RESEARCH ARTICLE
Biodegradation of Selected Nigerian Fruit Peels by the use of a Non-pathogenic Rhizobium species CWP G34B
Bolatito Esther Boboye*, George Olarewaju Ajayi
Article Information
Identifiers and Pagination:
Year: 2012Volume: 6
First Page: 88
Last Page: 93
Publisher ID: TOMICROJ-6-88
DOI: 10.2174/1874285801206010088
Article History:
Received Date: 02/11/2011Revision Received Date: 09/12/2011
Acceptance Date: 24/1/2012
Electronic publication date: 2/11/2012
Collection year: 2012

open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
Abstract
This study was carried out to determine the ability of Rhizobium species CWP G34B to degrade the peels of selected Nigerian fruits. The potential of the bacterium to digest some carbon sources (lactose, maltose, sucrose and mannitol) and peels of some Nigerian fruits (pineapple, orange, plantain, banana, pawpaw and mango fruits) was investigated by growing the organism on the substances separately after which DNSA reagent method was used to quantify glucose released into the medium. The results showed that the bacterium was able to degrade all the carbohydrates with the highest and the lowest glucose concentrations of 5.52 mg/ml for lactose and 0.50 mg/ml for mannitol. The carbohydrate-catabolic-enzyme (CCE) activity ranged from 0.169 mg/ml to 1.346 mg/ml glucose per mg/ml protein. Mannitol exhibited the highest CCE activity while the lowest activity was observed in the presence of sucrose. The amount of extracellular protein synthesized was highest (9.803 mg/ml) in the presence of maltose and lowest (0.925 mg/ml) in mannitol. The mean polygalacturonase activity was 0.54 unit/ml when the bacterium was grown in pectin in contrast to 0.28 unit/ml when it was grown in mannitol. The bacterium showed ability to breakdown the peels of the Nigerian fruits with the highest capability in banana and pineapple (0.42 and 0.41 mg/ml glucose per mg/ml protein respectively). The fruit-peel-degrading enzyme activity was lowest in orange peel (0.75 unit/ml).
INTRODUCTION
Peel, also known as rind or skin, is the outer protective layer of a fruit or vegetable. Botanically, the rind is usually the exocarp, which includes the hard shell in fruits such as nuts. Depending on the thickness and taste, peel is sometimes eaten as part of the fruit, as seen with apples. In some fruits such as banana or grape, the peel is unpleasant or inedible; thus, it is removed and discarded [1]. Oladiji et al. [2] reported that most fruit peels are discarded as waste after the inner fleshy portions have been eaten.
It is vital that peels be removed from most fruits before eating; and more importantly before using them in fruit juice industries to prevent contamination. Processing of fruits into juices reduces and prevents wastage when fruits are in season. Olukunle et al. [3] opined that fruit juice is the next best thing to fresh fruit, and can be packaged in aseptic, easily transportable containers that are less susceptible to damage and have a relatively long storage life. Juice extraction and separation therefore open up new market opportunities for tailoring fruit products to modern consumer demands.
At the time of producing fruit juice, a lot of peels are produced. This could cause environmental pollution and health problems if left untreated. Peels can be removed manually, mechanically and by the use of enzymes. A lot of money, time, equipment and other resources are used to remove the peels in the industry. Enzymatic removal of peels could be cheaper and more effective than manual and mechanical methods. The use of enzyme in the manufacture of various industrial products is wide spread [4]. Development of microbes that will synthesize these enzymes will then be useful to man. Pectate lyase (PL) is one of those enzymes. The PL is otherwise known as pectate transeliminases; it catalyses the eliminative cleavage of de-esterified pectin, which is a major component of the primary cell walls (peels) of many higher plants [5, 6]. The backbone of this pectic polysaccharide (de-esterified pectin) is built up with blocks of α–1,4 linked polygalactosyluronic acid residues inter-spersed with regions of alternating galactosyluronic acid and rhamnosyl residues [7].
