All published articles of this journal are available on ScienceDirect.
A Simple Method for Assessment of MDR Bacteria for Over-Expressed Efflux Pumps
Abstract
It is known that bacteria showing a multi-drug resistance phenotype use several mechanisms to overcome the action of antibiotics. As a result, this phenotype can be a result of several mechanisms or a combination of thereof. The main mechanisms of antibiotic resistance are: mutations in target genes (such as DNA gyrase and topoisomerase IV); over-expression of efflux pumps; changes in the cell envelope; down regulation of membrane porins, and modified lipopolysaccharide component of the outer cell membrane (in the case of Gram-negative bacteria). In addition, adaptation to the environment, such as quorum sensing and biofilm formation can also contribute to bacterial persistence. Due to the rapid emergence and spread of bacterial isolates showing resistance to several classes of antibiotics, methods that can rapidly and efficiently identify isolates whose resistance is due to active efflux have been developed. However, there is still a need for faster and more accurate methodologies. Conventional methods that evaluate bacterial efflux pump activity in liquid systems are available. However, these methods usually use common efflux pump substrates, such as ethidium bromide or radioactive antibiotics and therefore, require specialized instrumentation, which is not available in all laboratories.
In this review, we will report the results obtained with the Ethidium Bromide-agar Cartwheel method. This is an easy, instrument-free, agar based method that has been modified to afford the simultaneous evaluation of as many as twelve bacterial strains. Due to its simplicity it can be applied to large collections of bacteria to rapidly screen for multi-drug resistant isolates that show an over-expression of their efflux systems. The principle of the method is simple and relies on the ability of the bacteria to expel a fluorescent molecule that is substrate for most efflux pumps, ethidium bromide. In this approach, the higher the concentration of ethidium bromide required to produce fluorescence of the bacterial mass, the greater the efflux capacity of the bacterial cells. We have tested and applied this method to a large number of Gram-positive and Gram-negative bacteria to detect efflux activity among these multi-drug resistant isolates. The presumptive efflux activity detected by the Ethidium Bromide-agar Cartwheel method was subsequently confirmed by the determination of the minimum inhibitory concentration for several antibiotics in the presence and absence of known efflux pump inhibitors.
EXISTING METHODS TO ASSESS BACTERIAL EFFLUX ACTIVITY
The increasing multi-drug resistance reported in bacterial clinical isolates has become a major concern in the public health area given that the lack of treatment options for these infections, with the existing antibiotics, is becoming more and more problematic [1-3]. As multi-drug resistance can be the result of an over-expression of the bacterial efflux pump systems, used to extrude unrelated antibiotics prior to their reaching the intended targets [3], there is a need to develop and implement new and improved methods for a real-time and quick identification of efflux mediated multi-drug resistant (MDR) phenotypes [4, 5]. Primarily, assessment of efflux activity has been conducted using a common substrate of efflux pumps, such as ethidium bromide (EtBr). This compound is widely used to “monitor” efflux in a given bacteria and its increased retention after an uncoupler of the proton motive force or other efflux inhibitor has been added [4-6]. The baseline fluorescence or the increase of EtBr that is retained by the bacteria prior to and after the addition of the uncoupler (or other type of inhibitor), is usually determined using specialized and expensive fluorometric instrumentation that may not be available in a clinical bacteriology laboratory [5, 7]. The principle of these fluorometric assays is the passage of EtBr across the cytoplasmic membrane and its subsequent intracellular accumulation inside the bacterial cell [8]. EtBr traverses the bacterial cell wall (in the case of Gram-negative bacteria via porin channels) and once inside, it can be concentrated to a point where it fluoresces when excited by ultraviolet (U.V.) light. Efflux pumps of MDR bacteria recognize this substrate and are able to extrude it to the medium [6, 8]. These efflux systems are temperature dependent [9] and this process will continue if the concentration of EtBr in the culture medium does not overcome the capacity of the bacterial efflux pump itself. Therefore, loading of the bacteria with EtBr has to take place at a concentration that is well below its minimum inhibitory concentration (MIC) [4, 5]. This