All published articles of this journal are available on ScienceDirect.
Beyond the Virus: Exploring Coinfections in the COVID-19 Pandemic
Abstract
The global impact of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), responsible for the COVID-19 pandemic, cannot be understated. Amidst the relentless focus on this viral adversary, we must not lose sight of an equally formidable challenge – the lurking threat of bacterial coinfections that can exacerbate morbidity and mortality.
Various factors, including ICU admissions, age demographics, microbiota disturbances, and empirical antibiotic use, contribute to the specter of bacterial coinfections. Respiratory tract coinfections, often featuring Streptococcus pneumoniae, precede bacteremia and urinary tract involvement. However, the bacterial landscape in COVID-19 coinfections is a diverse tapestry with regional and institutional variations.
Unlike its viral counterparts, COVID-19 exhibits a lower incidence of bacterial coinfection, underscoring the urgency of judicious antibiotic administration to curb the looming threat of antimicrobial resistance. Pandemics have historically witnessed an upsurge in coinfection-related morbidity and mortality.
This comprehensive review delves into the multifaceted realm of bacterial, viral, and fungal coinfections amidst the COVID-19 pandemic. We scrutinize their impact on the respiratory and urinary tracts, blood, microbiota, and the ominous emergence of drug-resistant microorganisms. In conclusion, we explore nuanced treatment strategies in the quest for effective pandemic management.
1. INTRODUCTION
One of the current century's most severe pandemics was brought on by SARS-CoV-2 [1, 2]. The COVID-19, severe acute respiratory infections, and gastroenteritis are all brought on by this virus, which is a member of the Coronaviridae family, with a mortality rate of about 3-6% [3]. It has also been reported that this virus, similar to SARS-CoV and MERS-CoV, uses angiotensin-converting enzyme 2 (ACE2) as a receptor to invade the target cell [3]. The infection ultimately contributes to respiratory symptoms and lymphopenia, cytokine cascades, and immune reactions in the target tissue, leading to extreme respiratory symptoms and impaired immune functions [3-5]. The novel SARS-CoV-2 emerged from Wuhan, China, in December 2019 and is currently responsible for more than 544 million infections and 6.34 million deaths in 228 countries [6, 7]. ACE2 acts as a receptor for the SARS-CoV-2 virus in alveolar epithelial cells, and infection with this virus can contribute to acute respiratory distress syndrome [8]; furthermore, the SARS-CoV-2 RNA has been reported in human feces, indicating the presence of ACE2 receptors in the intestinal epithelial cells [9-11]. Studies have also shown that secondary bacterial infections, particularly Staphylococcus pneumoniae, was the primary cause of mortality from bacterial pneumonia in the influenza pandemic, occurred in 1918 [12, 13]. In general, secondary bacterial infections have been documented to be a complication of viral respiratory diseases that increase the severity of respiratory infections and pneumonia [14]. Due to the severity of the disease, 15% of patients are admitted to the intensive care unit (ICU), which provides an opportunity for bacterial infections [15]. If it occurs 24 hours after admission, they are considered community-acquired coinfections, but if they occur 48 hours after admission, they are called hospital-acquired superinfections [16].
Furthermore, researchers have found that common viral coinfections, such as influenza, rhinoviruses, and enteroviruses, were present in 17.2% of cases. Additionally, bacterial coinfections caused by both gram-positive and gram-negative species, like Mycoplasma pneumoniae, were identified in 7.11% of cases. The most commonly isolated bacterial species from ICU patients include Staphylococcus aureus, Legionella pneumophila, Haemophilus spp., Klebsiella spp., Pseudomonas aeruginosa, Chlamydia spp., Streptococcus pneumoniae, and Acinetobacter baumannii [17].
Furthermore, it has been shown that 1.3% of patients admitted to ICU due to drug resistance to S. aureus, Klebsiella pneumoniae, A. baumannii were susceptible to bacterial superinfection [8, 18-22]. Additionally, secondary bacterial infections such as S. pneumoniae, Haemophilus influenzae, and S. aureus, which are associated with influenza pandemics, have been illustrated as the most common causes of 11% to 35% bacterial coinfection [14]. It is worth noting that viral infections can promote complementary bacterial infections due to the host immune system's failure [23, 24]. Furthermore, the evidence shows that concomitant bacterial infection occurred in COVID-19 hospitalized patients. In these cases, an enhancement in the levels of pro-inflammatory cytokines and other biological markers associated with the disease indicates a secondary bacterial infection, which is attributable to the host immune system dysfunction [24-29]. Immune system disorders in COVID-19 patients can also assist coinfection occurrence with various opportunist bacteria [30].