Various microbes are known to synthesize pectolytic enzymes. Xanthomonas campestris, Erwinia and Streptomyces have been reported to produce pectate lyases [8-10]. Microorganisms that produce pectinases are mostly pathogenic in nature [11, 12]. Until recently, it was thought that pectolytic enzymes were secreted mainly by plant pathogens, which actions result to maceration of plant tissues. Non-pathogenic microorganisms will be safer for use in the production of these enzymes than pathogens. Rhizobia, a group of non-pathogenic bacteria known to form nodules on legumes through the digestion of external walls of legume’s roots, may be a potential candidate that can be used to synthesize these enzymes. Therefore, this study was designed to evaluate the ability of Rhizobium species CWP G34B to synthesise enzymes that can breakdown the peels of selected Nigerian fruits. This investigation will provide scientific information on the potential of Rhizobium species CWP G34B for possible use industrially and in the control of environmental pollution.
MATERIALS AND METHODS
Samples and Test Bacterium
Dried peels of pineapple, orange, plantain, banana, pawpaw and mango were ground using Marlex Electroline Blender. Lactose, maltose, sucrose, mannitol and Rhizobium species CWP G34B were provided by the Department of Microbiology at the Federal University of Technology, Akure, Nigeria.
Culture Media
The commercial media used in this study were nutrient agar, nutrient broth and agar agar (Lab M., Topley House, England). All the media were prepared according to the manufacturer’s specification.
Minimal medium (MM) was prepared by dissolving 1.0g of disodium hydrogen phosphate (Na2HPO4), 0.2g of potassium chloride (KCl) and 0.2g of magnesium sulphate (MgSO4) in 1 litre of distilled water. All the media were sterilized by autoclaving at 121oC for 15 minutes. The agar was allowed to cool before pouring into sterile Petri dishes.
Determination of Carbohydrate-Degrading Ability and Carbohydrate-Catabolic-Enzyme (CCE) Activity of Rhizobium species CWP G34B
Sterile minimal medium (MM) containing each carbohydrate (1.0 w/v), was inoculated with an overnight (18 hours) grown culture of Rhizobium species CWP G34B and incubated at 28oC for 24 hours. Each grown culture was centrifuged at 3600rpm for 15 minutes. The supernatant was transferred into a fresh tube as a source of extracellular glucose and enzyme. Dinitrosalicylic acid (DNSA) reagent method [13] was used to assay the amount of glucose released into the growth medium as described by Boboye and Alao [14]. The supernatant (0.5ml) was mixed with 0.5ml DNSA reagent, left for 15 minutes at 28oC and boiled for 5 minutes. The tubes were rapidly cooled under tap water and the optical density of the reaction mix was measured at 540 nm. The OD values were referred to a glucose standard curve to estimate the amount of glucose released into the medium by the bacterium during growth.
The CCE activity was based on the concentration of glucose released from the carbohydrate per milligram of protein produced by the bacterium [14]. Glucose concentration was measured according to DNSA reagent method [14]. The concentrations of extracellular glucose and protein formed by the Rhizobium were quantified by Biuret method [14, 15].
Measurement of Fruit-Peel-Degrading (FPD) Ability of the Rhizobium
A 10ml of each fruit peel (10% w/v) was dispensed into test tubes. They were autoclaved at 121oC for 15 minutes and allowed to cool. Each fruit peel was inoculated with 0.2ml of 24 hours old inoculums and incubated at 28oC for 24 hours. Each culture was centrifuged 3600 rpm for 15 minutes. The glucose and protein concentrations of the supernatant were determined using DNSA and Biuret reagent methods [14]. The extracellular Fruit-Peel-Degrading Enzyme (FPDE) activity of bacterium was defined as mg/ml glucose per mg/ml protein in the supernatant.
Statistical Analysis of Data
The data collected were analyzed using the analysis of variance (ANOVA) technique and expressed as means of values. Duncan Multiple Range Test was carried out to determine differences in the means using SPSS Software package [16].