way, the signal that is generated can be continuously recorded under specific conditions, such as time, temperature and different properties of the medium (for example: pH; calcium availability; etc.) [5, 6]. Efflux of EtBr can be readily shown by the use of standard fluorometers or cytometers [7, 10, 11]. However, when efflux is due to over-expressed efflux pumps, it cannot be easily shown/assessed by some of the conventional methods previously referred to [5, 12]. The current fluorometric systems used to access efflux activity are in general non-physiological; usually the control of the temperature is restricted; the methods are cumbersome and do not yield data that can be subjected to standardization for intra-laboratory comparison. Moreover, due to the wide variety of instrumentation, reagents, media, etc., used by the many laboratories working in this area, it is difficult to compare and standardize the results obtained from these studies [4, 5]. In the recent years there have been several methods published that can assess efflux in a more dynamic and physiological way making use of fluorescent probes, such as EtBr, Hoechst-33342 (bisbenzimide) or fluorogenic compounds (for example: Fluorescein-di-β-D-galactopyranoside) or using lipophilic dyes, such as Nile Red [4, 5, 6, 12-15]. The majority of these methods use a liquid system and need equipment that may or may not be available in all the clinical laboratories. Therefore, there is still an obvious need to establish new and optimized assays for the assessment of efflux pumps of distinct bacteria, in particular those of clinical origin showing an MDR phenotype. Additionally, there is still a lack of methods that make use of a solid system and simple equipments. A new assay should be simple, quick to perform, reproducible and should excuse any specialized instrumentation for its conductance [16]. In MDR bacteria, the assessment of efflux pump activity can help to characterize the basis by which novel/existing compounds inhibit efflux activity, consequently rendering the organism susceptible to one or several of the antibiotics to which it was initially resistant [8, 17-19]. If those compounds were available for clinical therapy, it would be expected that the assay would have extensive implementation within the hospital/clinic and provide much of the guidance needed for the administration of the compounds to patients infected with an MDR bacteria [19]. Moreover, the availability of such compounds brings benefits to the therapeutic regimens, resulting in the opportunity to use outdated, inexpensive and safe antibiotics that had been made redundant as a consequence of the emergence of MDR bacteria [20]. To describe and characterize in detail the activity of an efflux inhibitor, the method by which that activity is defined and quantified must be reliable and feasible to apply to a large number of inocula, yielding inter-laboratory standardization.
ETHIDIUM BROMIDE (ETBR)-AGAR CARTWHEEL METHOD
This simple, instrument-free, agar-based method utilizes EtBr for the demonstration of efflux pump activity in bacteria [21]. It is an improved and optimized version of the previously described EtBr-agar method [16]. This new optimized method can be applied simultaneously to up to twelve bacterial strains to identify clinical isolates that have an over-expressed efflux activity [21]. Each plate should include at least one reference strain that will serve as a comparative control for fluorescence analysis. However, the number of reference strains can be increased to two or more, depending on a given experiment. In addition, it allows the evaluation of agents that may inhibit this same activity. The effect of temperature on the efflux pump system can also be assessed/confirmed. The methodology used is straight-forward employing the preparation of two sets of Trypticase Soy Agar (TSA) plates containing EtBr concentrations ranging from 0.0 to 2.5 mg/L (these concentrations may vary according to the bacterial strain in study). The TSA plates should be prepared fresh on the previous or same day of the experiment and kept protected from light. Overnight cultures of the bacterial isolates to be tested are prepared in liquid media and in the following day their concentration adjusted to 0.5 of a McFarland standard. The TSA plates are then divided into as many as twelve sectors by radial lines, forming a cartwheel pattern (Fig. 1). The adjusted bacterial cultures are then swabbed on the EtBr-TSA plates starting from the center of the plate to the margin. The TSA plates are then incubated at 37ºC for 16 hours. After this period, the TSA plates are examined under a gel-imaging system (or a U.V. transilluminator); the minimum concentration of EtBr that produces fluorescence of the bacterial mass recorded; and the TSA plates photographed (Fig. 1).
