All published articles of this journal are available on ScienceDirect.
Subpathotypes of Avian Pathogenic Escherichia coli (APEC) Exist as Defined by their Syndromes and Virulence Traits
Abstract
Avian pathogenic Escherichia coli (APEC) strains cause different types of systemic extraintestinal infections in poultry, collectively termed colibacillosis, which can cause significant economic losses in the poultry industry. To date, there have been no descriptions of genes or characteristics that allow for the classification of avian strains pathotypes responsible for causing specific diseases in their hosts. In this study we aimed to characterize avian E. coli strains representing 4 groups, including one of commensal strains (AFEC – Avian Fecal Escherichia coli) and 3 groups of APEC strains, where each group is responsible for causing a different disease syndrome in their respective hosts (septicemia, omphalitis and swollen head syndrome). We chose to examine several biological characteristics of these strains including: adhesion to eukaryotic cells, pathogenicity levels according to the lethal dose (50%) assay, phylogenetic group and virulence gene profiles. The comparison of strains based on these genotypic and phenotypic traits, using multivariate statisticals tools and complex networks, allowed us to infer information about the population structure of the studied groups. Our results indicate that APEC strains do not constitute a unique homogeneous group, but rather a structured set of subgroups, where each one is associated with a specific infectious syndrome which can possibly be used to define pathotypes or subpathotypes within APEC strains. These results offer new possibilities with which to study the genes responsible for various pathogenetic processes within APEC strains, and for vaccine development. It may be important to consider these subgroups when developing a vaccine in an effort for obtain cross protection, which has not yet been successfully accomplished when working with APEC strains.
INTRODUCTION
Escherichia coli are probably one of the most ubiquitous microorganisms on Earth, existing as part of the commensal intestinal flora, where it is responsible for the production of many beneficial metabolic products. It is also capable of causing different disease syndromes in animal and human hosts [1]. Avian pathogenic Escherichia coli (APEC) [2] strains cause many different types of extraintestinal systemic infections in poultry, which are collectively termed colibacillosis [3]. These diseases, including septicemia, omphalitis, cellulitis, yolk-sac infection and swollen head syndrome, are responsible for high morbidity, mortality and carcass condemnation, all of which cause significant annual economic losses within the avian industry [3]. For many years it has been widely accepted that APEC strains are opportunistic pathogens [4]. Recent studies, however, have revealed significant differences in the distribution of virulence factors among APEC strains and avian fecal commensal E. coli (AFEC) strains, suggesting that APEC strains are well adapted to a pathogenic lifestyle [5, 6]. It is reasonable to expect that APEC would utilize virulence mechanisms similar to that of other extraintestinal pathogenic E. coli (ExPEC) [7] strains, such as uropathogenic E. coli (UPEC) and neonatal meningitis E. coli (NMEC), both human diseases, because they are capable of causing disease in similar environments. A comparative analysis of the first complete APEC genome sequence [8] and genomes from various Ex-PEC subpathotypes [5] have revealed strong similarity between these strains of E. coli. If all APEC strains cause a common disease in poultry with a common set of virulence factors they may constitute a specific pathotype [1, 6]. We hypothesize that APEC strains like human ExPEC strains, which encompasses the UPEC and NMEC strains that cause distinct syndromes, may harbor subtypes with unique genotypes that cause distinct disease syndromes in poultry. To our knowledge, no effort has been made to determine if distinct subpathotypes of APEC strains exist, or if specific traits determine the root causes of pathogenesis in these distinct syndromes. APEC pathogenesis is still poorly understood in general, when compared to that of human pathogenic E. coli [4, 6]. Despite recent advances in our understanding of APEC pathotypes, a method that allows for the differentiation of APEC from AFEC has not yet been developed, partially due to the diversity of APEC populations [5, 6] and the likelihood that some are commensal strains with known virulence characteristics. In this study, we investigated the possibility that if APEC strains isolated from poultry showing specific disease symptom, such as septicemia (S), omphalitis (O), and swollen head syndrome (H), are actually member of distinct subpathotypes, which possess different virulence traits and capacities to cause disease. For the purposes of comparison, AFEC strains (N), isolated from the cloaca of presumably healthy chickens were also studied. For the first time our results indicate, that at least some of these strains can potentially be grouped into subpathotypes responsible for specific infectious dieases in poultry.