RESULTS
Carbohydrate-Degrading Ability of Rhizobium species CWP G34B
There was considerable (at 95% confidence limit) degradation of each carbohydrate by the Rhizobium species CWP G34B within 24 hours of incubation. The amount of glucose released into the minimal growth medium is shown in Fig. (1). Concentration of the glucose is lowest in the growth medium containing mannitol (0.50 mg/ml) and highest (5.52 mg/ml) in lactose.
 |
Fig. (1). Quantity of glucose released by Rhizobium species CWP G34B during growth in some carbohydrates. |
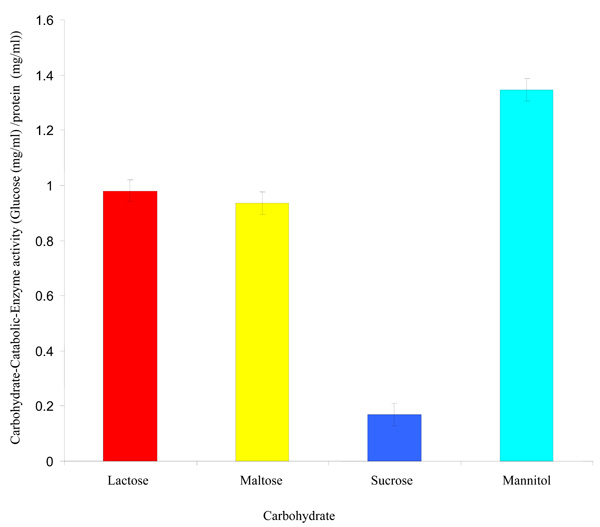 |
Fig. (2). Carbohydrate-Catabolic-Enzyme activity of Rhizobium species CWP G34B. |
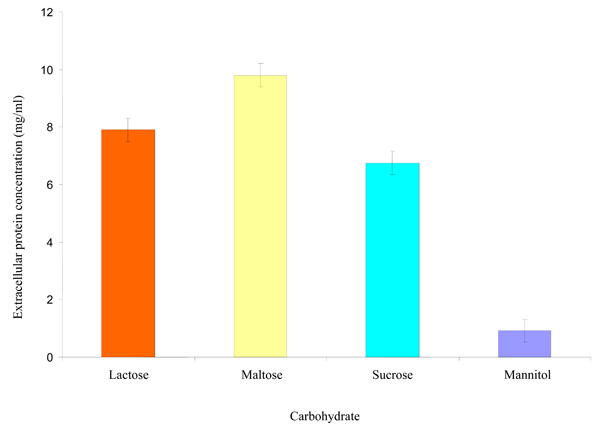 |
Fig. (3). Effect of some carbohydrates on extracellular protein synthesized by the Rhizobium species CWP G34B. |
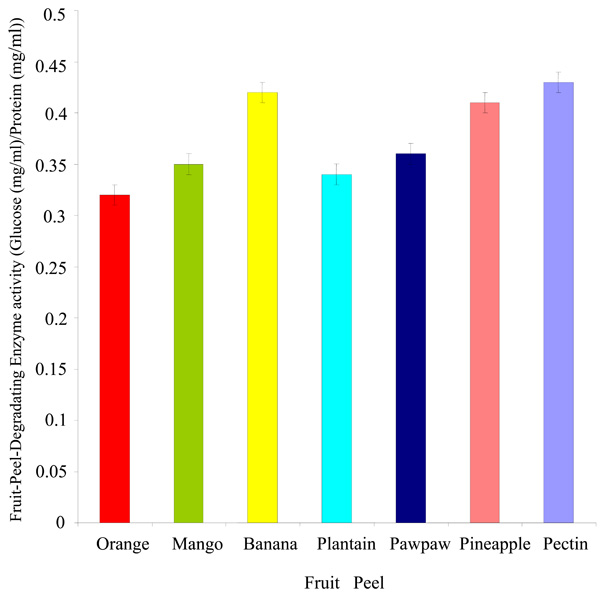 |
Fig. (4). Degradation of fruit peels by Rhizobium species CWP G34B. |
Carbohydrate-Catabolic-Enzyme (CCE) Activity
The Rhizobium species CWP G34B grown in minimal medium containing each carbohydrate (lactose, sucrose, mannitol and maltose) showed varying degrees of the carbohydrate-catabolic-enzyme (CCE) activity at a probability level of ≤ 0.05 (Fig. 2). The CCE activity ranged from 0.169 mg/ml to 1.346 mg/ml glucose per mg/ml protein. Mannitol exhibited the highest CCE activity while the lowest activity was shown in the presence of sucrose.