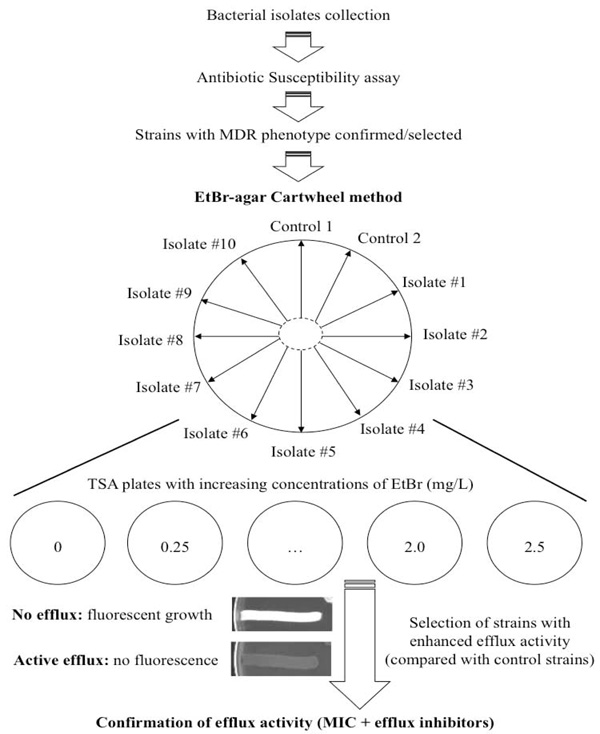
Flowchart followed to test bacterial strains using the EtBr-agar Cartwheel method. MDR strains were swabbed in TSA plates con-taining different concentrations of EtBr and incubated for 16 hours at 37ºC. Controls and clinical isolates were swabbed on EtBr-containing TSA plates, according to the diagram. Each EtBr-TSA plate can accommodate as many as twelve bacterial strains. The distribution of the strains in the TSA plate can be altered according to the desired experiment. After this, fluorescence was recorded and strains that showed lower fluorescence than the control (indicative of efflux activity) were selected. The efflux activity was further confirmed by determining the MIC of antibiotics in the presence of efflux inhibitors. Two control strains can be inserted in this study. For example, Control 1 should pre-sent the highest fluorescence (evidence of no efflux activity or physiological activity) and Control 2 should be a strain showing no fluores-cence or very low levels of fluorescence (indicative of an active over-expressed efflux system). These strains should be previously well char-acterized for their efflux systems.
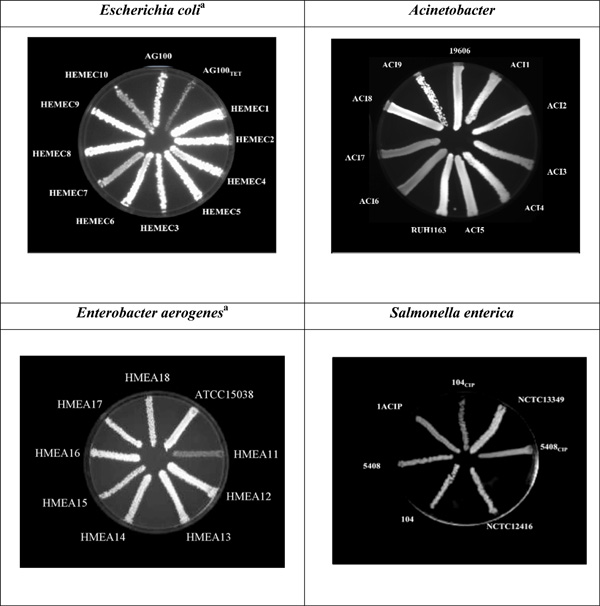
The EtBr-agar cartwheel method applied to Gram-negative bacteria.
TET – Tetracycline; CIP – Ciprofloxacin. For the E. coli and Acinetobacter strains, the TSA plates shown contain 2 mg/L of EtBr while for Enterobacter aerogenes and Salmonella the agar plates contain 1.5 mg/L and 2.5 mg/L of EtBr, respectively. adata previously published in [21].
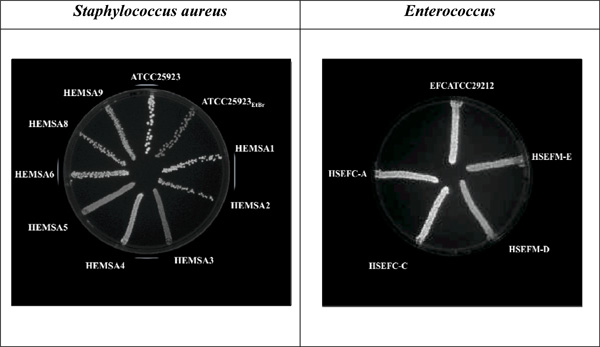
The EtBr-agar cartwheel method applied to Gram-positive bacteria.