MATERIAL AND METHODS
Bacterial Strains
Escherichia coli strains were isolated from commercial adult chickens from different regions of the São Paulo State in Brazil, which had clinical signs of septicemia (S: n = 24) and swollen head syndrome (H: n = 14), as well as being from one-day-old chicks with omphalitis (O: n = 11). Thirty AFEC strains (N) were isolated with cloacal swabs from chickens with no signs of illness. All strains belong to the Microbial Molecular Biology Laboratory of the Department of Genetics, Evolution and Bioagents, at Campinas State University (UNICAMP). The strains from septicemic cases were isolated from the liver, air sacs and lungs. Strains that caused Swollen Head Syndrome were isolated from the infraorbital sinuses of affected birds, and strains that caused omphalitis were isolated from the yolk sacs of one-day-old chicks. MacConkey agar medium was used for the primary isolation of these strains and further confirmation that these strains were E. coli was accomplished using the Enterobacteria test kit from Laborclin. For routine tests, Luria Bertani broth (LB), or agar (LBA) [8], was used. All isolates were stored at -70 °C in LB containing 30% glycerol.
DNA Extraction
DNA extraction was performed with the Wizard Genomic DNA Purification Kit (Promega, EUA), according to manufacturer's instructions. To confirm the presence of a number of plasmids, the alkaline extraction technique described previously by Sambrook et al. [8] was used.
Detection of Virulence Genes
Virulence genes were detected using Polymerase Chain Reaction (PCR), following protocols described elsewhere (Table 1).
Primers used in PCR Assays. Their Descriptions, Functions, Annealing Temperatures, Expected Amplicon Fragment Sizes and References
| Gene | Function | Annealing Temp. (°C) | Amplicon Size (bp) | References |
|---|---|---|---|---|
| yjaA | Unknown function gene | 55 | 211 | [2] |
| tspE4.C2 | DNA fragment | 55 | 152 | [2] |
| iucA | Aerobactin synthetase | 55 | 1100 | [21] |
| irp-2 | Iron- controlling gene associated to yersiniabactin | 59 | 287 | [22] |
| fep C | Iron transport enterobactin | 55 | 347 | [23] |
| sitA | Iron transport | 59 | 608 | [25] |
| fyuA | Yersiniabactin | 63 | 787 | [24] |
| fimA | Structural subunit of type 1 fimbria | 50 | 331 | [25] |
| papA | Structural protein of P fimbria | 65 | 328 | [26] |
| crl | Regulatory gene of “curli” fimbria | 54 | 250 | [27] |
| csgA | Structural protein of “curli” fimbria | 55 | 200 | [27] |
| tsh | Temperature-sensitive hemagglutinin | 61 | 420 | [27] |
| lpfAO141 | LPF fimbria | 55 | 412 | [28] |
| lpfAO154 | LPF fimbria | 55 | 525 | [29] |
| iha | Adhesion protein | 55 | 602 | [30] |
Strains Positive for the Presence of Virulence Genes as Determined by PCR. These Values are Presented as Percentages
| Virulence Genes | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Group | fimA | papA | crl | csgA | tsh | lpfAO141 | lpfAO154 | iha | fyuA | irp-2 | iucA | fepC | sitA |
| N | 16.7 | 0.0 | 10.0 | 0.0 | 10.0 | 3.3 | 20.0 | 13.3 | 13.3 | 13.3 | 6.7 | 26.7 | 26.7 |
| S | 29.2 | 25.0 | 29.2 | 87.5 | 25.0 | 25.0 | 58.3 | 20.8 | 16.7 | 20.8 | 45.8 | 29.2 | 12.5 |
| H | 64.3 | 35.7 | 50.0 | 64.3 | 50.0 | 0.0 | 100.0 | 0.0 | 57.1 | 71.4 | 42.9 | 92.9 | 28.6 |
| O | 27.3 | 0.0 | 36.4 | 54.5 | 0.0 | 0.0 | 81.8 | 18.2 | 36.4 | 36.4 | 0.0 | 45.5 | 18.2 |
N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
Strains Positive for Adhesion to Eukaryotic Cells. These Values are Presented as Percentages
| Adhesion Assays | ||||||
|---|---|---|---|---|---|---|
| Group | TraqNM | TraqM | HeLaNM | HeLaM | HEp-2NM | HEp-2M |
| N | 66.7 | 33.3 | 53.3 | 20.0 | 93.3 | 63.3 |
| S | 45.8 | 33.3 | 70.8 | 54.2 | 58.3 | 41.7 |
| H | 71.4 | 42.9 | 92.9 | 92.9 | 92.9 | 78.6 |
| O | 36.4 | 9.1 | 27.3 | 27.3 | 18.2 | 18.2 |
N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains. TraqM: adhesion to the trachea in the presence of D-mannose; TraqNM: adhesion to the trachea in the absence of D-mannose; HeLaM: adhesion to HeLa cells in the presence of D- mannose; HeLaNM: adhesion to HeLa cells in the presence of D- mannose; HEp-2M: adhesion to HEp-2 cells in the presence D- mannose; HEp-2NM: adhesion to HEp-2 cells in the absence of D- mannose.
