All published articles of this journal are available on ScienceDirect.
Monitoring and Evaluation of Antibiotic Resistance Pattern of Escherichia coli Isolated from Drinking Water Sources in Ardabil Province of Iran
Abstract
Background and Objective:
Antibiotic resistance is rapidly spreading among bacteria, such as Escherichia coli, which have been discharged into water sources by humans and animals, in part due to genes carried by integrons. The goal of this study was to track and assess the pattern of antibiotic resistance among Escherichia coli strains isolated from drinking water sources in Iran’s Ardabil province.
Methods:
Escherichia coli strains were isolated and identified using standard biochemical and microbiological techniques from Ardabil province’s water supply sources in 2019 and 2020. The class 1 integron gene was detected using the polymerase chain reaction (PCR). The disk diffusion method was used to determine antibiotic resistance and sensitivity.
Results:
The Int I gene was found in 47 out of 200 isolates (23.5%). The antibiotic streptomycin had the highest resistance (100%), whereas gentamycin, ciprofloxacin, tobramycin, tetracycline, cefipime, imipenem, meropenem, chloramphenicol, and trimethoprim-sulfamethoxamide had the lowest resistance. Gentamycin, ciprofloxacin, tobramycin, tetracycline, cefipime, imipenem, meropenem, and trimethoprim-sulfamethoxazole, on the other hand, had the highest sensitivity of 100%, while streptomycin had the lowest (0%). Also, the highest level of pollution of Escherichia coli in drinking water supply sources in the province is in Parsabad city (42.86%), and the lowest level of pollution is in Kosar city (6.67%).
Conclusion:
According to the findings of this study, the high prevalence of Escherichia coli strains resistant to the antibiotic streptomycin circulating in water sources should be considered a major problem in terms of antibiotic resistance spreading among bacteria in water.
1. INTRODUCTION
Antibiotics are the most effective kind of chemotherapy produced in the 20th century, saving many lives every day [1]. Antibiotic resistance limits clinical antibiotic use, and as resistant bacteria become more common, there is a growing fear that existing antibiotics will become ineffective and more expensive against these diseases [2]. Antibiotic-resistant genes have been found in a number of aquatic habitats, including drinking water, in both industrialized and developing countries [3]. The most serious hazard to public health is the transfer of resistance genes from environmental bacteria to human pathogens. Drinking water’s ability to spread microbial infections to a larger number of individuals, resulting in sickness, has been widely documented in countries of all economic levels [4, 5]. Furthermore, having access to safe drinking water is critical for avoiding epidemic disease and enhancing the overall quality of life [6, 7]. According to the World Health Organization, contaminated water is responsible for 80% of all infections [8]. Water-related disorders, such as diarrhoea, are particularly prevalent in developing nations, accounting for 10% of the disease burden [9].
Escherichia coli (E. coli) is a type of fecal coliform that can contaminate drinking water when it comes into contact with human or animal feces. For decades, E. coli has been the most reliable indicator of faecal contamination in water quality monitoring. These coliforms may be swept into creeks, rivers, streams, lakes, or groundwater during heavy rains. Integrons are bacterial genetic components that allow the shuffling of smaller mobile elements known as gene cassettes, and they have been dubbed a genetic assembly kit for bacteria because they are found in untreated drinking water from these sources [10]. Integrons have a function in the evolution and transmission of antibiotic resistance genes in gut bacteria. Integrons containing intI1 are classified as “class I,” intI2 as “class II,” intI3 as “class III,” etc., based on the amino acid sequences of the integrases. Most of the integrons detected in clinical isolates are class I integrons, which are common in gram-negative bacteria and have been related to the global spread of antimicrobial drug resistance. As a result, integron-positive isolates can resist antibiotic treatment [11].
