All published articles of this journal are available on ScienceDirect.
The Complete Genome Sequence of Bacillus safensis BRM1 Isolated from Brazilian Mangrove Sediment: A Potential Source of Biomass Converting Enzymes
Abstract
Background:
Bacillus safensis BRM1 was isolated from Brazilian mangrove sediment and selected for its ability to grow in xylan as the sole carbon source. To identify genes encoding biomass conversion enzymes, the genome of this bacterium was sequenced.
Methods:
Genome wide analysis revealed 99% nucleotide identity to the Bacillus safensis genome. The isolated strain was named B. safensis BRM1, and its genome consists of a circular chromosome of 3.74 Mb with a GC content of 41.8%. Genes encoding a plethora of hydrolytic enzymes are present in the BRM1 genome but absent from the other B. safensis genomes.
Results:
A total of 23 genes encoding putative cellulases or hemicellulases were identified.
Conclusion:
These data support that B. safensis BRM1 is an interesting candidate for the prospection of enzymes that can be applied in the conversion of cellulosic biomass to biofuel.
1. INTRODUCTION
The genus Bacillus is comprised of aerobic gram-positive, rod-shaped bacteria. Species of this genus can be found in many different habitats, such as soil, water, food, diseased animals (B. anthracis or B. larvae), hot springs, and plants [1]. These bacteria have wide biotechnological applications, being the main source of a variety of hydrolytic enzymes such as amylases and proteases which are used in the industry worldwide [2]. Many Bacillus strains are able to produce cellulases and hemicellulases that may be applied in biofuel production; some of these enzymes showed particular interesting properties such as a wide range of pH and temperature stability [3-5]. Amongst the species of the genus, Bacillus safensis has demonstrated the potential of producing enzymes with different biotechnology applications, including inulinases, laccases, galactosidases, oxidoreductases, and keratinases [6-10].
Searching for new enzymes with biotechnology potential, we collected a sample of a Brazilian mangrove, located at Paranaguá bay, Paraná (25°29'S 48°27'W). The microorganisms from the sample were isolated after growth in a selective medium, and a strain capable of growing on xylan as the sole carbon source, and in LB (Luria-Bertani) containing 5% NaCl (m/v) was isolated and named BRM1 (BRazilian Mangrove). In order to identify genes encoding biomass conversion enzymes, the genome of this strain was sequenced.
2. MATERIALS AND METHODS
2.1. Sample Extraction and Strain Selection
Samples were extracted from the sediments of mangroves at Paranaguá Bay, Paraná, Brazil, during the summer season (March 14th, 2014), near Antonina city. The upper layer of sediment (around 10 cm) was discarded and around 100 grams of the surfaced material was aseptically collected and stored in ice until processing.
At the laboratory, 1 g of the sediment sample was resuspended in 5 mL of 5% NaCl. An aliquot of 200µL of the serial dilutions at 1:25 and 1:125 was spread on an agar plate containing a selective medium for cellulases or xylanases producing microorganisms. The selective medium used had the following composition: 1.6g/L of K2HPO4, 0.2g/L of KH2PO4, 0.2 g/L of MgSO4.7H2O, 0.01g/L of FeSO4.7H2O, 0.2 g/L of yeast extract, 0.2 g/L casamino acids, 1.2x10-2 g/L of MnSO4.H2O, 0.02 g/L of CaCl2.2H2O, 1.4x10-2 g/L of H3BO3, 4.0x10-4 g/L of CuSO4.5H2O, 1.2x10-3 g/L ZnSO4.7H2O, 0.01 g/L of Na2MoO4.2H2O, 20 g/L of NaCl, 15 g/L of agar, and 5 g/L of xylan or carboxymethyl cellulose (CMC).
After inoculation, the plates were maintained at 30°C for 4 days, and morphologically different colonies were picked, streaked onto a novel selective medium agar plate, and incubated again at 30°C for 4 days. A total of 20 isolates were selected and kept at 4 °C on solid medium or stored in skim milk at –80 °C [11]. One isolate showing higher xylan degrading activity was named BRM1 and selected for further analysis.
2.2. Enzyme Activity Measurements
To detect enzymatic activity on Petri dishes, the solid medium was the same selective medium used during the isolate selection stage. The presence of CMC hydrolytic activity was verified by the formation of hydrolytic halo using the red congo staining [12]. For the xylanase activity, the halo could be identified directly without the need for a staining step.