Number of Strains Associated with Each Phylogenetic Type. These Values are Presented as Percentages
| Phylogenetic GROUPS | ||||||
|---|---|---|---|---|---|---|
| Group | A | D | B1 | B2 | P (B2 + D) | NP (A + B1) |
| N | 83.3 | 0.0 | 13.3 | 3.3 | 3.3 | 96.7 |
| S | 45.8 | 45.8 | 8.3 | 0.0 | 45.8 | 54.2 |
| H | 21.4 | 64.3 | 0.0 | 14.3 | 78.6 | 21.4 |
| O | 72.7 | 18.2 | 9.1 | 0.0 | 18.2 | 81.8 |
N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
Number of Strains Associated with Each 50% Lethal Dose Interval. These Values Presented are as Percentages
| LD50 Assay | |||
|---|---|---|---|
| Group | LD50 > 1010 | LD50 = 109 - 107 | LD50 = 106 - 105 |
| N | 100.0 | 0.0 | 0.0 |
| S | 41.7 | 45.8 | 12.5 |
| H | 0.0 | 100.0 | 0.0 |
| O | 100.0 | 0.0 | 0.0 |
N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
K-means Calculated Composition of two and three Clusters with all Variables Included
| K-means for Two Groups | K-means for Three Groups | ||||
|---|---|---|---|---|---|
| Cluster 1 | Cluster 2 | Cluster 1 | Cluster 2 | Cluster 3 | |
| N | 0% | 100% | 90% | 0% | 10% |
| S | 54.17% | 45.83% | 33.33% | 33.33% | 33.33% |
| H | 100% | 0% | 0% | 92.85% | 7.14% |
| O | 0% | 100% | 18.18% | 0% | 81.81% |
N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
Classification Matrix using a DA Model to Discriminate Between g Groups N, S, H and O
| Correct assignment (%) | N | S | H | O | |
|---|---|---|---|---|---|
| N | 100.00 | 30 | 0 | 0 | 0 |
| S | 70.83 | 2 | 17 | 2 | 3 |
| H | 100.00 | 0 | 0 | 14 | 0 |
| O | 90.91 | 1 | 0 | 0 | 10 |
| Total | 89.87 | 33 | 17 | 16 | 13 |
Rows: Observed classifications. Columns: Predicted classifications. N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
Classification Matrix using the DA Model to Differentiate AFEC and APEC Strains
| Correct Assignment (%) | AFEC | APEC | |
|---|---|---|---|
| AFEC | 96.67 | 29 | 1 |
| APEC | 95.92 | 2 | 47 |
| Total | 96.20 | 31 | 48 |
Rows: Observed classifications, Columns: Predicted classifications.
Phylogenetic Analysis
Isolates were assigned to phylogenetic groups according to the method of Clermont et al. [9]. In this method, isolates are assigned to one of four groups (A, B1, B2, or D) based on the presence of two genes (chuA and yjaA) and a specific DNA fragment (TSPE4.C2).
Adhesion to Tracheal Epithelial Cells
The ability to adhere to tracheal epithelial cells was determined as described by Dho-Moulin and Lafont [10]. Briefly, tracheas were aseptically removed from 18-day-old specific-pathogen-free, embryonated eggs, rinsed in phosphate buffered saline (PBS, pH 7.4) and cut in 5-mm sections. Adherence studies were performed in 96-well-round-bottom microtiter plates containing two tracheal rings, 25 mL of Eagle’s medium with 5% calf serum, and a bacterial suspension (109 cells/mL) which was previously grown in LB (37°C, 18 h). The plates were incubated at 37°C for 30 min and were washed twice with PBS. Fresh Eagle’s medium was then added, and the plates were incubated for 4 h at 37°C. After incubation the tracheal sections were rinsed with 0.1M PBS containing 10% formalin. The trachea rings were dehydrated, treated with xyline and embedded in paraffin. Five-micrometer sections were removed using a microtome, mounted on glass slides, hydrated and stained with Giemsa stain. The adherence assays were performed in the presence and in the absence of 1% D-mannose.