Cultures were swabbed in TSA plates containing increasing concentrations of EtBr. Following overnight incubation at 37°C for 16 hours, fluorescence was detected under UV light. The concentration shown above as an example for both strains is 2.0 mg/L of EtBr. Note: Data previously published in [21]. EFCATCC - Enterococcus faecalis reference strain; HSEFC – Enterococcus faecalis; HSEFM – Enterococcus faecium.

The effect of the temperature in the efflux activity of Acinetobacter strains screened by the EtBr-agar cartwheel method. The strains were streaked in the EtBr-TSA plates following the following scheme:
In the case that one wants to check for the temperature effect, then an additional incubation step should be performed. In this case, one of the TSA plate sets should be re-incubated at 37ºC, whereas the duplicate set is transferred to 4ºC. At the end of an additional 24 hours period, the TSA plates are observed and photographed again and the minimal concentration of EtBr that produced fluorescence at each temperature will be compared to that evident after the first incubation (at 37ºC).
TESTING OF THE METHOD IN CLINICALLY RELEVANT MDR GRAM-NEGATIVE BACTERIA
All the clinical isolates evaluated by the EtBr-agar cartwheel method and inserted in this study have a confirmed MDR phenotype. This MDR was defined as resistance to three or more different classes of antibiotics [22]. Identification and antibiotic susceptibility profile of the isolates was done using the VITEK2 system and provided by the microbiology laboratories of the collaborating hospitals (bio Mérieux, Marcy l´Etoile, France). The antibiotic susceptibility data was also confirmed in our laboratory by the Kirby-Bauer method, following the Clinical and Laboratory Standards Institute (CLSI) guidelines (CLSI, 2006). Results were evaluated according to the CLSI breakpoints.
In a first approach it is necessary to establish the conditions in which the assays should be performed. Therefore, minimum fluorescence values shown by the bacterial isolates should be determined. We have tested a collection of clinical isolates including, Gram-positive and Gram-negative strains. Reference strains were used in each of the assays for comparison terms and to help determine the fluorescence baseline of each isolate. Clinical isolates from Gram-negative bacteria included: Escherichia coli, Acinetobacter baumannii, Acinetobacter genomic species 3, Enterobacter aerogenes and Salmonella enterica serovars Enteritidis and Typhimurium.
Escherichia coli K12 and Escherichia coli AG100TET strains were used as controls due to the previous well characterised expression of their efflux pumps system. Escherichia coli K12 AG100 wild-type strain (argE3 thi-l rpsL xyl mtl ∆(gal-uvrB) supE44), contains a fully functional AcrAB-TolC efflux pump system. E. coli AG100TET, is a AG100 progeny strain that was induced to high levels of resistance to tetracycline (TET) (MICTET of 12 mg/L) and over-expresses acrAB [23]. E. coli K-12 AG100 has been previously characterized and was kindly offered by Hiroshi Nikaido, University of California, Berkeley, California, USA [4, 24]. When tested by the Ethidium bromide (EtBr)-agar method E. coli AG100 (wild-type) shows fluorescence at a concentration of 0.5 mg/L of EtBr while its isogenic E. coli AG100TET (induced to high level resistance to TET and expressing the AcrAB efflux pump 6-fold higher than that of its parent [16, 21]) does not fluoresce with a concentration of EtBr as high as 2.5 mg/L (data not shown). In the case of the MDR E. coli clinical isolates, two of the ten isolates show less fluorescence than the remaining ones at the concentration of 2.5 mg/L of EtBr (Fig. 2).
A. baumannii ATCC19606 and Acinetobacter genomic species 3 RUH1163 were used as reference strains. As visible in Fig. (2), the two Acinetobacter strains resistant to carbapenem (namely meropenem (MER) and imipenem (IMI)), ACI6 and ACI7 don’t show fluorescence at the maximum concentration of EtBr tested (2 mg/L).
The method was also applied to Enterobacter aerogenes MDR clinical strains. Enterobacter aerogenes ATCC15038 was used as a reference. Enterobacter aerogenes HMEA11 shows no fluorescence at 1.5 mg/L EtBr while the ATCC and the other Enterobacter aerogenes strains show fluorescence at this same concentration (Fig. 2).