For enzymatic assay in a liquid medium, the isolate was cultured aerobically in a rotary shaker at 30oC and 120 rpm. The medium composition was the same as described by COMMICHAU and collaborators [13] but diluted 2-fold in water. Different carbon sources (0.5% w/v) were added as indicated: glucose (control), cellobiose, xylan, or carboxymethyl cellulose (CMC). A 300 μL aliquot was removed daily from each cultivation flask and the cell density was monitored by measuring the optical density at 600 nm.
2.3. Genome Sequences
For genome’s comparison, we used 13 complete Bacillus genome sequences available at public database: B. pumilus 145, B. subtilis ATCC 19217, B. safensis KCTC, B. altitudinis P-10, B. pumilus SAFR-032, B. altitudinis SGAir0031, B. pumilus SH-B9, B. safensis U14-5, B. safensis U17-1, B. safensis U41, B. xiamenensis VV3, B. altitudinis W3, and Bacillus sp. WP8 (Table 1).
2.4. Genome Sequencing, Annotation, and Analysis
Genomic DNA was purified using phenol-chloroform extraction of cells lysed with lysozyme and SDS. Genome sequencing of BRM1 was performed using an Illumina MiSeq Sequencer. Library construction was done with Nextera DNA Library Prep Kit, and sequencing generated 3,185,036 end-paired reads (~448 Mb), yielding a 120-fold genome coverage. The data were assembled using CLC Genomics Workbench 6.5.1 [14] and Roche GS De Novo Assembler (Newbler, release 2.8). Contigs were ordered in one scaffold using the jContigSort program [15] and the gaps were closed with the FGAP program [16] and in house scripts.
| Accessiona | Organism | Isolation Source | Size (Mb) | GC (%)b | Identity (%)c |
|---|---|---|---|---|---|
| CP024204.1 | B. altitudinis P-10 | Rhizosphere of rice | 3.76 | 41.5 | 87.94 |
| CP022319.1 | B. altitudinis SGAir0031 | Air sample | 3.84 | 41.4 | 87.95 |
| CP011150.1 | B. altitudinis W3 | Raw gallnut honey | 3.74 | 41.4 | 87.88 |
| CP027116.1 | B. pumilus 145 | Sediment top | 3.94 | 41.2 | 90.15 |
| CP000813.4 | B. pumilus SAFR-032 | Spacecraft facilities | 3.70 | 41.3 | 89.90 |
| CP011007.1 | B. pumilus SH-B9 | Rhizosphere of beet | 3.87 | 41.4 | 90.38 |
| CP018197.1 | B. safensis KCTC | Marine sponge | 3.97 | 41.4 | 97.31 |
| CP015607.1 | B. safensis U14-5 | Antarctic Lake | 4.15 | 41.4 | 97.56 |
| CP015611.1 | B. safensis U17-1 | Antarctic Lake | 3.74 | 41.5 | 95.97 |
| CP015610.1 | B. safensis U41 | Antarctic Lake | 3.73 | 41.5 | 95.99 |
| CP009749.1 | B. subtilis ATCC 19217 | Industrial strain | 3.95 | 46.4 | 78.25 |
| CP017786.1 | B. xiamenensis VV3 | Fermented rice | 3.63 | 41.4 | 88.35 |
| CP010075.1 | Bacillus sp. WP8 | Rhizosphere of wheat | 3.70 | 41.6 | 95.90 |
bPercentage of G and C in the genome sequence, defined as 100 x (G+C)/(A+T+C+G).
cIdentity with B. safensis BRM1.
Genome annotation was performed using the NCBI Prokaryotic Genome Annotation Pipeline [17] on the fly during the genome deposit process at GenBank. Annotation was also performed using the RAST genome annotation service [18].
The carbohydrate-active enzymes coding genes in the strain BRM1 genome were identified using the CAZY online [19]. Ribosomal sequences were identified using RNAmmer [20] and ARAGORN [21].
The Bacillus safensis BRM1 genome map was generated using DNAPlotter [22]. Clusters of orthologous group (COGs) analysis were performed using the EggNog 4.5.1 software [23]. PSORTb v3.0 was used to predict the protein subcellular localization [24].
