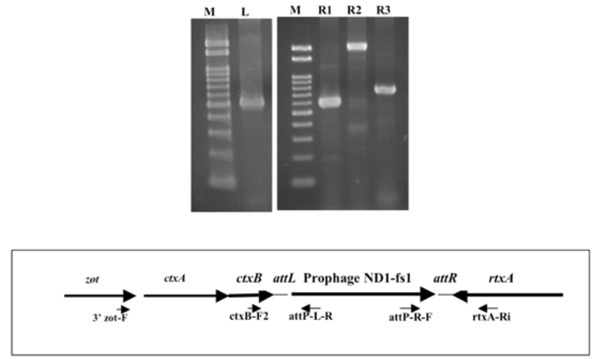
Determination of the left and right junctions of the ND1-fs1 by PCR. Lane M: marker DNA, 100 bp ladder, lanes L, R1, R2, R3: PCR products obtained using ctxB-F2/ attP-L-R (left junction), attP-R-F/ rtxA-Ri (right junction), 3’ zot-F/ rtxA-Ri, and ctxB-F2/ rtxA-Ri primers (ND1-fs1 is not integrated). Each primer site is shown in the lower box.
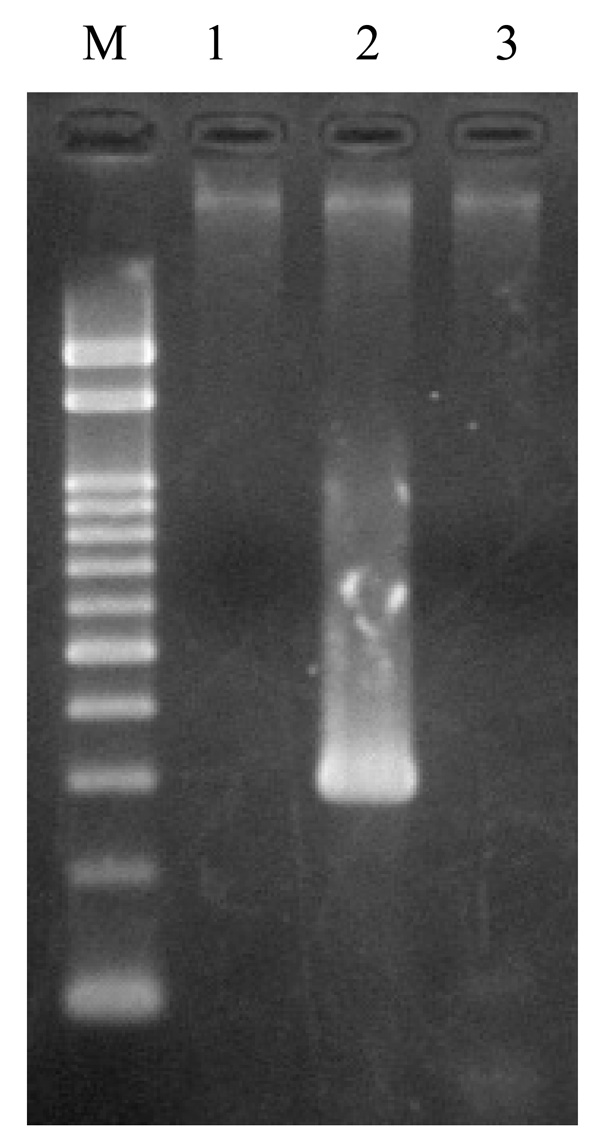
Detection of rstRCal, rstRET, and rstRClass by PCR in V. cholerae O139, strain ND1. Lane M: marker DNA, 100 bp DNA ladder, lanes1, 2, 3: rstRCal, rstRET, and rstRClass, respectively.
