All published articles of this journal are available on ScienceDirect.
Mutants of Saccharomyces cerevisiae and Bacillus citri Changed the Protein Content of the Nigerian Oryza sativa variety “Igbimo” during Fermentation
Abstract
Effect of mutation on protein production by Saccharomyces cerevisiae and Bacillus citri, the best protein producing yeast and bacterium isolated during a previous natural fermentation of a Nigerian rice (“Igbimo”). The two microorganisms were grown to logarithmic phase and mutagenized separately using ethylmethyl sulphonate (EMS). The wild-types and variants were inoculated individually into sterile “Igbimo” rice. Fermentation was allowed to take place at 27°C for 7 days after which protein released into the rice was quantified using the Biuret reagent method. The data obtained showed that the mutants are different from each other. Some mutants did form the protein at lower concentrations, others at the same and higher concentrations than the mother strains. The parental strains of S. cerevisiae and B. citri synthesized 0.89 mg/mL and 0.36 mg/mL protein respectively. Four groups of the mutants are recognized: classes I, II, III and IV which are the Poor, Average, Good and Super Protein Producers with 0-0.20, 0.21-0.50, 0.51-1.0 and 1.0 mg/mL protein respectively The yeast mutants produced higher amounts of protein than those of the bacterium.
INTRODUCTION
Rice, the seed of the monocot plant Oryza, is the second highest worldwide producing grain after maize [1]. It has two domesticated species namely: O. sativa and O. glaberrima. The O. sativa is most common and often cultivated in Africa, America, Australia, China, New Guinea and South Asia [2]. A small amount of the O. glaberrima is grown in Africa [3]. The most important rice producing countries are China, India and Indonesia [4]. In many smaller countries, rice is the leading food crop. In Nigeria, rice is grown on 1.77 Millions hectares ranking sixth after sorghum, millet, cowpea, cassava and yam cultivations [5]. Rice is cultivated in virtually all the agro-ecological zones of Nigeria [6].
Rice is the most important staple food for a half of the world’s population especially in the East, South, South-east, Middle East, Latin America and West Indies [1]. It is the staple food for many homes in Nigeria and ranks first socially being a festive meal generally in the country [5]. Since the mid-1970s, rice consumption has increased tremendously (≥10.3% per annum) because of accelerating growth population rate (+2.8% per annum) and rising per capita consumption (+7.3% per annum) due to an increase in the consumer’s preference for rice [7]. Besides direct consumption of the grain, rice is utilized for the production of laundry starch, wine, beer, cakes, snack foods, breakfast cereals, face powder, polish for camera lenses and expensive jewelleries. The hull and straw are used to make construction materials and production of oil, animal feeds, rope and paper [3]. Rice is used to make starch which is employed in the manufacture of baby foods and extruded noddles. It serves as a glucose substitute in oral dehydration solution used for infants suffering from diarrhoea [8]. Rice consumption is important for the supply of energy. One fifth of the calories intake is provided by rice being rich in carbohydrate [9]. Rice is also rich in vitamins and minerals such as thiamine, niacin, iron, riboflavin, vitamin D and calcium [10]. Rice is poor in protein, sodium and fats, and free of cholesterol which makes it useful in treating hypertension [11-13].
The low content of protein in rice causes nutritional diseases. The nutritional problems prevalent in some countries are protein-energy malnutrition (PEM), nutritional anaemia, vitamin A deficiency and iodine disorders [14-16]. Fermentation of foods is one of the techniques used to improve the nutritional quality of rice. Rice fermentation is commonly practised in the Asian countries where fermentation of food is regarded beneficial. Potentialities of fermented foods are less appreciated in Africa, South and Central America and not valued in the Europe, thus rice fermentation is not well documented in many parts of the world unlike in the Asia [17]. Natural fermentation and the use of starters are employed in rice fermentation [18, 19]. Chemical analysis of a Nigerian rice (‘Igbimo’) naturally fermented showed that there was an improvement in the nutrient contents of the rice [20]. Boboye and Alabi [21] showed that individual microorganisms isolated during the natural fermentation of the rice carried out specific functions in the synthesis and catabolism of nutrients. Saccharomyces cerevisiae and Bacillus citri were identified as the best protein producing yeast and bacterium. As a means of improving the protein forming capability of the yeast and rod bacterium, the organisms were mutagenized using ethylmethyl sulphonate (EMS).
MATERIALS AND METHODOLOGY
Microbial Strains
Saccharomyces cerevisiae and Bacillus citri used in this study were provided by the Department of Microbiology, Federal University of Technology, Akure, Nigeria. They were isolated during the natural fermentation of this rice and have been proven to be the best protein producing microbes among the isolates [21].