3. RESULTS AND DISCUSSION
3.1. Genome Analysis
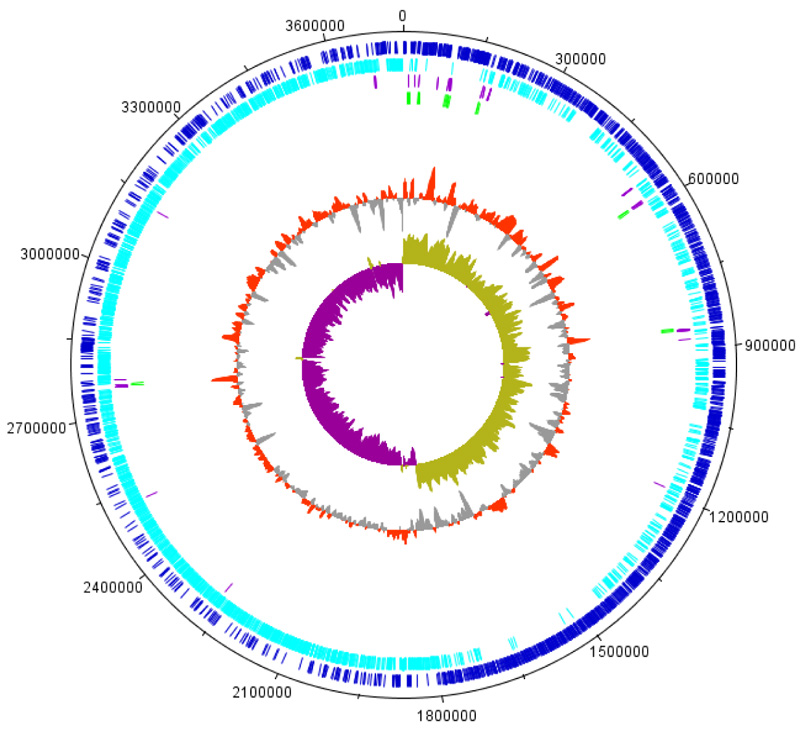
The genome of BRM1 consists of one circular chromosome with 3,740,842 pb (Fig. 1) and a GC content of 41.80%. BRM1 strain has eight rRNA operons: eight 5S rRNAs, eight 16S rRNAs, and eight 23S rRNAs. There were 91 tRNAs, covering all 20 amino acids. The number of coding sequences (CDS) was 3.699, with an average length of 837 bp. The content of non-coding regions was 10.82%. The genome features of the BRM1 strain were similar to those of other Bacillus safensis strains.
3.2. Phylogenetic Analysis
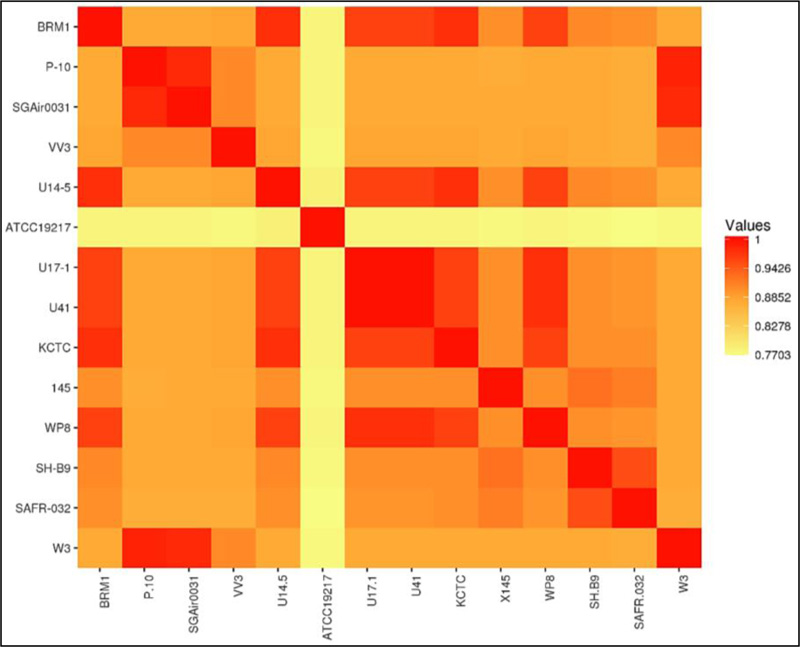
Genome similarity analysis was performed using the Average Nucleotide Identity tool [25]. Table 1 shows the genome features of the 13 Bacillus strains included in this study, selected due to their identity of the 16S rRNA of BRM1 and their genomes were completely sequenced. The genome sizes ranged from 3.63 Mb to 4.15 Mb, G+C content ranged from 41.2% to 46.4%, and CDS numbers ranged from 3556 to 3979. The results were plotted as a heatmap (Fig. 2). The identity between BRM1 and 13 complete Bacillus genomes varied from 77% to 97.56%.
The identity of BRM1 with B. safensis strains varied from 95.99% to 97.56%, being the highest identify degree with B. safensis strain U14-5 (97.56%). The next closest species (B. pumilus) had identities of 90% or lower. Since the identities with B. safensis were above 95%, these results support the classification of the BRM1 within the B. safensis species [25].