Effect of some Carbohydrates on Extracellular Protein Synthesized by Rhizobium species CWP G34B
The Rhizobium species showed ability (P = ≤ 0.05) to synthesize protein when cultivated separately in minimal medium containing each of the carbohydrates separately. The amount of extracellular protein synthesized was highest in maltose with a concentration of 9.803 mg/ml followed by lactose (7.900 mg/ml) and sucrose (6.753 mg/ml) (Fig. 3). Concentration of the protein synthesized was lowest in mannitol.
Degradation of Fruit Peels by Rhizobium species CWP G43B
The Rhizobium spp. CWP G34B degraded the peels of fruits used in this study (Fig. 4). The bacterium showed a greater ability to breakdown the peels of banana and pineapple (0.42 and 0.41 mg/ml glucose per mg/ml protein respectively). The fruit-peel-degrading enzyme (FPDE) activity expressed in the peels of orange, mango, plantain and pawpaw was between 0.32 and 0.36 mg/ml glucose per mg/ml protein. The lowest degrading enzyme activity was exhibited in orange. In comparison with the fruit peels, the polygalacturonase enzyme activity in the presence of pectin appeared to be higher than the values of FDPE activity of any of the fruit peels.
DISCUSSION
The experiment showed that the organism was able to breakdown all the carbohydrates significantly (P = ≤0.05), indicating that R. spp. CWP G34B was capable of using any of the carbohydrates as carbon and energy source. This means that the Rhizobium synthesized enzymes necessary for catabolism of the carbohydrates. It then suggests that the enzyme that degraded each carbohydrate in the bacterium recognized the bonds between monomers in the macromolecules. This is based on mechanisms of digesting similar carbohydatrates by microbes [17].
The Rhizobium species CWP G34B showed considerable (P = ≤ 0.05) degradative ability on the peels of the orange, mango, banana, plantain, pawpaw and pineapple used in the experiment. This showed that the organism was able to attack the pectin derivatives in the peels. This is related to the high fruit-peel-degrading enzyme (FDPE) activity expressed by the Rhizobium in the presence of the fruit peels. This enzyme exhibited its action best in the presence of pectin, meaning that each fruit peel contains one substance or the other that reduced the enzyme activity. It could also be that the amount of reducing sugar units present in the individual peels is lesser than that in the pectin. Indeed, the amount, structure and chemical composition of pectin differs between plants, within a plant over time and in different parts of a plant. The pectin content of some food stuffs are 0.2-1.0% (grapes), 0.5-1.6% (apples), 1.6-4.5% (grape fruit), 3.0-4.0% (lemon), 6.0% (lemon seeds), 32% (lemon rind), 25.0% (lemon pulp), 10.0% (turnip), 30.0% (sugar beet pulp), 20.0% (pineapple/orange peel), 29.0% (pineapple/orange membrane) and 16.0% (pineapple/orange juice sac) 30% (oranges) and 25% (fleshy roots like sugar-beets) [4, 18-20].
The bacterium was able to synthesize protein in the minimal medium containing the carbohydrates. This result indicates that the synthesis of protein by the organism was not repressed by the presence of lactose, maltose, sucrose and mannitol used in this work. The non-deleterious effect of the carbohydrates on the protein synthesis suggests that the production of fruit-peel-degrading enzyme (a protein) will not be significantly affected by these carbohydrates and possibly complexes made up of the disaccharides.