Some strains of E. coli emerge as pathogens due to the presence of pathogenic characteristics and virulence genes situated in the transmissible genetic element that distinguishes them from regular commensal strains [12]. E. coli was also discovered in the investigation, and it is a helpful marker when researching resistance in bacterial populations because it contains a large number of genes coding for antimicrobial drug resistance. Antibiotics used indiscriminately in agriculture and released from sewage treatment plants have accelerated the establishment of antibiotic-resistant bacteria in environmental water bodies, necessitating accurate and frequent monitoring [13, 14]. Given that water is closely tied to human activities and health [15], it is critical to assess pathogenic microorganisms and antibiotic resistance in human drinking water sources.
The main objective of this study is to determine the antibiotic resistance pattern and multiple antibiotic resistance index of E. coli strains recovered from various water sources over a year of bacteriological monitoring.
2. MATERIALS AND METHODS
2.1. Sample Collection
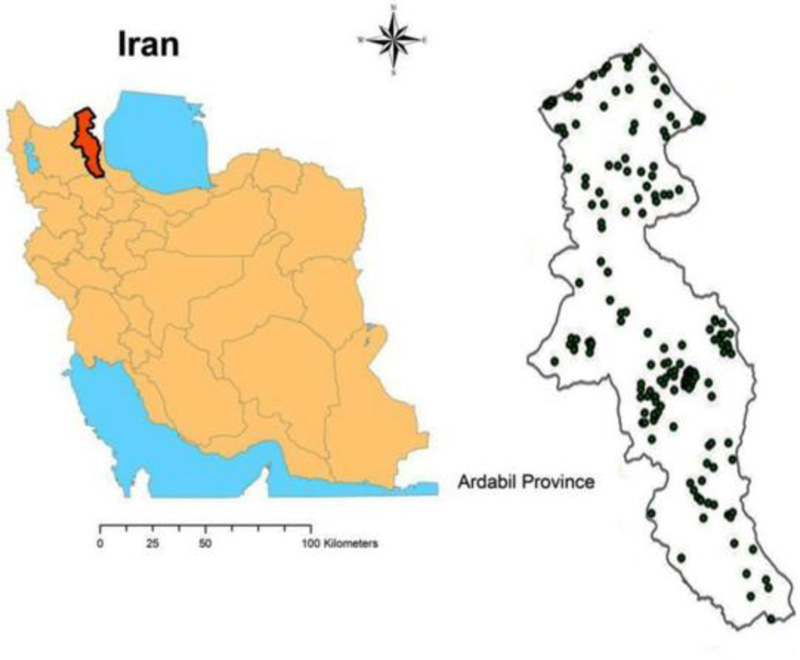
A total of 200 samples were taken from 10 counties (Ardabil, Namin, Sareyn, Nir, Kowsar, Khalkhal, Meshgin Shahr, Germi, Bileh Savar, and Parsabad) from various water sources such as dams, stream sources, rivers, and wells. Samples were gathered from places that were indicative of the water sources and/or distribution networks from which water is provided to the inhabitants and/or points of use, based on criteria such as population and the extent of use or patronage of these sources. The majority of the communities are dominated by farmers. The principal source of water for the residents of each of the communities was a borehole or a stream.
2.2. Site Observation Details
Important observations were obtained surrounding the sampling sites prior to water sampling. These observations covered sanitary conditions as well as potential sources of pollution that could affect the quality of water from the sources analyzed.
Water clarity/turbidity (visual clarity in the water, i.e., leaves, debris, and algae), weather conditions (temperature, wind, and rainfall), the presence of animals (birds/ducks), and other comments (e.g., system problems, i.e., disinfection /filtration equipment, and fecal accidents) were also recorded in the field.
2.3. Sample Size and Sampling Frequency
Between October 2019 and September 2020, 200 water samples were collected for analysis from various water sources. Standard Methods for the Examination of Water and Wastewater [16, 17] and WHO recommendations for drinking water quality [18, 19] were used for all water sampling and preservation methods. Sampling for bacteriological investigation was done aseptically and carefully, ensuring that samples were not contaminated from the outside. Within two hours, all samples were transported to the laboratory.