Using mechanical ventilation as a supportive treatment for COVID-19 patients can increase the risk of hospital-acquired infections caused by bacteria likeEscherichia coli, K. pneumoniae, P. aeruginosa, A. baumannii, and S. aureus [31]. According to several studies, macrophage hyperactivity has been implicated as the cause of the low rate of bacterial concomitant infection in COVID-19 patients [16, 32]. Another study showed that due to a weakened immune system and reduced type I interferons (IFN), bacterial coinfection is common in male and female COVID-19 patients [33, 34]. It should be noted that mechanisms such as reducing mucociliary clearance, cell destruction by viral enzymes, the release of planktonic bacteria from biofilms, and agitation of dysbiosis in respiratory tract microbiome and gut microbiota are also involved in secondary infection with bacteria [10, 35]. Depending on the type of virus or bacterial species and the extent of the immune system response to the pathogen, the various molecular pathways contribute to each of the above modifications. In general, bacterial invasion in the airways occurs as a result of viral infections [36].
Furthermore, other upper respiratory tract viruses increase the susceptibility of immortalized epithelial cells to bacterial pathogens [38]. According to 80% of reported studies, the most common cause of bacterial coinfection is lymphopenia [36]. In this regard, coinfection can occur in the respiratory tract, the bloodstream, and the urinary tract [37]. Several bacterial infections have been reported concurrently with COVID-19. However, studies have suggested that the rate of improvement in COVID-19 patients with secondary bacterial infections is dramatically decreased, particularly when admitted to ICU [17]. In this study, we reviewed some medical articles about COVID-19 that reported bacterial coinfection.
2. BACTERIAL COINFECTION IN OTHER VIRAL RESPIRATORY TRACT INFECTIONS
In a report by Novotny et al., adenovirus and respiratory syncytial virus have been found to have enhanced intercellular adhesion molecule 1 (ICAM-1) expression in primary respiratory epithelial cells [38]. Also, owing to the propensity of Type IV pili (T4P) of non-typeable H. influenzae (NTHI) to the ICAM-1 receptor, this expression contributes to the sensitivity of cells expressing this receptor to bacterial pathogens [38]. Besides, the ability of P. aeruginosa to adhere to normal epithelial cells and cells affected by cystic fibrosis increases as a result of respiratory syncytial virus infection [35]. Moreover, dysregulation of pro-inflammatory cytokines has increased cell vulnerability to bacterial coinfections following viral infection. For example, IFNs stimulate the immune system and antiviral responses during viral infections; however, excessive secretion of IFNs can result in the destruction and damage to host cells [35, 39]. The production of anti-inflammatory cytokines, interleukins (IL), including IL-10 and IL-6 are induced by interferons, inhibiting the secretion of pro-inflammatory cytokines such as IL-17 and IL-23, which constitute innate and acquired immunity [35]. Additionally, macrophage, dendritic cells, natural killer cells’ activity and the percentage of CD4+ and CD8+ T-cells are decreased by IFNs, which interferes with the clearance of bacterial infections [40-43] (Fig. 1).
Despite extensive studies on viral and bacterial coinfections, the issue of infectious coronaviruses is debatable. On this basis, the human coronaviruses, including 229E, NL63, OC43, SARS-CoV-1, MERS-CoV, and SARS-CoV-2, have been reported to cause pneumonia in addition to influenza virus, as well as concurrent bacterial infections and extreme respiratory symptoms [44-46].
Human coronavirus NL63 (HCoV-NL63) has been reported to be one of the most significant viral pathogens in the upper and lower respiratory tract [47-49]. Besides, Golda et al. have demonstrated that HCoV-NL63-induced infection increases the susceptibility of virus-infected epithelial cells in the respiratory tract to bacterial pathogens such as S. Pneumoniae, which in turn exacerbates respiratory symptoms [48].