The pectin and fruit peels degradative activities expressed by this Rhizobium species CWP G34B are similar to that reported on many microorganisms including Xanthomonas, Erwinia and Streptomyces. Boboye and Shonukan [10] and Boboye and Shonukan [21] documented that Xanthomonascampestris campestris formed pectate lyase (PL) and the PL synthesis can be genetically controlled. Erwinia species [22, Erwinia aroideae [23], Streptomyces nitrosporeus [8], Erwinia chrysanthemi [9], Bacillus gibsonii S-2 and Clostridium species [24, 25] are known to produce pectin and pectin-derivatives degrading enzymes. Besides bacteria, other microorganisms such as fungi (Penicillium sclerotigenum) [26] and human oral spirochaetes [27] produce pectin degrading enzymes. Many of the microbes reported to produce enzymes capable of degrading pectin and pectin-derivatives are pathogens. These pathogenic microorganisms have been used in the production of industrial pectinases including Clarizyme, Irgazyme, Spark-1 and Ultrazym SE604 for extraction and clarification of fruit juices and grape must, maceration of vegetables and fruits, application in fruit juice and wine technology and Olive oil extraction [4].
The data obtained in this research that Rhizobium species CWP G34B has the ability to degrade fruit peels by synthesizing enzyme/s that catabolize pectin and the peels suggests that the bacterium can be used beneficially for similar purposes as other microbes or organisms. This rhizobium can thus be developed in a process for enzymatic removal of peels from fruits for fruit juice production. Also, it can be applied to control environmental pollution that is caused by piles of fruit peel wastes thrown on the streets or wastes dump site. This is because enzymatic degradation of peels has many benefits including the fact that it is less cumbersome, faster and cheaper at the long run than manual or mechanical means of removing peels. In addition, the merit of using non-pathogenic bacterium can be interesting to develop this organism for safe industrial use, human health and environment application. However, further work is considered to purify and characterize the enzyme.
CONFLICT OF INTEREST
The author(s) confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
We are grateful to Prof. F. C. Adetuyi, Mr. F. C. Akharaiyi and Mrs. F. M. Ilori of the Department of Microbiology and Department of Biochemistry, Federal University of Technology, Akure, Nigeria for provision of materials and instrumentation.
REFERENCES
| [1] | Wikipedia.org [homepage on the internet] Peel (fruit). Available from: http://en.wiki/peel–fruit 2009; 1-7. |
| [2] | Oladiji AT, Yakubu MT, Idoko AS, Adeyemi O, Salawu MO. Studies on the physicochemical properties and fatty acid composition of oil from ripe plantain (Musa parasidiaca) peel Afr Sci 2010; 11: 73-8. |
| [3] | Olukunle JO, Oguntunde PG, Olukunle OF. Development of a system for fresh fruit juice extraction and dispensary In: Proceeding of the Conference on International Agricultural Research for Development; 2007 October 9-11; Tropentag, University of Kassel-Witzenhausen and University of Göttingen, Germany. 2007; pp. 1-4. |
| [4] | Forgatty WM, Kelly CT. Microbial enzymes and biotechnology. London: Enviromnental and Applied Science Publishers 1983; pp. 131-82. |
| [5] | Dutta AC. Botany for degree students. 5th ed. Delhi: Oxford University Press 1981; pp. 106-61. |
| [6] | Capita NC, Gibeaut DM. Structural models of primary cell walls in flowering plants: Consistency of molecular structure with the physical properties of cell walls during growth Plant J 1993; 3: 1-30. |