In the case of the Salmonella strains, Salmonella Typhimurium NCTC12416 and Salmonella Enteritidis NCTC13349 were used as reference strains since these are fully characterized strains. Salmonella Enteritidis 104 and 5408 and their respective progeny 104CIP, 1ACIP, and 5408CIP that were induced to high level resistance to ciprofloxacin (CIP) by serial passage in media containing increasing concentrations of the antibiotic [25] were also inserted in the study. In the case of the Salmonella Enteritidis, the CIP induced strains 5408CIP and 104CIP do not fluoresce at the highest concentration of EtBr used in the assay (2.5 mg/L EtBr) (Fig. 2). These strains have been shown to over-express their AcrAB efflux pump 6-fold over that of their CIP susceptible isogenic parental strains [26].
APPLICATION OF THE METHOD TO CLINICAL RELEVANT MDR GRAM-POSITIVE BACTERIA
The EtBr-agar cartwheel method was also applied to Gram-positive pathogens causing infections in the clinical setting, namely, Staphylococcus aureus, Enterococcus fae-cium and Enterococcus faecalis. S. aureus ATCC25923EtBr (MIC for EtBr of 200 mg/L) is a progeny strain derived from S. aureus ATCC25293 (MIC for EtBr of 6 mg/L) that was induced to increased resistance to this dye [27] and was also inserted in this study as reference for fluorescence comparison. The fluorescence of the clinical isolates identified as HEMSA3, HEMSA4, HEMSA5 and HEMSA9 is greater than the one produced by the reference strain (S. aureus ATCC25923) and is almost as great as that needed for the fluorescence of the S. aureus ATCC25923EtBr strain that had been induced to high level resistance to EtBr and which over-expresses its NorA efflux pump [27] (Fig. 3). This fluorescence can also be analyzed based on the Index for efflux activity of the isolates. This index allows classifying the level of efflux demonstrated by these strains, in order to rank the capacity to efflux EtBr of each bacterial strain (in comparison with the reference strain) [21].
Efflux of EtBr was also analyzed among reference and clinical enterococci. It was visible the difference in fluorescence between the three E. faecalis strains and the two clinical E. faecium (HSEFM-D and HSEFM-E). While at a concentration of 2 mg/L EtBr the Enterococcus faecalis strains show fluorescence, the E. faecium strains are starting to fluoresce [5, 21] (Fig. 3).
THE EFFECT OF TEMPERATURE IN THE EFFLUX LEVEL OF THE CLINICAL ISOLATES
The EtBr-agar cartwheel method was also applied to test the effect of temperature on the efflux activity of the clinical isolates. Drug efflux is an energy-mediated transport and by using a low temperature that decreases the membrane energy we can decrease the efficiency of efflux transporters [6, 8]. As an example, two sets of TSA plates were streaked with the Acinetobacter strains and incubated at 37°C for a period of 18 hours. After this period, the fluorescence of the strains or the absence thereof was recorded (Fig. 4). The TSA plates were then re-incubated for an extra 24 hours; one set was returned to 37°C and the other transferred to 4°C to test if the efflux activity was the major contributor to the low levels of fluorescence previously registered. After this period the TSA plates were photographed again (Fig. 4). After a period of incubation at 4°C, strains that didn’t show fluorescence at 37°C were highly fluorescent at the higher concentrations of EtBr tested in the agar plate. The same was obtained for all the other strains that didn’t show fluorescence or showed lower fluorescence than the reference strains (data not shown).

ACI – Acinetobacter clinical isolate.
The effect of the temperature on the efflux activity of the bacterial strains can be assessed by the EtBr-agar cartwheel method. As an example, a set of Acinetobacter strains is illustrated in the figure. As visible, after a period of incubation at 4°C, strains that didn’t show fluorescence at 37°C are highly fluorescent at the higher concentrations of EtBr tested in the TSA plate.
CONFIRMATION OF THE RESULTS PROVIDED BY THE ETBR-AGAR CARTWHEEL METHOD
The results provided by the EtBr-agar cartwheel method were further explored by the additional determination of the MIC values for selected antibiotics, known to be efflux pump substrates (e.g. tetracycline, norfloxacin, imipenem, etc.), in the presence of compounds described as having an inhibitory effect on efflux activity. These compounds can include a variety of classes, such as efflux inhibitors, permeabilizers, chemosensitizers, etc. Ultimately, the intended use of these compounds is to confirm the activity of over-expressed efflux system(s) in the MDR clinical strains identified by this method. This confirmation is important to rule out other factors that may affect fluorescence such as a decrease in the cellular permeability to EtBr [6, 8, 28, 29]. Isolates that fluoresced at concentrations of EtBr significantly greater than their reference strains were evaluated for their susceptibility to a panel of antibiotics known to be substrates of efflux pump systems described for each bacterial species, in the presence and/or absence of compounds known to inhibit efflux activity. For comparative terms, representative strains showing fluorescence at low EtBr concentrations were also evaluated (Tables 1 and 2).