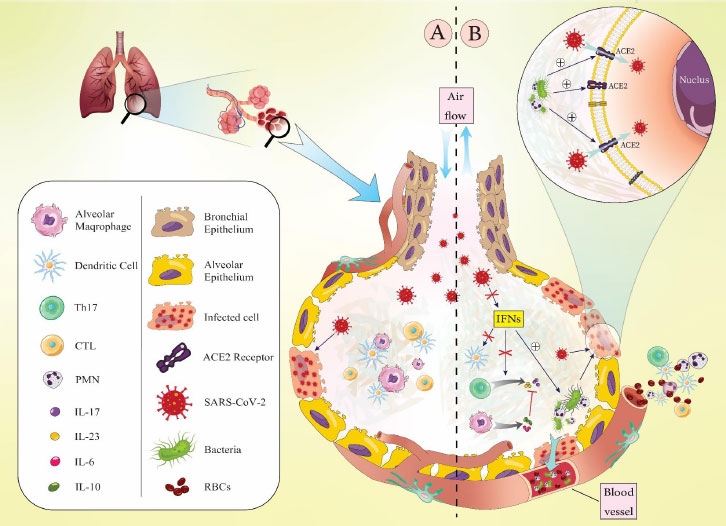
(A). Infection with SARS-CoV-2 alone can lead to an increase in the level of immune cells in the lung alveoli and subsequently cause inflammation through cytokine secretion. Also, during infection with SARS only clinical respiratory symptoms are mild.
(B) Coinfection of SARS-CoV-2 with respiratory pathogenic bacteria can enhance the migration of inflammatory cells (macrophages and neutrophils) and increase the secretion of pro-inflammatory cytokines such as IL-10, IL-17, and IL-23 in alveoli, which lead to an aggravation of the disease severity and tissue damage promotion in the alveoli. INFs increase the trapping of bacteria by PMNs, which promote inflammation. In this situation, bacteria have a greater chance of reaching the bloodstream through damaged cells and capillary vessels and causing bacteremia.
In another study on SARS, Zahariadis et al. indicated that a noticeable percentage of patients with SARS had bacterial coinfection with Chlamydophila Pneumoniae and Mycoplasma Pneumoniae [35]. Furthermore, Alfaraj et al. discovered MERS-CoV coinfection with tuberculosis in two cases [50], and Wang et al. illustrated seven patients with SARS-CoV-related deaths and secondary bacterial infection [51].
Following SARS-CoV infection, the accumulation of macrophages and mononuclear neutrophils increases in the body, which subsequently increases and decreases Th1 cells and NK cells, respectively [52, 53]. It also leads to the increase of cytokines IL-6, IL-1, IFN-γ, TGF-β, IL-12, IL-8, CXCL9, CXCL10 and CCL2 [52, 54, 55]. It is worth mentioning that in MERS-CoV infection, the cytokines IFN-β and IP-10 were significantly lower, while the level of IL-10 was increased. Also, CD4+ T cells, Th1 cells and Th2 cells were significantly decreased in MERS-CoV infection [56].
Based on the studies in SARS-CoV-2 infection, T regulatory cells (Treg) and CD4+ T cells decreased. Th1 cells were considerably reduced, while Th17 and Tfh cells were elevated. In the group with moderate symptoms compared to the seriously ill patients, there were more DCs, macrophages, CD4+ T cells, and TGF-+CD28 naive CD8+ T cells [57-59].
The body releases IL-10, IL-7, IL-2, MCP1, IP10, G-CSF, MIP1, and TNF-α in response to COVID-19.3 In contrast to SARS-CoV infection, SARS-CoV-2 infection also led to an increase in the release of anti-inflammatory Th2 cytokines such IL-4 and IL-10 [60, 61].
3. BACTERIAL COINFECTION IN COVID-19
Immunodeficient individuals have been documented as more vulnerable to COVID-19 and are more likely to develop secondary bacterial infections than other patients. The evidence suggests that hospitalization and consequent antibiotic resistance in some bacteria may cause secondary infections [25-27, 35].