Mutation Experiment
Saccharomyces cerevisiae and Bacillus citri were separately grown in 5 mL nutrient broth at 37oC for 18 hours with agitation at 80 rpm. Using ethylmethyl sulphonate (EMS), the culture was mutated at 28oC according to the method of Parkinson [22] modified as described by Boboye and Alao [23]. Mutational rates were estimated. Mutants were screened for the synthesis of protein in the fermentation of rice.
Fermentation of Rice with Mutants and Wild-type Strains
Each variant was inoculated into 5 mL of nutrient broth and incubated at 37oC for 24 hours. The cells were harvested by centrifuging at a force of 168 x 103 g for 15 min (MSE Minor 35 Centrifuge). Cells were washed and resuspended in sterile distilled water. Four grams of the rice were sterilized for 15 min at 121oC and cooled down to 27oC. Four millilitres of the cell load (1.64 at 670 nm) of each mutant and wild-type strain were separately inoculated into different rice sample. Fermentation was carried out for 7 days at 27oC after which protein contents were determined. Based on the amount of total protein synthesized during the fermentation, mutants were categorized thus: 0-0.20 mg/mL: Class I (Poor Protein Producers, PPP); 0.21-0.50 mg/mL: Class II (Average Protein Producers, APP); 0.51-1.0 mg/mL: Class III (Good Protein Producers, GPP); 1.1 mg/mL and above: Class IV (Super Protein Producers SPP).
Quantification of Protein
Protein concentration in every rice sample fermented with each mutant and wild-type was determined by Biuret method [24] as reported by Boboye and Alao [23]. Bovine Serum Albumin (BSA) in a range of 0 to 2.0 mg. Biuret reagent (0.5 mL) was mixed with each BSA concentration at ambient temperature of 28oC for 15 min. This procedure was repeated using 2 mL of each fermented rice sample. Absorbance was read at 540 nm. Standard protein curve was plotted with the values of the optical density and concentrations of the BSA samples. Corresponding protein concentration in each rice sample was read on the standard protein curve using the OD values.
RESULTS
The mutation rate was 87.5% in the Saccharomyces cerevisiae and 69.64% in the Bacillus citri. This shows that the mutation was successful although with more positive impact on the survival of the latter organism. The variants of the yeast and bacterium synthesized protein to varying levels. Relative to the wild-type strains, some mutants synthesized protein at the same, lower and higher levels ranging from 0.05 (Mu 9) to 1.55 mg/mL (Mu 20) for mutated colonies of S. cerevisiae and 0.02 (Mu 18, Mu 69) to 1.16 mg/mL (Mu 22) for the variants of B. citri (Fig. 1). These various mutants constituted classes I, II, III and IV which are the Poor, Average, Good and Super Protein Producers respectively (Tables 1 and 2). Majority (42%) of S. cerevisiae variants belong to the category of Average Protein Producers (class II). Next to this class in decreasing order of population are the yeast variants (28%) that formed low amounts of the protein. The fewest (7%) yeast mutants are Super Protein Producers while 24% variants are good in producing the protein. In contrast, the highest population (48%) of the B. citri mutants fall into class I followed by class II with 42%, 7% in Class III and 3% in the last class. The mother strain of S. cerevisiae formed 0.89 mg/mL protein, meaning that it is naturally a good protein producing microbe unlike the B. citri that made the total protein averagely at 0.36 mg/mL.
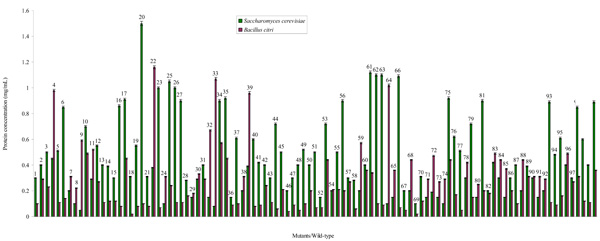
Protein synthesized by mutants and wild-type of Saccharomyces cerevisiae and Bacillus citri during the fermentation of a Nigerian rice (Oryza sativa variety “Igbimo”).