The Effect of Selected EPIs on the Resistance of the Induced and MDR Gram-negative Bacteria
| Species | Strain | MIC (mg/L) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| TET | CIP | ||||||||
| - | + TZ | + CPZ | + PAN | - | + TZ | + CPZ | + PAN | ||
| E. colia | AG100 | 1 | 0.5 | 0.5 | 0.5 | 0.015 | 0.003 (5x) | 0.003 (5x) | 0.0075 |
| AG100TET | 12 | 0.375 (32x) | 3 (4x) | 3 (4x) | 0.06 | 0.015 (4x) | 0.03 | 0.03 | |
| HEMEC7 | 128 | 32 (4x) | 16 (8x) | 32 (4x) | 0.25 | 0.125 | 0.125 | 0.125 | |
| HEMEC10 | 64 | 32 | 16 (4x) | 32 | 16 | 8 | 4 (4x) | 2 (8x) | |
| Enterobacter aerogenesa | ATCC15038 | TET | NOR | ||||||
| - | + TZ | + CPZ | + PAN | - | + TZ | + CPZ | + PAN | ||
| 0.5 | 0.25 | 0.125 (4x) | 0.5 | 0.025 | 0.0125 | 0.025 | 0.025 | ||
| HMEA11 | 2 | 1 | 0.25 (8x) | 1 | 0.03 | 0.007 (4x) | 0.003 (10x) | 0.015 | |
| Salmonellaa | NCTC13349 | TET | CIP | ||||||
| - | + TZ | + CPZ | + PAN | - | + TZ | + CPZ | + PAN | ||
| 1 | 0.5 | 0.5 | 0.5 | 0.015 | 0.007 | 0.015 | 0.007 | ||
| 5408CIP | 1 | 0.25 (4x) | 0.5 | 0.5 | 32 | 8 (4x) | 16 | 8 (4x) | |
| Acinetobacter | MER | IMI | |||||||
| - | + TZ | + CPZ | + PAN | - | + TZ | + CPZ | + PAN | ||
| ATCC 19606 | 4 | 2 | 2 | 0.25 (3x) | 8 | 4 | 4 | 0.5 (4x) | |
| ACI7 | 128 | 64 (2x) | 0.25 (9x) | 16 (3x) | ≥ 256 | 128 | 128 | ≥ 256 | |
| ACI3 | 128 | 128 | 128 | 64 (2x) | ≥ 256 | 128 | 128 | ≥ 256 | |
TET - tetracycline; CIP - ciprofloxacin; NOR - norfloxacin; MER – meropenem; IMI – imipenem; TZ - thioridazine; CPZ - chlorpromazine; PAN - Phe-Arg-napthylamide. aparcial data previously published in [21]. Solutions of TZ, CPZ and PAN (Sigma Aldrich, SA, Madrid, Spain) were prepared in deionized water. Aliquots of stock solutions of the phenothiazines CPZ and TZ were protected from light and stored at -20ºC. Working solutions were prepared on the day of the experiment. To assure that the efflux inhibitors do not compromise the bacterial viability, each compound was employed at a concentration of 1/2 of the MIC. Exceptions to this was PAN, known to affect the cellular viability even at 1/2 of the MIC [30] and in this case the compound was used at a maximum of 20 mg/L (value well below their pre-determined MIC). Values in bold-type correspond to a decrease of 4-fold or higher on the MIC values in comparison to the ones in the absence of inhibitor. Values in parenthesis indicate the MIC decrease relative to the one of the original culture. A reduction in the MIC of at least one-fourth of their original values in the presence of the efflux inhibitor was considered indicative of the presence of efflux activity. All assays were performed in triplicate.