| [7] | Willats WG, McCartney L, Makey W, Knox JP. Pectin: Cell biology and prospects for functional analysis Plant Mol Biol 2001; 47: 9-27. |
| [8] | Sato M, Kaji A. Purification and properties of pectate lyase produced by Streptomyces nitroporeus Agric Biol Chem 1977; 41: 2193-7. |
| [9] | Hugouvieux-Cotte-Pattat N, Riverchon S, Condemine G, Robert- Boudouy J. Regulatory mutations affecting the synthesis of pectate lyase in Erwinia chrysanthemi J Gen Microbiol 1986; 132: 2099-106. |
| [10] | Boboye BE, Shonukan OO. Regulatory mutations affecting the synthesis of pectate lyase in Xanthomonas campestris World J Microbiol Biotechnol 1993; 9: 240-. |
| [11] | Cleveland TE, Cotty PJ. Invasiveness of Aspergillus flavus isolates in wounded cotton bolls associated with production of specific fungal polygalacturonase Phytopathology 1991; 81: 155. |
| [12] | Ajayi AA, Olutiola PO, Fakunle JB. Studies on polygalacturonase associated with deterioration of tomato fruits (Lycopersicon esculentum Mill) infected by Botryodiplodia theobromae Pathol Sci Focus 2003; 5: 68-77. |
| [13] | Bernfeld P. Amylases α and β In: Colowick SP, Kaplan NO, Eds. Method in enzymology . New York: Academic Press Inc 1955; pp. 149-58. |
| [14] | Boboye BE, Alao A. Effect of mutation on trehalose-catabolicenzyme synthesized by a tropical Rhizobium species FI Res J Microbiol 2008; 3: 269-75. |
| [15] | Gornall AJ, Bardawill CS, David MM. Quantitative determination of protein J Biol Chem 1949; 177: 151. |
| [16] | Duncan DM. Multiple Range and Multiple F-test Biometrics 1955; 11: 1-42. |
| [17] | Wikipedia.org [homepage on the internet] Disaccharides. Available from: http://en.wiki/disaccharides 2010; 14. |
| [18] | Ihekoronye AI, Ngoddy PO. Integrated food science and technology for the tropics. College ed. London & Basingstaoke: Macmillan International Publishers Ltd 1985; p. 386. |
| [19] | Alana A, Alkota I, Dominguez JB, Liama MJ, Serra JL. Pectin lyase activity in a Penicillium italicum strains Appl Environ Microbiol 1990; 56: 3759. |
| [20] | Favelas–Torres E, Aguilar CN, Contreras–Espuivel JC, Viwegra–Gondalez G. Pectinases In: Pandy A, Webb C, Soccol CR, Larroch C, Eds. Enzyme technology. New Delhi, India: Asia Tech. Publishers Inc 2005; p. 26. |
| [21] | Boboye B, Shonukan OO. Isolation purification and some properties of pectate lyase from synthesized by Xanthomonas campestris campestris Biosci Res Commun 2002; 14: 365-9. |
| [22] | Lyon GD, Smith KJ, Heilbronn J. The calcium requirement of polygalacturonic acid lyases produced by Erwinia species Lett Appl Microbiol (Great Br) 1986; 2: 127-9. |
| [23] | Kamimuya S, Itoh , Iziak K, Takahashi H. Purification and properties of pectate lyase in Erwinia aroideae Agric Biol Chem 1977; 41: 975-81. |
| [24] | Li ZM, Bai ZH, Zhang BG, et al. Newly isolated Bacillus gibsonii S-2 capable of using sugar beet pulp for alkaline pectinase production World J Microbiol Biotechnol 2005; 21: 1483-6. |
| [25] | Sharma DC, Satyanarayana T. A marked enhancement in the prduction of a highly alkaline and thermostable pectinase by Bacillus pumilus dcsr1 in submerged fermentation by using statistical methods Bioresour Technol 2006; 97: 727-33. |
| [26] | Olutiola PO. Polygalacturonase and pectate lyase from Penicillium sclerotigenum Niger J Microbiol 1982; 2: 154-7. |
| [27] | Weber FH, Canale-parola E. Pectolytic enzymes of Oral Spirochaetes from human Appl Environ Microbiol 1984; 48(1 ): 61-7. |