Adhesion to Epithelial Cells Cultured In Vitro
The ability of strains to adhere to HEp-2 and HeLa cells was determined according to a technique described previously by Scaletsky et al. [11], with slight modifications. Cultures of HEp-2 and HeLa cells were grown in 24-well tissue culture microplates (BD Falcon, Bedford, MA, USA) into which sterile round cover slips (13mm in diameter) had been placed prior to inoculation with the cells. The growth medium in each well of the microplate consisted of 0.9mL of Eagle’s minimal essential medium (MEM) supplemented with 10% fetal calf serum, either with or without 1% D-Mannose, and an 1% antibiotic solution (penicillin 100,000 U and streptomycin 100 μg/ml). Microplates were incubated in 10% CO2 at 37°C until a semi-confluent monolayer was formed. Afterwards, the monolayers were washed 3 times with sterile PBS (0.05M; pH 7.2) and 0.1mL aliquots of bacterial test cultures [(grown for 18 h at 37°C, and diluted 1:10 in PBS (approximately 107 CFU)] were added to the wells. After 3 h of incubation at 37°C, monolayers were washed 10 times with PBS buffer, fixed with methanol for 10 min, stained with the May-Grunwald and Giemsa stains, and observed under bright field microscopy (1000x magnification).
Fifty % Lethal Dose (LD50) Assay
Pathogenicity tests for all strains were carried out as described by Silveira et al. [12]. Briefly, 0.5 mL from various 10-fold dilutions of bacterial culture; grown overnight at 37°C in LB medium, followed by 1:100 dilution in the same medium and growth at 37°C for 4 h, with shaking at 150 rpm; were injected subcutaneously into the neck of groups six one-day-old male chicks. At the same time, these dilutions were plated on LBA to determine the number of colony forming units (CFU) present in each injection. Controls chicks were injected with LB medium. The injected groups and controls were observed for 7 days and the LD50 was calculated by the method described by Reed and Muench [13]. Bacterial strains were grouped into three categories, accordingly to 50% Lethal Dose value: LD50>1010, 109 > LD50< 107 and 106 > LD50< 105.
Statistical Analysis
To analyse binary variables, all data were organized in a binary incidence matrix, wich indicated the presence or absence of the traits tested in each isolate examined. These traits included the 13 virulence genes, fimA, papA, crl, csgA, tsh, lpfAO141, lpfAO154, iha, fyuA, irp-2, iucA, fepC, sitA; and 11 phenotypic characteristics. The phenotypic characteristics included the following adhesion properties: adhesion to the trachea in the presence (TraqM) or absence (TraqNM) of D-mannose; adhesion to HeLa cells in the presence (HeLaM) and absence (HeLaNM) of D- mannose, and adhesion to HEp-2 cells in the presence (HEp-2M) and absence (HEp-2NM) of D- mannose. Phylogenetic groups were also included in the matrix as phenotypic traits. Phylogenetic groups B2 and D were considered a single category (B2+D), because virulent extra-intestinal strains mainly belonged to groups B2 and D. Groups A and B1 were considered a separate category (A+B1) [9]. Additionally, three categories of 50% Lethal Dose characteristics, LD50>1010, 109 > LD50< 107 and 106 > LD50< 105 were included in the matrix as phenotypic data. The frequencies of these traits were compared among the groups (N, S, H and O) using the G-test.
The structure of the data was visualized by a Correspondence Analysis (CA), performed with the frequency matrix for the presence or absence of the genes and phenotypic characteristics in each group (N, S, H and O). CA is an exploratory technique, which allowed us to bidimensionally visualize the structure of categorical variables on Chi-squared values. To accomplish this, the total variance (total inertia) of the data is decomposed arriving at a lower-dimensional representation of the variables that allows for the reconstruction of most of the variance matrix variables. The distance between the points in the bidimensional representation indicates the association between them, the closer they are, the most associated they are with each one [14].
Principal component analysis (PCA) is a factor analysis that reduces the number of variables and classifies variables according to their relationship. PCA uses an orthogonal transformation to convert a set of possibly correlated variables into a set of values with uncorrelated variables known as principal components. In this manner, PCA is commonly used to make predictive models which are useful for classification of variables [14]. A PCA model was constucted to obtain the best separation among the groups, using all 24 binary variables.