2.4. Bacteria Isolation and Identification
To simultaneously detect coliform and E. coli bacteria, depending on the type of sample, the samples were cultured in 10 tubes, 15 tubes, and P / A in LMX Broth culture medium. Eosin methylen blue (EMB) agar plates (Merck, Germany) were streaked with positive samples and incubated at 37°C for 24 hours. Suspected E. coli colonies were identified by standard methods based on colonial appearance and bacterial morphology. Colonies that are indicative of E. coli in the EMB agar plate showed a metallic sheen (green) color [20]. Mixed cultures were restreaked for purity on plates of sterile EMB agar, and biochemical identification was done on isolates. E. coli was detected as a pure colony that was positive for indole, negative for citrate, positive for methyl red, and negative for Voges-Proskauer [21].
2.5. DNA Extraction
DNA extraction was performed by boiling method. Nanodrop and electrophoresis were used to assess the quantity and purity of the isolated DNA, which was then kept at -20ºC [21]. Two pairs of Sinaclon primers were prepared (Table 1). The Int I gene primers used in the present study are capable of amplifying the 287 bp fragment.
| Target Gene | Primer Sequence | Size (bp) | Reference |
|---|---|---|---|
| F.Int I | 5ʹ-TCTCGGGTAACATCAAGG-3ʹ | 287 | Kheiri et al. [22] |
| R.Int I | 5ʹ-GTTCTTCTACGGCAAGGT-3ʹ |
2.6. The PCR Reaction for E. coli Isolates
The PCR reaction was used to examine E. coli isolates using previously published conditions and techniques [23]. The 287 bp segment of the int1 gene was amplified using only one pair of primers, as shown in Table 1.
A 25 µl reaction mixture containing Master Mix (12.5 µl), template DNA (2 µl), a primer (1 µl) and 8.5 µl of sterile distilled water was used for the PCR reaction. In a DNA thermocycler (Model CP2-003; Corbett, Australia), the PCR reaction was carried out as follows: initial denaturation at 95°C for 5 min, followed by 35 cycles at 95°C for 5 min, annealing at 54°C for 1 min, extension at 72°C for 1 min, and the last extension step at 72°C for 5 min, followed by chilling at 4°C. On a 1% agarose gel containing ethidium bromide, the PCR products were electrophoresed for 30 min at 90 volts.
2.7. Antibacterial Susceptibility Testing of E. coli
From a fresh culture of 18-24 hours of bacteria grown in nutrient broth medium, a suspension equivalent to half McFarland was prepared in nutrient broth medium. Subsequently, Müller Hinton agar was completely grassed by swap. After culture, antibiogram disks were selected, and antibiotic disks were placed 22 mm apart and 16 mm from the plate wall. They were then heated at 35 °C for 16-20 hours [18]. The diameter of the growth inhibition zone was then assessed for all drugs. CLSI [24] was used to interpret the findings. Antibiogram tests were classified as sensitive, resistant, or semi-sensitive for each antibiotic.
The E. coli isolates were tested against the following antibiotics: amikacin (30 µg), gentamycin (10 µg), ciprofloxacin (5 µg), tobramycin (10 µg), tetracycline (30 µg), cefipime (30 µg), imipenem (10 µg), meropenem (10 µg), chloramphenicol (30 µg), streptomycin (10 µg), trimethoprim-sulfamethoxazole (1.25/23.75 µg), and ampicillin (10μg).
3. RESULTS
The pattern of antibiotic resistance in E.coli isolated from drinking water sources in Iran’s Ardabil province was investigated in this study. E. coli was detected in 200 samples collected over the course of the investigation. E. coli was found in samples collected from 10 counties. The locations where water samples were collected are shown in Fig. (1).