About 50% of mortality during the COVID-19 pandemic is associated with secondary bacterial infections and often contribute to acute symptoms in COVID-19 patients [18, 62]. As mentioned, the ICU hospitalization of COVID-19 patients promotes coinfection frequency, where, due to antibiotic resistance, 1.3% of cases developed bacterial superinfection [17]. It has been illustrated that the lung cells destroyed during SARS-CoV-2 infection contribute to an enhancement in the propensity and colonization of bacteria [63].
Furthermore, the immune response rate in SARS-CoV-2 infection is different from the response rate in pneumonia due to concomitant bacterial-viral disease. Given the severity of clinical manifestations of COVID-19, which enhances coinfection, it facilitates the attachment and invasion of bacterial pathogens and eventually increases tissue damage [64]. Furthermore, airway dysfunction, cytopathology, and tissue disruption have been shown to occur during both single SARS-CoV-2 infection and bacterial coinfection [64]. This may promote the systemic dissemination of the virus, the pathogenicity of concomitant bacterial infections, blood infections, and sepsis [64]. Besides, epithelial cell disorders by respiratory pathogens have been documented to increase following rhino and influenza virus infections [65]. Moreover, it has been reported that structural and non-structural proteins of the SARS-CoV-2 inhibit the response of IFNs, which in turn predisposes the host to secondary bacterial infections [66, 67]. Proteins encoded by the SARS-CoV-2, including NSP1, ORF6, and N, inhibit IFN-related signaling pathways. It has also been reported that bacterial coinfections can disrupt host signaling pathways, increasing SARS-CoV-2 and exacerbating clinical symptoms. For instance, inhibition of NF-κB-associated signaling responses by K. pneumoniae leads to disruption of the host antiviral response [68]. Since the ACE2 receptor is an IFN-stimulated gene, the secreted IFNs during bacterial infections promote SARS-CoV-2 infection [69].
A study by Goncalves Mendes Neto et al. has reported that 57% of COVID-19 patients became coinfected with E. coli and E. cloacae, causing urinary tract infections (UTI). Furthermore, asymptomatic bacteriuria and lower UTI to acute pyelonephritis have been illustrated in these patients [70].
It has been shown that the intestinal-pulmonary axis plays a significant function in preventing bacterial pneumonia [71]. Also, in COVID-19 sufferers, intestinal cells are damaged and causing gastrointestinal symptoms and eventually stimulating the immune defense [72, 73]. In this regard, during severe infections, the intestinal microbiota can contribute to host susceptibility to secondary bacterial infections and vulnerability to concomitant infections [64].
Concurrent bacterial infections have been reported in other viral respiratory diseases, such as influenza H1N1 and H3N2, which cause problems in diagnosis and treatment and may eventually be associated with high morbidity and mortality [74]. Typically, after the virus spreads and induces infection, it disturbs the respiratory system both functionally and histologically [75]. These diseases range from mild to severe based on the type of virus infection; they include alterations in mucosal secretion, cell death, hyperplasia, reduced alveolar gas exchange, and compromised surfactant secretion [75]. The cited references are summarized in Table 1.