K-means is a clustering tool that produces different k clusters of the greatest possible distinction. The best number of clusters k that lead to the greatest separation is not known as a priori, and must be computed from the data [14]. The algorithm K-means was used to verify the clustering of the 79 strains according the traits analyzed. K-means was performed for 2 and 3 clusters using all of the binary variables.
Discriminant function analysis (DA) is used to determine what variables can be used to differentiate between two or more groups, and is determinated a priori [14]. A DA was performed to determine what variables discriminate among the groups N, S, H and O, among clusters 1, 2 and 3 obtained by the K-means algorithm, as well as between the APEC and AFEC groups. All the statistical analyses were performed using STATISCA 6.0 (Statsoft).
Complex Network
In this network, there are two sets of nodes: one is representing the samples and another representing the traits. Each node representing a sample is connected to all nodes that represent the traits specific to it. We used an approach derived from the physics of complex networks to identify the modules in the network. This module identification algorithm searches for partitions in the empirical network, in which modules maximize a modularity function [15]. This function accounts for the difference between the fraction of links inside each module, and the fraction of links that are expected to be assoicated with the modules based on chance [16]. The algorithm uses the simulated annealing (SA) method, a stochastic optimization technique that retrieves the lowest cost partition, and thus the optimal configuration [17]. Many methods are available for cluster identification, such as hierarchical clustering or principal component analysis; however, this maximization algorithm provides a single solution with well-defined boundaries that can be used to separate groups [16]. The SA algorithm also calculates the significance of the modularity degree by comparing it with that of similarly-sized random networks. The SA analysis was run using 100 random networks to examine whether the empirical network was significantly more modular than the random ones. The number of fixed randomizations was low because the simulated annealing algorithm is time-consuming, and 100 randomizations were enough to yield small confidence intervals (average SD < 0.01).
RESULTS
AFEC and APEC strains responsible for different syndromes were compared on a genetic basis using the presence of the genes fimA, papA, crlA, csgA, tsh, lpfAO141, lpfAO154, iha, fyuA, irp-2, iucA, fepC, and sitA as variables. Phenotypic characteristics were also used to Campare AFEC and APEC strains including the ability to adhere to tracheal epithelial cells in presence (TraqM) or absence (TraqNM) of D-mannose, the ability to adhere to HeLa cells cultured in vitro in the presence (HeLaM) or in the absence (HeLaNM) of D-mannose, and the ability to adhere to HEp2 cells cultured in vitro in the presence (HEp2M) or in the absence (HEp2NM) of D-mannose. Phenotypic characteristics were also considered if they were assigned to a particular pathogenic phylogenetic type, (A+B1 and B2+D), and where shown in the thress categories of pathogenicity levels according to the LD50 assay as strains of high virulence (106 > LD50< 105), strains of intermediate virulence (109 > LD50< 107) and strains of low virulence (LD50>1010). The results, per group (N, S, H and O), including the presence of virulence genes, adhesion assays, pathogenic-phylogenetic grouping and the 50% lethal dose assay are shown, respectively, in Tables 2, 3, 4 and 5.
Statistical Analysis
According to the G-Test, the groups N, S, H and O are different in regard to the distribution of the analyzed characteristics. The G-test value was 211.3361 with 63 d.f. and a p<0.001.
A bidimensional plot in Fig. (1) represents the results for the Correspondence Analysis (CA).
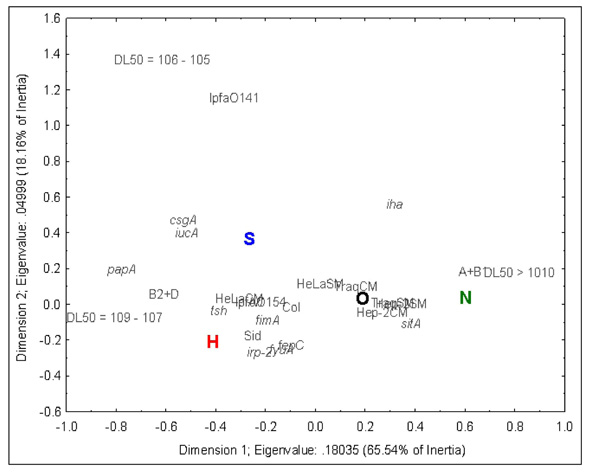
Correspondence Analysis (CA). N = commensal strains (AFEC). S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
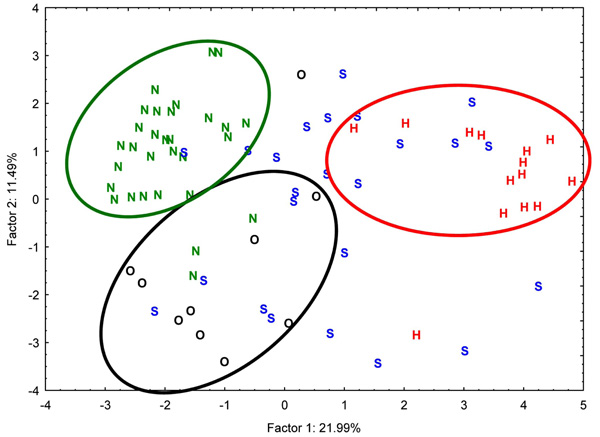
Principal component analysis (PCA). N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.