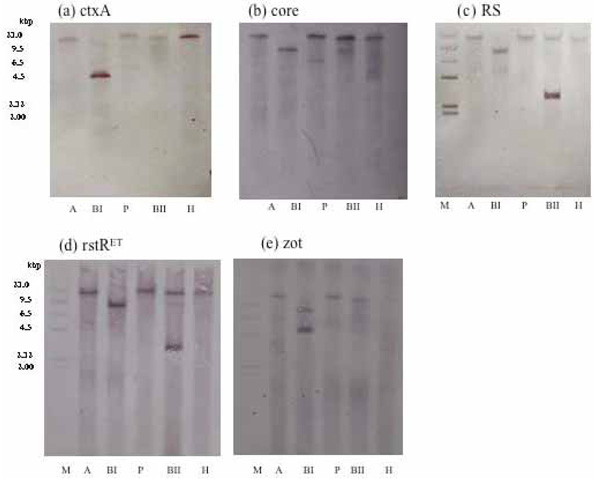
Results of Southern blot hybridization obtained with DNA from the ND1 strain of V. cholerae O139 digested with restriction enzymes, AvaI, BglI, PstI, BglII, and HindIII, and probed with (a) ctxA, (b) core, (c) RS, (d) rstRET, and (e) zot.

Schematic representation of genetic organization of CTXϕ of V. cholerae O139, strain ND1.
Surprisingly, a positive sample was found in the ice, which was produced locally under poor hygienic conditions, but the water from a deep well, used for making ice, was negative for VCO1, VCO139, toxR, ctxA. However, six of the environmental water samples were positive for VCO139, toxR, ctxA. This finding suggested that the ice was contaminated with environmental water, but the actual source of contamination remained unknown. The sampling sites were located along the same canal. The first sampling site is located about 10 km away from the last sampling site. When an environmental water sample is shown to be positive for ctxA, VCO139, the source of contamination is most likely from clinical cases. The clonality of environmental strains of V. cholerae O1 and O139 with ctxA gene is closely related with clinical strains [16]. Therefore, the finding of multiplex PCR suggested the presence of sporadic cases of cholera due to V. cholerae O139 in Namdinh province.
Isolation of V. cholerae O139
Out of seven samples found to be positive for VCO139, ctxA, toxR, we isolated V. cholerae O139 strain from only one sample (lane 1 in Fig. 1). This strain was named as V. cholerae O139, ND1. We don’t know exactly why only one sample culture was positive among 7 specimens positive for VCO139, ctxA, toxR by PCR But we experience great hardships in isolating V. cholerae O1 and O139 from the environmental water samples which show positive for VCO1 or VCO139 and ctxA by PCR. We speculate that the vibrio cell concentration may differ between 103-108 cfu/ml among the samples positive for ctxA,toxR, and VCO1 or VCO139. This strain was lysogenic and produced filamentous phages similar to fs1 [15] (Figs. 2 and 3). The phage was named ND1-fs1.
Genomic Organization of ND1-fs1
The ND1-fs1 DNA sequence is similar to sequences of other previously described phages, such as fs1 (96%) [15], VGJϕ (97%) [17], VSK (97%)[18], and VEJϕ (95%) [19]. The whole genome of ND1-fs1 consisted of 6856 nucleotides and had a G+C content of 42.9 mol%, which is slightly lower than the G+C content of its V. cholerae host (47.7% for chromosome I and 46.9% for chromosome II) [19]. The ND1-fs1 genome has 12 putative ORFs as shown in Fig. (4). It contained RstR, RstA, RstB, RstC, zot like protein, and minor and major coat proteins. This phage was integrated in an environmental V. cholerae O139. The role of this type of filamentous phage in pathogenesis remaines to be clarified. The other type of filamentous phage fs2 may play an important role in vivo. These two types of filamentous phages were found in fresh strains of V. cholerae O1 (manuscript in preparation).