| Study Focus | References |
|---|---|
| Immunodeficiency and secondary bacterial infections | [25-27, 35] |
| Mortality associated with secondary bacterial infections | [18, 62] |
| ICU hospitalization and bacterial superinfection | [17] |
| Lung tissue damage and bacterial colonization | [63] |
| Immune response, coinfection, and tissue damage | [64] |
| Epithelial cell disorders in respiratory infections | [65] |
| SARS-CoV-2 proteins and IFN response | [66, 67] |
| Bacterial coinfections and disruption of host response | [68] |
| ACE2 receptor and IFN-stimulated gene | [69] |
| E. coli and E. cloacae coinfections in COVID-19 patients | [70] |
| Intestinal-pulmonary axis and bacterial pneumonia | [71] |
| Intestinal cell damage, gastrointestinal symptoms, and immune defense | [72, 73] |
| Concurrent bacterial infections in other viral respiratory diseases | [74] |
| Study Focus | References |
|---|---|
| Bronchiectasis and NTM infection as predisposing factors | [76] |
| Bacterial opportunity due to inflammation and lung tissue destruction | [77] |
| Mucosal/epithelial destruction and infection | [78] |
| Mechanical ventilation and coinfection risk | [79] |
| Bacteria responsible for ventilator-associated pneumonia | [80] |
| Bacterial coinfection in Wuhan COVID-19 patients | [81] |
| Comparison of community-acquired and hospital-acquired coinfections | [16, 82] |
| S. pneumoniae coinfection and vaccine recommendations | [83] |
| S. aureus and necrotizing pneumonia in COVID-19 | [84] |
| Diagnosis of bacterial pneumonia with BBAL | [85] |
| Antibiotics for gram-positive and gram-negative bacteria | [86] |
| Use of vancomycin in COVID-19 | [87] |
4. RESPIRATORY TRACT AND BACTERIAL COINFECTION IN COVID-19
Bronchiectasis and previous nontuberculous mycobacteria (NTM) infection predispose the affected individuals to concurrent bacterial and viral infections [76]. In COVID-19, bacteria get a chance to impair their host due to inflammatory reactions and the destruction of lung tissue [77]. Another factor predisposing to secondary bacterial infection is mucosal/epithelial destruction [78]. Mechanical ventilation is the most critical risk factor for the respiratory tract’s coinfection with S. aureus, P. aeruginosa, Klebsiella spp., Enterobacter spp., and E. coli in intensive care units [79]. On this basis, the average time of bacterial coinfection emergence after positive results of SARS-CoV-2 infection is six days [79]. Bacteria responsible for causing ventilator-associated pneumonia [80] consist of oropharyngeal microbiota in the first 4-5 days of hospitalization, while after five days, multidrug-resistant bacteria prevail [31]. The mortality rate increases by about 60% when these bacteria are resistant to multiple drugs, so to reduce the risk of ventilator-associated pneumonia in COVID-19 patients, the World Health Organization [6] recommends that instead of nasal intubation, oral intubation must be used with the patient’s head at an angle of 30°–45° [31].
A study from Wuhan, China, has shown bacterial coinfection with common respiratory bacteria such as Mycoplasma pneumoniae, Bordetella pertussis, and P. aeruginosa; among these, M. pneumoniae was sensitive to moxifloxacin [81]. A study from Spain showed that the rate of community-acquired coinfection is lower than that of hospital-acquired superinfections; P. aeruginosa was the most isolated species among hospital-acquired superinfections, which is consistent with the study reported from Italy [16, 82]. Another study that reported the highest coinfection rate with S. pneumoniae, suggested pneumococcal conjugate and polysaccharide vaccines to prevent bacterial pneumonia in COVID-19 patients [83].
Different bacteria have been reported from various studies in COVID-19 patients. For instance, S. aureus is one of the most reported bacteria from these patients that causes severe necrotizing pneumonia due to toxins such as Panton-Valentine leucocidin, prompting the prescription of antibiotics with anti-toxin properties, such as linezolid or clindamycin [84].
Blind bronchoalveolar lavage (BBAL) is the best sample for diagnosing bacterial pneumonia in critically ill COVID-19 patients as it reduces the risk of contamination and does not require bronchoscopy [85].
Although antibiotics active against gram-positive bacteria effectively treat secondary bacterial pneumonia in influenza patients, the antibiotics active against gram-negative bacteria are more useful for parainfluenza virus and coronavirus infections [86]. It is worth noting that vancomycin is not approved to treat bacterial pneumonia in COVID-19 patients [87]. The cited references are summarized in Table 2.
5. BACTEREMIA IN COVID-19
Coagulase-negative Staphylococcus spp., Corynebacterium spp., Bacillus spp., and Micrococcus spp. are usually isolated from blood cultures of COVID-19 patients; still, it should be noted that these isolated bacteria may be the normal flora of the skin [88]. However, in infected cases, bacteremia is caused by E. coli, S. aureus, K. pneumoniae, and Enterobacter cloacae [88]. Due to some diseases' endemicity, COVID-19 and feverish bacteremia coinfection in some areas may cause problems in diagnosis; in this regard, coinfection with brucella and COVID-19 reported by Saudi Arabian investigators has been successfully treated with the combination of doxycycline and rifampin [89].