Principal component analysis (PCA). N = commensal strains (AFEC), S = septicemia strains, H = swollen head syndrome strains, O = omphalitis strains.
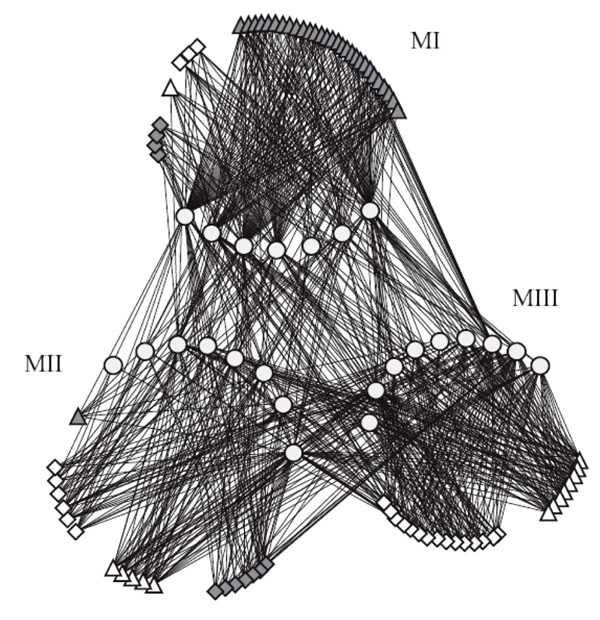
Network representation of the occurrence of traits analyzed in E. coli isolates. Circles represent each trait (see material and methods). Strains associated with different diseases are represented by different combinations of geometric figures and colors as follows: gray triangles = commensal strains (AFEC), gray diamonds = omphalitis strains, blank triangles = swollen head syndrome strains, blank diamonds = septicemia strains. An edge between a strain and a trait indicates that the trait was detected for that strain. Each module identified by the SA algorithm is labeled accordingly (MI, MII, and MIII).
The horizontal axis represents 65.54% of the total inertia and, the vertical axis is representative of 18.16%. Therefore, the horizontal axis is more representative of original data, and the distances between variables in this dimension are more informative. The total Chi-squared value was 209.137 (d.f. = 75, p<0.001), and the total Inertia was 0.30112. According to this analysis, similar traits were distributed between N and O samples, and traits were more dissimilar between H and N, when compared to results obtained from other comparisons.
The bidimensional plot of the PCA data is shown in Fig. (2).
The PCA model contained 22 individual factors. Factor 1, corresponding to the horizontal axis was responsible for 21.99% of the total inertia, and separated the swollen head syndrome-causing strains (H) from AFEC strains and omphalitis-causing strains (N and O), as shown in Fig. (2). Phylogenetic type and LD50 (both related to pathogenicity) were the most important traits in this factor comparison; they are the principal traits responsible for separating group H from groups N and O. Factor 2, corresponding to the vertical axis, encompassed 11.49% of the total inertia and separated the AFEC strains from the O strains. The adhesion assays were the most important traits in this factor comparison, because they were the major traits separating the AFEC and O strains.
The composition of the two clusters as calculated with the K-means algorithm, including all variables is presented in Table 6. The characteristics that had higher variability between groups (higher than the variance within the group) were LD50 >1010 and 109 > LD50 < 107. The composition of the three clusters obtained using the K-means algorithm with all variables is also presented in Table 6. The variables with the highest between-group variance (greater than the variance within an individual group) were LD50 >1010, 109 > LD50 < 107 and HEp-2 SM. These data corroborate the results obtained by PCA, and indicate that the pathogenicity and adhesion profiles are the major factor shaping the structure of this population.