Integration Site for ND1-fs1 in V. cholerae O139, Strain ND1
Since the ND1-fs1 phage is similar to fs1, VGJϕ, VSK, and VEJϕ, the integration site for ND1-fs1 was sought between ctxB and rtxA genes. The two amplicons, ca. 500 bp and 600 bp, produced by PCR with the two sets of primers ctxB-F2 / attP-L-R and attP-R-F / rtxA-Ri were exactly the left and right junction, respectively, as revealed by sequencing (Fig. 5). When amplified using the two sets of primers, 3’zot-F/ rtxA-Ri and ctxB-F2 / rtxA-Ri, two amplicons of 2 kb and 700 bp appeared (Fig. 6). The sequence data of these two amplicons showed that the two sequences were identical to those between zot and rtxA, and between ctxB and rtxA genes, respectively, without integration by ND1-fs1, even though DNA extracted from the lysogenic strain was used as a template. A similar result was reported from K139 phage [20], suggesting that the template DNA from any type of lysogenic strain is a mixture of two types of bacteria, lysogenic and non-lysogenic, probably due to spontaneous curring. The DNA regions with attP, attB, attL and attR sites are shown in Fig. (5). The attP sequence of ND1-fs1 was similar to that of VGJϕ, but the attB sequence of V. cholerae O139, strain ND1 was different from that of strain N16961 [17].
Genetic Organization of CTXϕ of V. cholerae 139, Strain ND1
Diverse CTX phages among Vibrio cholerae O139 were reported [21-26].In rstR allele specific PCR, this strain (ND1) produced an amplicon of 320 bp corresponding to El Tor type of rstR without amplicons of rstRCalc and rstRClass (Fig. 7). This prompted us to investigate the arrangement of CTX element in this strain. To determine the copy number and the arrangement of CTX genetic element, DNA from strain ND1 was digested with different restriction enzymes with recognition sites within the CTX element. Southern blot hybridization of chromosomal DNA digested with AvaI, BglI, PstI, BglII, and HindIII revealed a single band with ctxA probe (Fig. 8a), a single band with AvaI, BglI, BglII, HindIII, and two bands with PstI on hybridization with core probe (Fig. 8b). The hybridization patterns between RS and rstRET probes (Fig. 8c and d) were similar in the 2.7 kb band. The presence of 2.7 kb band in this strain suggests the presence of tandemly arranged RS1 element upstream of CTXϕ prophage. The higher intensity of 2.7 kb fragment revealed with the RS and rstRET probes (Fig. 8c and d, lane BII) suggested the presence of more than one copy of the rstRET gene. The restriction enzyme BglII has one site in rstRET The hybridization pattern with zot probe showed a single band with AvaI, PstI, BglII and two bands with BglI and HindIII. Based on the hybridization and PCR data, the organization of CTXϕ in V. cholerae O139, strain ND1 is presented in Fig. (9).
In conclusion, V. cholerae O139 seemed to spread widely in the environmental water sources in northern Vietnam. Our findings suggested the significance of testing diarrhea specimens with attention to V. cholerae O139. We found an environmental strain of V. cholerae O139 with an integrated filamentous phage. Filamentous phages of V. cholerae were reported after the appearance of V. cholerae O139 [15, 27]. We speculate some difference in maintaining the lysogenicity between V. cholerae O1 and O139. The strain of ND1 has an El Tor type of ctxB gene (AB15200) and a single copy of the ctx gene. Most strains isolated in India in 1993 had two copies of the ctx gene in tandem connected by two RSs [28]. Previously we reported the integration sites of vibriophage fs2 in a classical strain and in a strain of V. cholerae O139 [29]. It is noteworthy to examine whether two filamentous phages, fs1 and fs2 integrate into the chromosome of epidemic strains of V. cholerae O1 at the same time as shown in this study and the previous study [29] flanking the CTX phage.
CONFLICT OF INTEREST
The authors have no conflicts of interest to report.
ACKNOWLEDGMENT
We acknowledge the financial support received from the Ministry of Science and Technology, Vietnam. We are grateful to the Japan Society for the Promotion of Science (CORE UNIVERSITY PROGRAM) and Global COE Program of Ministry of Education, Culture, Sports, Science and Technology of Japan for supporting this study. This work was also supported by the Program of Japan Initiative for Global Research Network on Infectious Diseases(J-GRID, MEXT, Japan).