Furthermore, some diagnostic factors, such as procalcitonin level among COVID-19 patients are low and indicate that bacterial coinfection is rare in these patients [88]; however, this factor alone is insufficient for diagnosis [90]. In general, bacterial infection has higher white blood cell (WBC) and neutrophil counts and a more elevated level of C-reactive protein than SARS-CoV-2 infection [91, 92]. Bacteremia in other viral infections has been reported to be very low, although most of the lethal cases were positive for bacteremia [88].
6. VIRAL COINFECTION IN COVID-19 PATIENTS
Researches show that COVID-19 coinfections are common in patients infected with blood-borne viruses such as human immunodeficiency virus (HIV) or hepatitis C virus (HCV), with respiratory viruses having the lowest rate of coinfection. More research is needed to determine the incidence rate of viral coinfection in COVID-19 patients due to the prevalence of influenza virus or RSV in society. Another significant feature that should be studied is their association with the patient's morbidity and death. In summary, more research into viral coinfection with SARS-CoV-2 is critical [93].
Patients with viral infections such as influenza, severe acute respiratory syndrome (SARS) in 2002, and Middle East respiratory syndrome (MERS) in 2012 have also been reported to be susceptible to concomitant bacterial infections; for example, in 2009 pandemic caused by the H1N1 influenza virus, 30-55% of the mortality was attributed to bacterial pneumonia [37]. Moreover, researchers have discovered that common viral coinfections such as influenza, rhinoviruses, and enteroviruses were involved in 17.2% of cases, and gram-positive and gram-negative species causing bacterial coinfections like M. pneumoniae, have been identified in 7.11% of cases [94].
The highly variable character of SARS-CoV-2 itself, as well as a lack of knowledge about host-pathogen interactions, made it difficult to develop efficacious remedies for the sickness. Viral coinfection in COVID-19 patients may complicate the patients' recovery from the illness; therefore, the interaction of various viruses with SARS-CoV-2, as well as their synergistic influence on illness clinical symptoms, should be studied [93]. In Wuhan, 5.8 percent of confirmed COVID-19 patients were infected with different kinds of respiratory viruses as well. COVID-19 coinfections can be brought on by the majority of respiratory viruses, including respiratory syncytial virus, human metapneumovirus, and rhinovirus, and having information about these coinfections can be useful in applying antiviral therapy. Patients with co-infections may respond differently to therapy than those with solely COVID-19 infection. It is also thought that the interaction of COVID-19 with respiratory viruses may exacerbate the illness severity, which is critical for vulnerable individuals, such as immunodeficient or immunosuppressed patients [93].
When compared to single infections, coinfections may result in changes in pathogen transmission, development of clinical symptoms, and the unfavorable consequences associated with any particular infection, which ultimately impacts the management of infectious illnesses. In the lack of appropriate data, the failure of standard approaches to detect coinfection might lead to underdiagnosis of coinfections [94, 95]. The cited references are summarized in Table 3.
7. FUNGAL COINFECTION IN COVID-19
Despite having a significant influence on human morbidity and mortality, fungi are still underappreciated for their negative effects on human health. The majority of fungi are naturally members of the human microbiome; however, some are opportunistic pathogens because they may cause serious illness in immunocompromised hosts. In healthy individuals, the opportunistic pathogens cause superficial, mild, and localized illnesses. However, immune-suppressive diseases such as diabetes and lung disease increase mortality and cause systemic morbidity [96].
A noticeable percentage of COVID-19 patients become severely sick and require ICU hospitalization, and these patients are more likely to develop fungal infections [97]. Severe COVID-19 is linked to immunological dysregulation, impacting both T-helper cell 2 (Th2) and Th1 responses, including the cytokine release syndrome, which promotes pulmonary microbial growth and infection [98]. Severe patients with COVID-19 have enhanced levels of pro-inflammatory (IL-1, IL-2, IL-6) and anti-inflammatory (IL-4, IL-10) cytokines. The likelihood of developing major fungi infections is increased by the mentioned clinical state [99].