A genome comparison was performed at the SEED Viewer platform [27]. The results revealed a set of genes exclusively present in BRM1, encoding putative enzymes such as chitinase and ketoglutarate semialdehyde dehydrogenase, the DNA repair methyltransferase, glycerol dehydrogenase, nitrilase, and an exonuclease (Table 2).
3.3. Annotation and Protein Analysis
To annotate gene functions in B. safensis BRM1 genome, we use the EggNog 4.5.1 program [23], which performs a cluster of orthologous group (COGs) analysis classifying the proteins/ORFs into specific categories related to different aspects of physiology and cellular metabolism. The category distribution is depicted in the supplementary material (Table S1). Genes related to amino acid (E, 9.0%), transcription (K, 8.6%) and carbohydrate metabolism (G, 6.7%) comprised 24.3% of total protein coding regions identified. Genes for a protein belonging to unknown functions represented 24.6% of the sequences. Other categories, such as cell wall/membrane (5.3%), translation (5.1%), and ion transport and metabolism (5.1%) proteins, were also highly prevalent.
3.4. Hydrolytic Enzymes
Glycosyl hydrolases (GHs) are widely used in biomass-to-sugar conversion in food and biofuel producing processes. Analyses of BRM1 strain genome using carbohydrate-active enzyme (CAZy) database [19] identified 35 genes encoding putative biomass degradation enzymes (Table 3 and Table S2) such as cellulases, xylanases, and acetyl xylan esterases (involved in xylan degradation). The encoded enzymes were classified into 18 different families of glycosyl hydrolases (GHs) and carbohydrate esterases (CEs).
|
Number ID at SEED Viewer |
Probably Function at Seed Viewer | Length (pb) | Sequence ID at NCBI GenBank |
|---|---|---|---|
| 1027 | Chitinase | 276 | WP_155889681.1 |
| 893 | Ketoglutarate semialdehyde dehydrogenase | 927 | WP_065214177.1 |
| 556 | DNA repair methyltransferase | 1194 | WP_070326855.1 |
| 225 | Glycerol dehydrogenase | 1083 | WP_081123921.1 |
| 2266 | Nitrilase | 1008 | WP_170825868.1 |
| 3802 | Exonuclease SbcC | 1974 | WP_034283567.1 |
| Probable Activity | Conserved Domains (# of Genes Found) |
|---|---|
| Cellulases | - |
| Endo/exoglucanases | GH 5 (1), 9 (1), 48 (1), 51(1) |
| β-Glucosidase | GH 1 (6), 3 (1) or 42 (1) |
| Hemicellulases | |
| Xylanases or β-Xylosidases | GH 10 (1), 11 (1), 16 (1), 30 (1), 43 (2) |
| α-galactosidases | GH 4 (4) |
| Galactanases Acetyl xylan esterase |
GH 53 (1) CE 4 (9), 6 (1), 7 (1), 12 (1) |
| - | (Total: 35 genes for putative hydrolases) |
To infer which biomass degradation enzymes could be secreted to the extracellular medium, we use the PSORTb v3.0 software (yu et al. 2010). As a result, 12 of those enzymes were predicted to be targeted to the extracellular medium (Table S2).
Additionally, RAST analysis [18] showed that the xylan utilization pathway is complete in the BRM1 strain genome: xylose isomerase (XylA, ARD56435.1), xylulose kinase (XylB, ARD56434.1), xylose-responsive transcription regulator (XylR, ARD56436.1), xyloside transporter (XynT, ARD56438.1), and three xylan hydrolases (putatives XynA, XynB and/or XynC: ARD56360.1, ARD56433.1 and ARD56437.1). These results show that the BRM1 strain genome encodes the enzymatic machinery to degrade recalcitrant carbon sources including xylan, cellobiose, and CMC, hence, confirming the biomass degrading potential of BRM1.
3.5. Growth On Specific Substrates
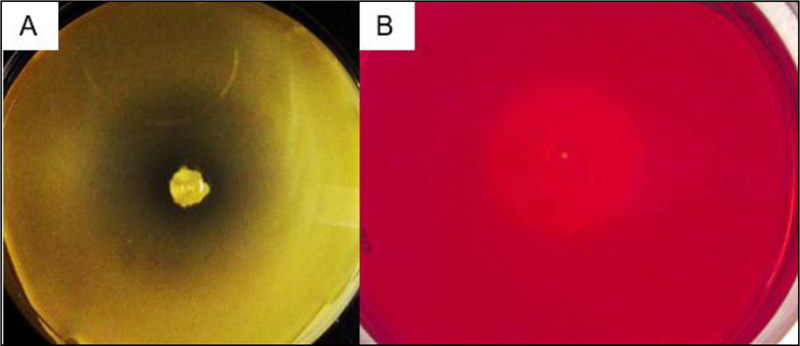
The BRM1 strain was isolated from a Brazilian mangrove sample plated on solid media containing CMC or xylan as a sole carbon source (as described in the Material and Methods section). This strain is capable of growing in 2% (w/v) NaCl using either substrate as carbon source, and the hydrolysis halo could be detected after 4 days of growth at 30°C (Fig. 3).