Results from Table 2 show that from the 200 water samples cultured, on 47 samples, PCR tests were performed, all of which contained the Int I gene. The highest number of E. coli was isolated from Parsabad 15 (32%), followed by Bilasavar 9 (19%), Ardabil 8 (17%), Germi 5 (12%), Khalkhal, Sarein, Meshginshahar, and Namin 2 (4% each), and also Kowsar and Nir 1 (2% each).
| Water Sources | Number of Samples Analyzed | Number of Strains of E. coli isolated |
Percentage Distribution (%) |
|---|---|---|---|
| Ardabil | 40 | 8 | 17 |
| Parsabad | 35 | 15 | 32 |
| Bilasavar | 25 | 9 | 19 |
| Khalkhal | 15 | 2 | 4 |
| Kowsar | 15 | 1 | 2 |
| Sarein | 10 | 2 | 4 |
| Nir | 10 | 1 | 2 |
| Meshginshahar | 15 | 2 | 4 |
| Germi | 20 | 5 | 12 |
| Namin | 15 | 2 | 4 |
| Total | 200 | 47 | 100 |
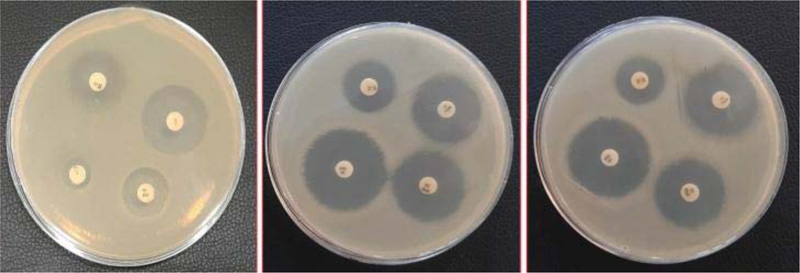
In this study, the PCR product appeared as a single band DNA with a size equal to 287 bp fragment corresponding to the int1 gene in E. coli strains (Fig. 2). All isolates of drinking water sources in Ardabil province were tested for antibiotic susceptibility using the disk diffusion method (Fig. 3).
Table 3 shows the antibiotic resistance pattern of E. coli isolates from Ardabil province in drinking water sources. The E. coli isolates showed the highest resistance to streptomycin 47(100%), followed by Amikacin 15 (31.9%) and ampicillin 10 (21.3%), while it was more sensitive to tobramycin, tetracycline, cefipime, imipenem, meropenem, gentamycin, ciprofloxacin and trimethoprim-sulfamethoxazole 47 (100%), followed by ampicillin 28 (59.6%), amikacin 23 (48.9%) and chloramphenicol 18 (38.3%) (Fig. 4).
4. DISCUSSION
The presence of E. coli in various water sources could result in health risks such as diarrhoeal illnesses, which cause a significant amount of morbidity and death in both adults and children [25-29]. Antibiotics may be used for the treatment of diarrhoea. Nonetheless, multiple E. coli strains have been identified. Antibiotic resistance has been documented in E. coli [30-32].