According to research, SARS-CoV and SARS-CoV-2 are members of the same species and have similar biology and clinical features. Moreover, literature shows that the prevalence rate of fungal infection in SARS patients was 14.8-27 percent, which was the leading cause of mortality in SARS patients. However, fungal infection swab test was untended at the onset of the SARS-CoV-2 pandemic [100]. Fungal coinfection can aggravate the COVID-19 patients’ situation [101]. Some of the fungal pathogens observed in severe COVID-19 patients are Aspergillus, Candida, Mucor and Cryptococcus [100].
7.1. Invasive Pulmonary Aspergillosis (IPA)
IPA is a well-known consequence in immunocompromised patients, yet 50% of the incidences occur in individuals who are frequently non-neutropenic when brought to the ICU. Severe influenza is a recognized risk factor for developing IPA in these people. The disturbance of the respiratory epithelium, as well as defective mucociliary clearance and local immunological impairment, are important pathophysiological aspects in the development of IPA [99].
First-line therapy options for IPA include voriconazole, posaconazole, and isavuconazole. Echinocandins or nebulized amphotericin B in conjunction with anti-mold azoles are potential treatment options [99]
7.2. Invasive Candidiasis
The risk of infection with Candida species may increase significantly in severe COVID-19 patients who have more opportunities to be treated with broad-spectrum antibacterial drugs or in patients with immune impairment factors [100]. Critically ill COVID-19 patients are exposed to additional fungal infections such as Candida species and Pneumocystis jirovecii. All of the risk factors for developing candidemia in a critically ill patient are present in COVID-19 patients admitted to the ICU, including mechanical ventilation, parenteral nutrition, broad-spectrum anti-bacterial treatment, older age, comorbidities, lymphopenia, corti-costeroids, and so on [99].
7.3. Invasive Mucormycosis
Mucormycosis is more common in COVID-19 individuals who have had trauma, diabetes, glucocorticoids usage, hemopoietic malignancy, persistent neutropenia, allogeneic hematopoietic stem cell transplantation (allo-HSCT), or solid organ transplantation (SOT). It recommends thorough surgical treatment of mucormycosis as soon as feasible, in addition to systemic antifungal therapy; amphotericin B lipid complex, liposomal Amphotericin B, and posaconazole oral suspension are used as first-line antifungal monotherapy, whereas isavuconazole is strongly recommended as salvage treatment [100]. Due to the nonspecific nature of the pulmonary and disseminated mucormycosis clinical signs, which may overlap with those of COVID-19, the diagnosis of COVID-19-associated mucormycosis is challenging. In addition, a number of Mucorales species, the most common of which is Rhizopus arrhizus, are responsible for mucormycosis, some of which are poorly sensitive to antifungal treatment [101].
7.4. Invasive Cryptococcosis
COVID-19 Cryptococcosis, which mostly manifests as meningoencephalitis, affects people with human immunodeficiency virus (HIV) infection and CD4 T lymphocyte count less than 200 [100].
Additionally, post-influenza coinfection with Cryptococcus has been linked to decreased IFN- γ levels. As a result, IFN- γ treatment has been shown to be helpful as an adjuvant therapy in patients with chronic granulomatous illness who are undergoing transplantation; however, its favorable effects during fungal coinfections are yet unknown [102]. The cited references are summarized in Table 4.
The complete list of microorganisms causing coinfection with SARS-CoV-2 is summarized in Table 5.
8. THERAPEUTIC APPROACH
Given the SARS-CoV-2 outbreak and the use of antibiotics in patients admitted to the ICU, the spread of multi drug resistant (MDR) bacterial strains has been documented in healthcare systems. Therefore, to avoid the development and increase of MDR strain, antibiotic therapy of patients with COVID-19 should be interrupted in the case of mild bacterial infection and is recommended only in patients with severe respiratory symptoms [103, 104].