The Effect of Selected Efflux Inhibitors on the Resistance of Induced and MDR Gram-positive Bacteria
| MIC for NOR (mg/L) | |||||
|---|---|---|---|---|---|
| Species | Strains | - | + TZ | + CPZ | + RES |
| S. aureusa | ATCC25923 | 0.5 | 0.125 (4x) | 0.125 (4x) | 0.25 |
| ATCC25293EtBr | 2 | 0.25 (8x) | 0.25 (8x) | 0.5 (4x) | |
| HEMSA 3 | 256 | 128 | 64 (4x) | 128 | |
| HEMSA 4 | 512 | 128 (4x) | 128 (4x) | 256 | |
| HEMSA 5 | 64 | 16 (4x) | 16 (4x) | 32 | |
| HEMSA 9 | 128 | 32 (4x) | 64 | 64 | |
| MIC for TET (mg/L) | |||||
| Enterococcusa | - | + TZ | + CPZ | + RES | |
| EFC ATCC29212 | 4 | 4 | 4 | 4 | |
| HSEFM-D | 16 | 4 (4x) | 4 (4x) | 8 | |
| HSEFM-E | 0.125 | 0.125 | 0.125 | 0.125 | |
TET - tetracycline; NOR - norfloxacin; CIP - ciprofloxacin; TZ - thioridazine; CPZ - chlorpromazine; RES - reserpine. aparcial data previously published in [21]. EFCATCC - Enterococcus faecalis reference strain; HSEFC – Enterococcus faecalis; HSEFM – Enterococcus faecium. Solutions of TZ and CPZ (Sigma Aldrich, SA, Madrid, Spain) were prepared in deionized water; RES (Sigma Aldrich, SA, Madrid, Spain) was prepared in dimethylsulfoxide. Aliquots of stock solutions of the phenothiazines CPZ and TZ were protected from light and stored at -20ºC. Working solutions were prepared on the day of the experiment. To assure that the efflux inhibitors do not compromise the bacterial viability, each compound was employed at a concentration of1/2 of the MIC. An exception to this is RES, known to affect the cellular viability even at 1/2 of the MIC [30] and in this case the compound was used at a maximum of 20 mg/L (value well below their pre-determined MIC). Values in bold-type correspond to a decrease of 4-fold or higher on the MIC values in comparison to the ones in the absence of inhibitor. Values in parenthesis indicate the MIC decrease relative to the one of the original culture. A reduction in the MIC of at least one-fourth of their original values in the presence of the efflux inhibitor was considered indicative of the presence of efflux activity. All assays were performed in triplicate.
Chemical Structures of EtBr, Antibiotics and Efflux Inhibitors Used in this Method
| Compounds | Designation | Acronym | Chemical Structure |
|---|---|---|---|
| DYE | Ethidium Bromide | EtBr |  |
| ANTIBIOTICS | Tetracycline | TET |  |
| Ciprofloxacin | CIP |  |
|
| Norfloxacin | NOR |  |
|
| Imipenem | IMI |  |
|
| Meropenem | MERO |  |
|
| EFFLUX INHIBITORS | Thioridazine | TZ |  |
| Chlorpromazine | CPZ |  |
|
| Phenylalanine-Arginine beta-Naphthylamide | PAN |  |
|
| Reserpine | RES |  |
The efflux inhibitors used for Gram-negative and Gram-positive bacteria can vary but some extensively studied compounds are usually used such as: thioridazine (TZ), chlorpromazine (CPZ), Phenylalanine-Arginine beta-Naphthy-lamide (PAN) (in the case of Gram-negative bacteria) and TZ, CPZ and reserpine (RES) (in the case of Gram-positive bacteria) Table 3. To assure that the bacterial viability is not compromised by the presence of the efflux inhibitor itself, each compound was employed at a concentration of 1/2 of the MIC. Exceptions to this were PAN and RES; in the case of PAN, it is known to affect the cellular viability or membrane permeability even at 1/2 of the MIC [13, 30]. In these specific cases (PAN and RES) the compounds were used at a maximum of 20 mg/L (value well below their pre-determined MIC). A reduction in the MIC of at least 1/4 of their original values in the presence of the efflux inhibitor was considered indicative of the presence of efflux activity. All assays were performed in triplicate.