| Antibiotics | Number of Isolates (%) | ||
|---|---|---|---|
| Resistant | Intermediate | Sensitive | |
| Streptomycin | 47 (100) | 0 (0) | 0 (0) |
| Amikacin | 15 (31.9) | 9 (19.2) | 23 (48.9) |
| Ampicillin | 10 (21.3) | 9 (19.2) | 28 (59.6) |
| Tobramycin | 0 (0) | 0 (0) | 47 (100) |
| Tetracycline | 0 (0) | 0 (0) | 47 (100) |
| Cefipime | 0 (0) | 0 (0) | 47 (100) |
| Imipenem | 0 (0) | 0 (0) | 47 (100) |
| Meropenem | 0 (0) | 0 (0) | 47 (100) |
| Chloramphenicol | 0 (0) | 29 (61.7) | 18 (38.3) |
| Gentamycin | 0 (0) | 0 (0) | 47 (100) |
| Ciprofloxacin | 0 (0) | 0 (0) | 47 (100) |
| Trimethoprim-Sulfamethoxazole | 0 (0) | 0 (0) | 47 (100) |
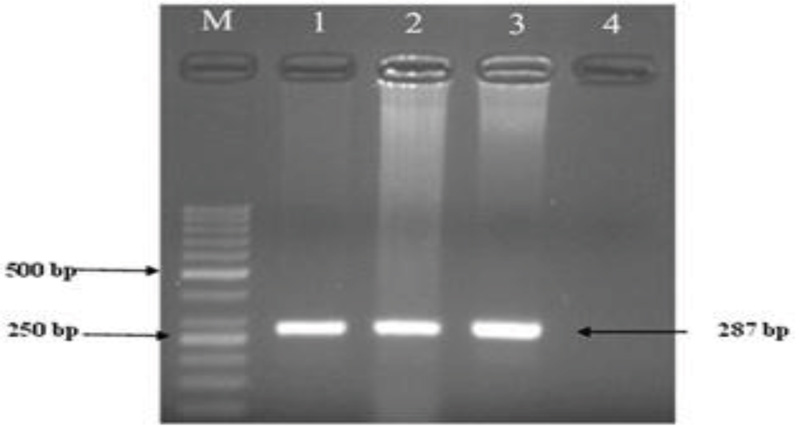
Note: From left to right: Column M: ladder 50 bp, marker scale; Columns 1 and 2: Positive specimens containing class I integron with a product size of 287 bp; Column 3: Positive control of class I integron; Column 4: Negative control.

From the 200 water samples cultured, 47 samples yielded growth of E. coli. The drinking water of the city of Parsabad accounts for 32%; thus, it had the highest number, followed by the drinking water of the city of Bilasavar 9 (19%). In a similar study done in Kaduna State, Nigeria, a total of 167 water sources for drinking were analysed, and 17 yielded growth of E. coli [33]. The result of the present study was also consistent with the work of Larson et al., who also isolated E. coli from water samples in rural Andean Households in Cajamarca, Peru [34]. The work of Chen et al. on the prevalence of multidrug-resistant E. coli in drinking water sources in Hangzhou city shows a high prevalence of antibiotics resistant E. coli [35].
The present study monitored and evaluated the pattern of antibiotic resistance of E. coli isolated from drinking water supply sources in Ardabil province of Iran. The prevalence of resistance to streptomycin 47 (100%), amikacin 15 (31.9%), and ampicillin 10 (21.3%) was significantly higher than other tested antibiotics in water sources. In contrast, the highest sensitivity (i.e., 100%) was related to gentamycin, ciprofloxacin, tobramycin, tetracycline, cefipime, imipenem, meropenem, and trimethoprim-sulfamethoxazole, and the lowest was related to streptomycin (0%). The E.coli isolated from the commercial water sources showed the highest resistance to streptomycin 4 (66.7%). This is in line with the work of Aasmae et al., where the E. coli isolated showed high resistance to streptomycin [36]. Larson et al. [37] reported that the E. coli antibiotic resistance profile showed the highest resistance against tetracycline (37.6%), ampicillin (34.2%), sulfamethoxazole-trimethoprim (21.4%), and nalidixic acid (13%); these findings are in contrast with our results.
Ranjbar et al. [38] investigated the genetic resistance of beta-lactam-related antibiotics among Escherichia coli strains isolated from different water sources in Iran. The highest resistance was related to streptomycin (28.1%), gentamicin (27.1%), tetracycline (24%), trimethoprim-sulfamethoxazole (22%), tobramycin (10.4%), ciprofloxacin (7.3%), chloramphenicol (7%), amikacin (6.7%), and imipenem (6.1 ٪). In the present study, the highest resistance was related to streptomycin (100%), amikacin (31.9%), and ampicillin (21.3%), and the lowest resistance was related to other antibiotics, including gentamicin, tetracycline, trimethoprim-sulfamethoxazole, tobramycin, ciprofloxacin, and chloramphenicol (0%).