The separation of the three clusters obtained by the K-means algorithm appeared to be the best way to look at the samples and data studied, because it was capable of distinguishing betwenn the H, O and N strains. Cluster 1, which was made up almost entirely of AFEC strains (N), was characterized by low pathogenicity, low prevalence of virulence factors, a predominance of phylogenetic groups A and B1, and many strains with adhesion capacity, especially to HEp-2 cells. Cluster 2, which was mostly comprised of strains causing swollen head syndrome (H), was characterized by intermediate pathogenicity, a high frequency of virulence factors, the predominance of phylogenetic groups B2 and D, many strains with general adhesion capacity, especially to HeLa cells in the absence of mannose. Cluster 3 was mostly comprised of omphalitis strains (O), and was characterized by low pathogenicity, a low frequency of virulence factors (with the exception of the lpfAO154 gene), a predominance of phylogenetic groups A and B1, a few strains with general adhesion capacity and the complete absence of adhesion to HEp-2 cells in the presence of mannose.
The DA, using all variables that allowed the discrimination of isolates from the N, S, H and O groups had a correct assignment rate of 89.87% (Table 7).
The variables used in the DA model to separate the isolates based on the characteristics of groups AFEC and APEC (formed by S, H and O strains) were csgA, lpfAO154, crl, sitA and fimA genes, groups A+B1,, HeLa CM, HEp-2 SM, TraqCM, LD50%>1010 and 106>LD50%<105 (Wilks' Lambda: 0.23952, F (11,67) = 19.338 p< 0.00001). These variables were used to correctly classify 96% of the isolates (Table 8).
Since the diversity was higher between isolates from group S, they were often incorrectly assigned (Table 4). A PCA was also performed using only isolates from groups N, H and O (Fig. 3).
Network Analysis
The modularity index M was significantly higher than that of similarly-sized random networks (P < 0.001). The SA algorithm identified three modules of similar size (Fig. 4). The first module, MI, was composed primarily of AFEC isolates. The second module, MII, contained more than 60% of the total omphalitis isolates, and septicemia isolates, but less than 40% of the total swollen head syndrome isolates, and just one AFEC isolate.The last module, MIII, was composed of many septicemic isolates, and more than 60% of the total swollen head syndrome isolates.
DISCUSSION
In this study, we aimed to test the hypothesis that APEC strains, isolated as causative agents of various syndromes, harbor different capacities to cause disease and can be grouped into distinct subpathotypes. We hypothesized that these isolates could be distinguished from one another by certain genotypic and phenotypic traits. The possibility that APEC strains could be categorized into subgroups has been suggested in previous studies [e.g., 18;6], based on data obtained by virulence factor-genotyping, but the relationship between these subgroups and the various disease syndromes caused by APEC strains has remained understudied.
Our results suggest such a relationship exists, and we have revealed the existence of at least two subgroups within APEC strains: one composed mainly of strains that cause swollen head syndrome and another composed mainly of strains that cause omphalitis. The septicemic strains appear to be dispersed and between these groups. These results also suggest that the difficulty in establishing an exact method with which to distinguish the AFEC and APEC strains may be partially due to the existence of structured subgroups within the APEC strains, including some of the more similar AFEC strains, such as the omphalitis-related strains. Thus, a resolution that allows differentiation between the extreme groups (groups more dissimilar from the AFEC strains, such as the strains that cause Swollen Head Syndrome) and the AFEC strains does not allow differentiation between more similar groups (again compared to AFEC strains), such as the omphalitis-causing and AFEC strains. Consequently, some APEC strains may be incorrectly classified as AFEC strains, and vice-versa [5, 6], whenever a unique cut-off parameter is established for differentiation, except if the factors involved in APEC pathogenesis are fully distinct, and used to develop cut-off parameters.
Other published data corroborate our studies. Rodriguez-Siek et al. [6] found a significant difference in the distribution of genes associated with virulence between strains causing septicemia and cellulitis, which suggests the presence of two distinct strains, each associated with a specific disease. Monkady et al. [19], when comparing their results to those obtained by Stocki et al. [20], observed that two strains belonging to the same serogroup (O2), one associated with septicemia and the other with cellulitis, and having different numbers of DNA sequences associated with virulence, which suggested that the strains associated with septicemia have acquired a larger array of virulence factors. It therefore appears that differences among strains that cause different disease exist, and these differences include variation in the number of virulence genes and phenotypic traits, such as adhesion and lethality, as demonstrated in this work.