| Coinfection | Examples | References |
|---|---|---|
| Bacteria | Staphylococcus pneumoniae | [17] |
| Haemophilus influenzae | [14] | |
| Staphylococcus aureus | [31] | |
| Escherichia coli | [31] | |
| Klebsiella pneumoniae | [31] | |
| Pseudomonas aeruginosa | [31] | |
| Acinetobacter baumannii | [31] | |
| Chlamydophila Pneumoniae | [35] | |
| Pseudomonas aeruginosa | [79] | |
| Mycoplasma Pneumoniae | [35] | |
| Bordetella pertussis | [81] | |
| Enterobacter spp. | [79] | |
| Enterobacter cloacae | [88] | |
| Virus | Human rhinovirus | [95] |
| Human metapneumovirus | [95] | |
| Respiratory Syncytial Virus (RSV) | [95] | |
| Parainfluenza viruses | [95] | |
| Influenza virus | [95] | |
| Adenovirus | [95] | |
| Human immunodeficiency virus | [95] | |
| Hepatitis B virus | [95] | |
| Dengue viru | [95] | |
| Cytomegalovirus | [95] | |
| Fungi | Aspergillosis fumigatus | [100, 102, 107] |
| Candida albicans | [100, 102, 108] | |
| Candida glabrata | [109] | |
| Rhizopus oryzae | [100, 102, 110] | |
| Mucor | [110] | |
| Cryptococcus neoformans | [111] | |
| Cryptococcus gattii | [111] |
Additionally, research indicates a very large percentage of patients with SARS-CoV-2 have been treated with a wide variety of antibiotics, such as third-generation cephalosporins, quinolones, and carbapenems. In this respect, it has a significant role in considering the local epidemiology of drug resistance, its influence on the patient, and the assessment of antibiotic side effects such as diarrhea. Treatment with a wide variety of antibiotics, mainly acquired nosocomial infections, has also been reported to have a limited impact due to bacteria’s resistance to at least one class of antibiotics. Besides, a lack of adequate care for sepsis can contribute to promoted mortality [105, 106].
Since steroid levels are elevated in concomitant bacterial infections and given the anti-inflammatory activity of glucocorticoids, steroids prevent the cytokine storm's progression and subsequently control the host immune system. It is worth mentioning that in patients suspected of bacterial coinfection, oral antibiotics have fewer side effects than intravenous antibiotics [91].
CONCLUSION
According to present review, coinfections have a significant role during SARS-CoV-2 infection. For example, chronic obstructive pulmonary disease (COPD) is one of the chronic conditions associated with this sickness; these patients may be colonized by bacterial pathogens during the stable phase of the illness, thereby resulting in the host’s susceptibility to SARS-CoV-2 infection. In general, there are three different forms of SARS-CoV-2 bacterial infections, including: 1) SARS-CoV-2 infection secondary to bacterial infection, 2) coinfection and bacterial-viral pneumonia, and 3) secondary bacterial “superinfection” after contamination with SARS-CoV-2.
Consequently, the identification of bacterial coinfections during COVID-19 is critical. Besides, acute respiratory distress syndrome (ARDS) is a clinical feature of COVID-19 that increases nosocomial pneumonia risk. Moreover, other viral and fungal coinfections have an impact on the disease symptoms, severity and mortality. It has been shown that a wide range of pathogens, such as their antimicrobial resistance profiles and MDR bacteria, can be quickly identified using the Next-Generation Sequencing (NGS) metagenomic method. Also, the recognition of molecular pathway disorders due to simultaneous bacterial and SARS-CoV-2 infection leads to the development of new effective drugs and therapeutic interventions that provide appropriate immunity during simultaneous bacterial and viral coinfections.
In general, compared to other viral diseases, the rate of concurrent bacterial infection in COVID-19 disease is meagre, revealing the importance of antibiotic stewardship to control the emergence of drug resistance during the pandemic. On this basis, it leads to the facilitation of hospitalization procedures and treatment.
LIST OF ABBREVIATIONS
| SARS-CoV-2 | = Severe Acute Respiratory Syndrome Coronavirus 2 |
| ACE2 | = Angiotensin-converting Enzyme 2 |
| HCoV-NL63 | = Human Coronavirus NL63 |
AUTHORS’ CONTRIBUTION
HBB conceived the idea for this manuscript, edited subsequent drafts; RR: Literature search, Manuscript preparation, Design of the figure; ESM: Literature search, Design of the table; PSA: Literature search; HF, JSN and TE: Manuscript preparation.
All authors have read and approved the final manuscript.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors wish to thank the professionals of the Infectious and Tropical Diseases Research Center, Tabriz University of Medical Sciences, Tabriz, Iran.