Odonkor et al. on the prevalence of multidrug-resistant E. coli isolated from drinking water sources, showed high resistance patterns to the tested antibiotics. They were most resistant to penicillin (32.99%), cefuroxime (28.87%), erythromycin (23.71%), and tetracycline (21.45%). In contrast, they were susceptible to nitrofurantoin (93.8%), cefotaxime and amikacin (91.75%), gentamicin (90.7%), nalidixic acid (89.65%), ciprofloxacin (74.2%), chloramphenicol (69.07%), pipemidic acid (65.97%), and cefuroxime (52.58%) [13].
Also, Udoh et al. [39] isolated 25 strains from drinking water sources in a tertiary institution in which 92% of the E. coli isolated showed resistance to the tested antibiotics. Resistance was higher with augmentin (64%), chloramphenicol (48%), and streptomycin 11 (44%), while most were sensitive to tarivid and perfloxacine (100%). Isolates from school storage water sources showed the highest resistance to augmentin (76.5%), while those from commercial storage water sources showed the highest resistance to streptomycin (66.7%). These findings contradict our findings and could be due to differences in sample source, cleanliness level, and geographical areas. As a result, it could be due to strict search-and-destroy and surveillance policies, as well as antibiotic prescription control. Antibiotic-resistant E. coli has been found in other research.
E. coli is found in surface waters; for example, Nontongana et al. [40] reported that E. coli recovered from the Kat river in South Africa were resistant to ampicillin (98%), tetracycline (13%), and streptomycin (8%). Dolejská et al. [41] reported that, on average, 17% of surface water E. coli isolates were resistant to one or more antibiotics in the Czech Republic. Toroglu et al. [42] isolated 67 strains from the Aksu river, with E. coli accounting for 67.2% of the total. Multiple antibiotic resistances were detected in 27 isolates (40%) that were resistant to five or more antibiotics, and Beta-lactamase generating bacteria were found in 49.3% of the isolates. This study found that E. coli has a significant level of antibiotic resistance.
According to Chatterjee et al. the susceptibility rate for cefixime was significant (85%). The resistance to streptomycin and ciprofloxacin was 75% and 30%, respectively, among the twenty E.coli isolates. Since all isolates were confirmed to be resistant to doxycycline, there was a high level of resistance recorded. Cefixime was shown to be the most effective antibiotic against E. coli isolates [43]. Furthermore, the results of 27,886 E. coli isolates from the Minjiang River in China showed that they were most resistant to cefoperazone, cefazolin, ciprofloxacin, norfloxacin, and rifampicin [44], which contradicts our findings.
Change of antibiotic resistance following the change in water quality leads to negative effects on public health. High frequency of resistant bacteria against amoxicillin and ampicillin has been reported by Muleta T. et al. [45].
From the 767 E. coli isolates in wastewater treatment plants, the highest resistance rates were found for ampicillin and piperacillin, cefalothin and cefuroxime-axetil, nalidixic acid, trimethoprim-sulfamethoxazole, and tetracycline [46].
CONCLUSION
According to the results of this study, the prevalence of streptomycin, amikacin, and ampicillin of E. coli (21-100%) is in all water sources commonly used by the people of Ardabil province. Water treatment is limited, and there is exposure to resistant bacteria. These results have raised concerns about the presence and distribution of these threatening factors in drinking water resources and their aftermath. Moreover, prevention and control strategies should be implemented immediately to prevent spread. Most of these species should be avoided.
In general, our results showed that the high prevalence of E. coli strains resistant to the antibiotic streptomycin circulating in water sources could be considered as a major concern regarding the spread of antibiotic resistance among bacteria in water.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
This work was supported by Islamic Azad University, Ardabil Branch, Ardabil, Iran.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors thank the Department of Biology, Islamic Azad University, Ardabil Branch. The authors also sincerely thank the Ardabil Water and Sewerage Company and the laboratory staff for their laboratory support. Finally, we would like to thank Dr. Ruhollah Kheiri for his sincere cooperation in this research.