The strains that cause septicemia, which do not constitute a specific group, overlap with the H and O isolates, and only slightly with the AFEC strains, which is consistent with there being a distinct difference between the AFEC and APEC strains [5]. This evidence also corroborates the theory that APEC strains are well equipped for a pathogenic lifestyle, and are not simply opportunistic pathogens [5]. Besides being important to our understanding of the APEC pathotypes, the dispersal of strains causing septicemia within other groups is important, because septicemic strains are often chosen as ‘model’ strains in APEC studies that aim to better define the pathogenic mechanisms used by APEC to cause colibacillosis. The apparent diversity of strains that cause septicemia may reflect a failure to identify a specific set of virulence factors among these strains. If there is a yet unknown set of properties that would allow for the better characterization of septicemic strains, its discovery may allow for this distinction in the future
Alternatively, it may be that septicemia is a result of the progression of primary omphalitis or swollen head syndrome. If these were the case, septicemic strains would not be expected to form a unique homogenous group, but would be comprised of a heterogenous mix of strains, varying in genotype and the number of virulence factors, as well as their capacity to cause disease and colonize different tissues. In humans, more than one subpathotype of ExPEC, including strains of UPEC and NMEC, can cause septicemia [7].
The multivariate statistical analyses used in this study did not determine the number of groups within the population, which was deduced by data interpretation, based on samples similarities. However, the complex network analysis revealed a structure based on three modules, indicating that the APEC population is structured, since one module is comprised almost entirely of AFEC strains. If this was not the case, only two modules would appear in the network, one comprise of the AFEC strains, ant the other comprised of the APEC strains. The appearance of three modules, one comprised of the AFEC strains and two other of the APEC strains, indicates that there are at least two structured APEC groups. Our data suggest that one group represents the APEC strains that cause swollen head syndrome; the other represents the strains that cause omphalitis. The strains that cause septicemia are distributed between the two groups.
The apparent differences between the results obtained from the multivariate statistical analyses, and the results obtained from the complex network analysis, may be explained by the fact that the network links each strain to a module. Therefore, differences between strains linked to the same module may not necessarily be apparent, and can become understated. The multivariate statistical analyses, which did not reveal the presence of distinct groups, allowed us to visualize that strains causing omphalitis are more geneticall and phenotypically related to strains than strains that cause swollen head syndrome.
As revealed by PCA, the phylogenetic groups and DL50 (both of which are related to pathogenicity) are the most important traits responsible for separating strains that cause swollen head syndrome (H) from AFEC strains and omphalitis-causing strains (N and O). These findings suggest that H strains are more pathogenic and that they have “aggressive” characteristics, including high lethality, the presence of virulence genes and a general ability to adhere to different cell types.
Additionally, it is important to note that the traits primarily responsible for the grouping of AFEC strains and omphalitis-causing strains were that of lethal dose categories and phylogenetic groups. The specific characteristics that distinguished them were the performance in adhesion assays and lpfAO154 gene. Thus, omphalitis-causing strains may be not opportunistic pathogens, but isolates of low pathogenicity that are particularly well adapted to a specific niche, resulting in their similarities to AFEC strains, but with a different adhesion profile. This observation is consistent with the fact that these strains may have a particular infection pathway, causing direct transmission from the cloacae through the egg shell.
Our results corroborate those of Johnson et al. [5] who found that APEC strains were not homogeneous, and could be assigned to subgroups based on differences in virulence gene distribution. In their work, strains were not analyzed by the disease they caused, or by the site of origin. However, they detected two groups, one harboring strains with low pathogenicity that had few known virluence factors, and another harboring strains with high pathogenicity that had a high number of known virulence genes. Interestingly, in that study, many APEC strains were misclassified as AFEC strains because they possessed a low number of virulence genes, prompting the authors to suggest that these were opportunistic pathogens. It is possible to hypothesize that such characterization reflects the population structure observed here, with these differences reflecting their virulence profiles based on the syndrome they cause.
Our results enhance the premise that avian hosts, in the same manner as their human counterparts, are biologically complex and provide different niches for pathogens to occupy. Because there are many pathotypes of E. coli associated with disease in humans [1], the same would be expected for avian pathogenic strains.
The work reported here has examined Brazilian APEC strains, but future work is required to determine if our observations can be repeated when analyzing non-Brazilian APEC strains. If there are distinct APEC subpathotypes, it may be important to consider when developing vaccines, because new vaccines may vary in ther ability to prevent infections caused by different pathotypes.
ACKNOWLEDGEMENTS
The authors thank Fundação de Amparo à Pesquisa do Estado de São Paulo - Fapesp (Grant number 08/56739-6)