In order to confirm the hydrolytic potential, the BRM1 strain was cultured aerobically in a liquid medium supplemented with glucose (control), cellobiose, xylan, or carboxymethyl cellulose (CMC). For this analysis, the minimal medium CSE described by Commichau et al. (2008) was used, which provided better homogeneity during cell growth, without the appearance of biofilm and bacteria flocculation observed using LB or M9 medium (data not shown).
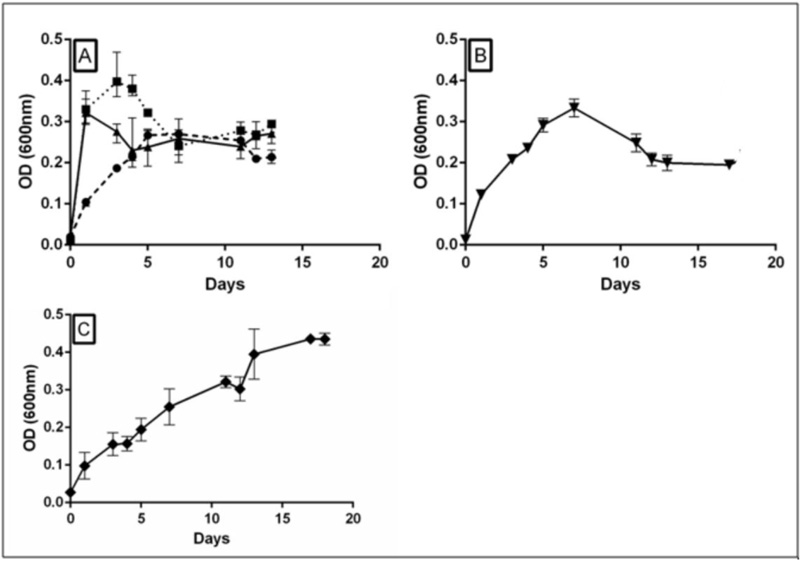
The growth curves in Fig. (4) indicate that BRM1 was capable of growing in CSE minimal medium with no addition until day 5, probably consuming the glutamate and succinate present in the broth. After this, growth stopped reaching a stationary phase. When cellobiose was added, the bacteria grew much faster, similarly to glucose, reaching maximum cell density 1-2 days of incubation. On the other hand, when xylan was used, the initial growth rate was somewhat faster than with only CSE, but the cells continued to grow even after 17 days of incubation. When CMC was used as a carbon source, the initial growth rate was marginally higher than the control condition (CSE only) and also reached maximum cell density higher than the control after 5-7 days. Since BRM1 produced a hydrolysis halo in a solid medium plate containing CMC, the growth profile in liquid medium with CMC suggests that cellulolytic activity is low, or perhaps aeration could interfere in CMCase activity.
CONCLUSION
Here we report the isolation of B. safensis BRM1 from the Brazilian mangrove soil, a strain with the capacity to grow using several recalcitrant polysaccharides. The genome sequence of this strain represents the sixth complete genome of Bacillus safensis deposited on the GenBank database. The other strains with complete genome described were also isolated from saline environments: Antarctic Lake and Chinese ocean.
The BRM1 strain genome encodes the enzymatic machinery to degrade different carbon sources. At least 12 enzymes potentially involved in the degradation of CMC/cellulose could be identified in the genome of BRM1; these enzymes could allow the efficient degradation of cellulose to cellobiose and then to D-glucose. Genes encoding enzymes involved in the hydrolysis of xylan to xylose were also found which would allow hemicellulose biomass degradation.
The complete genome of Bacillus safensis BRM1 revealed that this strain may have biotechnological potential due to the identified cellulases and hemicellulases. Future studies should be conducted to evaluate the activities of these enzymes separately and the possible usage in biomass-to-biofuel conversion processes.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
This work was supported by the National Institute of Science and Technology of Nitrogen Fixation/CNPq/MCT, Fundação Araucária and Coordination for the Improvement of Higher Education Personnel (CAPES).
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We thank Dr. Paulo da Cunha Lana who helped during the collecting of mangrove sediment.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publisher’s website along with the published article.