Legend: W: Wild-type strain
Groups of Mutants and Wild-type of Saccharomyces cerevisiae that Synthesized Protein during the Fermentation of a Nigerian Rice (Oryza sativa variety “Igbimo”)
| Class/Identity | Repression/ Induction Ratio | Protein Level (mg/mL) | Mutants |
|---|---|---|---|
| I (Poor Protein Producers, PPP) | ≤0.22 | 0-0.20 | 7, 8, 9, 24, 29, 32, 33, 36, 38, 46, 52, 54, 59, 64, 65, 67, 68, 69, 71, 72, 73, 74, 80, 82, 85, 88, 91, 92. |
| II (Average Protein Producers, APP) | ≤0.56 | 0.21-0.50 | 1, 2, 3, 4, 11, 13, 14, 15, 16, 17, 18, 19, 21, 22, 28, 30, 31, 39, 41, 42, 43, 45, 47, 48, 50, 51, 55, 57, 58, 60, 70, 78, 83, 84, 86, 87, 89, 90, 94, 96, 97, 100. |
| III (Good Protein Producers, GPP) | ≤1.12 | 0.51-1.0 | 5, 6, 10, 12, 23, 26, 27, 34, 35, 37, 40, 44, 49, 53, 56, 75, 76, 77, 79, 81, 93, 95, 98, 99. |
| IV (Super Protein Producers, SPP) | ≤1.74 | 1.1 and above | 20, 25, 61, 62, 63, 66. |
| Wild-type | 0.89 |
Groups of Mutants and Wild-type of Bacillus citri that Synthesized Protein during the Fermentation of a Nigerian Rice (Oryza sativa variety “Igbimo”)
| Class/Identity | Repression/Induction Ratio | Protein Level(mg/mL) | Mutants |
|---|---|---|---|
| I (Poor Protein Producers, PPP) | ≤0.56 | 0-0.20 | 1, 5, 6, 13, 14, 15, 16, 18, 19, 20, 21, 23, 26, 27, 28, 29, 36, 37, 40, 41, 43, 44, 46, 47, 48, 49, 50, 51, 52, 58, 62, 63, 66, 67, 69, 70, 76, 77, 79, 81, 82, 86, 87, 93, 94, 95, 99, 100. |
| II (Average Protein Producers, APP) | ≤1.39 | 0.21-0.50 | 2, 3, 7, 8, 10, 12, 17, 24, 25, 30, 31, 35, 38, 42, 45, 53, 54, 55, 56, 57, 60, 61, 65, 68, 71, 72, 73, 74, 75, 78, 80, 83, 84, 85, 88, 89, 90, 91, 92, 96, 97, 98. |
| III (Good Protein Producers, GPP) | ≤2.78 | 0.51-1.0 | 4, 9, 11, 32, 34, 39, 59. |
| IV (Super Protein Producers, SPP) | ≤3.22 | 1.1 and above | 22, 33, 64. |
| Wild-type | 0.36 |
DISCUSSION
The low levels in protein production of class I mutants means that they are weak in formation of the protein. This weakness was caused by repression in the expression of the DNAs coding for the protein. The repression in the average producers was less pronounced, hence increase in the amounts of protein formed was observed. The good and super protein producers were induced in their genes to synthesize protein similar and more than the mother strains. The genetic modifications are reflected in the repression and induction levels of the mutants (Tables 1 and 2). The ratios below 1.0 indicate that there was no induction in these strains but repression, since production of the same amount of protein as the wild-type resulted to a ratio of 1.0. Above 1.0 ratio indicates induction. The bacterial mutants were better induced than the yeast variants; although the protein concentrations in the latter were generally higher than in the former microorganism. This signifies that the genes encoding proteins in the yeast are naturally better in producing the substance than those in the bacterium.
The variations obtained in the two organisms used in this work are due to the ethylmethyl sulphonate (EMS) added to their genomes. Ethylmethyl sulphonate is an alkylating mutagen known to effect mutation through the addition of methyl to nucleotide [25, 26]. When the CH3 is added to a pool of genes, there is structural change in the base (guanine) to mispair with thymine; this causes mRNA transcribed from the DNA to carry altered message [27], resulting in incomplete synthesis of polypeptide chain in the protein, hence the lower protein concentrations observed in some mutants. Insertion of the methyl resulted to increased protein production in some mutants because their genes were altered to form more polypeptides. The repressor protein was no longer produced or the operator regions had been altered so that they no longer responded adequately to the action of the repressor protein [28].
This research has shown that there is genetic basis of synthesizing total protein by Saccharomyces cerevisiae and Bacillus citri in rice during fermentation and that the level of the anabolism of this substance can be controlled. In Nigeria, rice is not fermented for consumption and when natural fermentation sets into left-over cooked rice, it is considered spoilt and thrown away. In order to employ nutrient enrichment of fermentation in rice consumption in Nigeria, the indigenous rice variety “Igbimo” should be fermented using the genetically improved strain of Saccharomyces cerevisiae. However, further studies to identify and up- or down-regulate the gene/s concerned will be considered in the next aspect of this research.
CONFLICT OF INTEREST
The author(s) confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENT
The authors appreciate Federal University of Technology Akure, Nigeria for provision of equipments, materials and reagents used for this project. The assistance of the Mr. F. Akharaiyi to operate the equipments is acknowledged. We are indebted to late Prof. O. O. Shonukan for provision of ethylmethyl sulphonate.