As can be seen by the data presented in Tables 1 and 2, there were significant reductions in the MIC values in the presence of the efflux inhibitors. For example, in the case of the E. coli strains there was a considerable reduction of the MIC of TET in the presence of the three compounds tested. This was observed in the TET-resistant strain and one the clinical isolates that showed lower levels of fluorescence (when compared with the control strain). Other reductions of the MIC were also obtained with CIP in the presence of the three inhibitors. The same was obtained for the Enterobacter aerogenes clinical isolates. Examples for these strains are shown in Table 1. While the clinical isolate HMEA11 showed a reduction of the MIC to TET in the presence of CPZ, the MIC for NOR was reduced in the presence of CPZ as well as TZ. For the Salmonella strains an example is presented for TET and CIP, showing that TZ contributed to a reduction of the MIC (4-fold reduction) of these antibiotics (Table 1). In the case of the Acinetobacter strains showed a reduction of the MIC of MER in the presence of CPZ and PAN. In the case of the Gram-positive strains, Staphylococ cus and Enterococcus there was a reduction of the MIC for NOR in the presence of TZ and CPZ for almost all of the strains tested; this reduction varied from 4- to 8-fold. Reserpine was able to reduce the MIC of NOR in the case of the EtBr-induced strain. In general, it was possible to confirm various degrees of efflux activity in the strains tested and that were previously identified and selected by the EtBr-agar cartwheel method.
CONCLUDING REMARKS
Bacterial resistance to antibiotics has become a serious concern for the public health setting. Concomitantly, the role that efflux systems play in antibiotic resistance in MDR bacteria is an important subject that has been extensively discussed in recent years [1, 3, 22, 28]. Although high-level resistance may not occur as a result of MDR efflux pumps alone, the association of over-expression of specific genes among highly resistant clinical isolates cannot be ignored [3, 24-26]. Therefore, we should bear in mind that the intrinsic resistance to antibiotics of some isolates may be largely due to efflux systems [19, 24, 31]. Synergic increases in resistance seen with over-expression of efflux systems, as well as target site mutations can lead to highly resistant bacteria that are difficult to treat with the antibiotics that are currently available. The contribution of efflux to the resistance seen in some clinical strains needs to be considered as one of the parameter used in the design of future antibiotics or any other active compounds [32, 33]. In any event, alterations of the structure of a given antibiotic should be made to reduce the ability of that same antibiotic to be extruded from the bacterial cell, but without compromising its activity [8, 32, 34]. Another approach could be based on potentiating the activity of antibiotics by these inhibitors/novel compounds [19, 20, 30, 32, 33]. The development of compounds that can act as inhibitors/blockers of efflux thereby reducing the impact of these pumps on the efficacy of certain antibiotics, will be of clinical interest and could impact greatly on the clinical setting. However, the major challenge persists: to discover a compound that can be used as an efflux blocker in a non-specific manner, i.e., that can target a big range of efflux systems from different bacterial species. This could be considered a tangible goal as many of these systems show a high degree of structural homology between different bacterial species [28, 29, 31, 34]. This approach can contribute to the design of more realistic synergic assays that can combine antibiotics and blockers to reduce the MIC for a given antibiotic to a clinical relevant level [20, 32, 33]. That way, bacteria initially resistant to antibiotics can become susceptible. If this approach proves to be successful in the future it can be used as an important alternative in the treatment of some of the most important MDR infections that we have to deal with nowadays [19, 20, 32, 35].
In summary, the EtBr-agar cartwheel method is easy to perform, less time-consuming and can be used to screen large numbers of bacterial strains, thereby facilitating the rapid identification of isolates displaying an MDR phenotype. It can be used in both Gram-negative and Gram-positive clinical isolates to detect MDR mediated by efflux. It is also possible to establish a rapid comparison of efflux activity of laboratory derived isogenic mutants that were manipulated, in the case of strains that were adapted to a given antibiotic by sequential passage exposure; by deletion or interruption of a specific gene or cluster of genes; or by growing the strain in different conditions, for example: different temperature; different pH of the medium; others. Taken together, these factors may play a role in the efflux activity of the isolates, and all of these can be rapidly screened by the described Ethidium Bromide-agar Cartwheel method.
LIST OF ABBREVIATIONS
CONFLICT OF INTEREST
The authors confirm that this article content has no conflicts of interest.
ACKNOWLEDGEMENTS
This work was supported by grants EU-FSE/FEDER-PTDC/BIA-MIC/71280/2006 and EU-FSE/FEDER-PTDC/ BIA-MIC/105509/2008 provided by the Fundação para a Ciência e a Tecnologia (FCT, Portugal). M. Martins was supported by grant SFRH/BD/14319/2003 (FCT, Portugal). This work was also partially supported by Grant 06/TNI/ UCD/10 provided by the Department of Agriculture, Fisheries and Food (DAFF), Ireland, under the Food Institutional Research Measure (FIRM) Network & Team Building Initiative 2006 and by COST Action BM0701 (ATENS).


