All published articles of this journal are available on ScienceDirect.
Bacteriome Identified by Next-Generation Sequencing in Saliva, Dental Plaque, and Tumor Tissue of Patients with Oral Squamous Cell Carcinoma
Abstract
Background:
Oral squamous cell carcinoma (OSCC) is the sixth most common cancer in the world, and the bacterial microbiome has been considered a risk factor that could play an important role in carcinogenesis.
Objective:
A bacteriome study was performed by next-generation sequencing in dental plaque, saliva, and tumor samples of 10 OSCC patients and compared with bacteriome in dental plaque and saliva of 10 patients without OSCC.
Methods:
DNA was extracted from all samples and sequenced by Illumina technology MiSeq™. Bioinformatic analyzes were performed for evaluated sequence quality, alpha and beta diversity, bidirectional analysis of variance (p <0.05), and principal component analysis. After establishing bacterial profiles associated with each sample and population, intragroup and intergroup comparisons were carried out. For bacteria identification compatible with eubiosis and dysbiosis processes, a screening was performed based on the frequency of appearance in all patient samples with and without OSCC. Lastly, frequency, average, standard deviation, Chi-square, and Mann Whitney test were calculated.
Results:
Out of the identified 1,231 bacteria in the populations under study, 45 bacterial species were selected, of which 34 were compatible with eubiosis, and 11 were compatible with dysbiosis. Among the bacteria compatible with eubiosis were species of Lactobacillus and Streptococcus, Chromobacterium violaceum, Enterobacter asburiae, Mycobacterium chubuense, Mycoplasma penetrans, and Brachyspira intermedia. Among the species associated with dysbiosis, Providencia stuartii, Capnocytophaga canimorsus, Legionella pneumophila, and Mycoplasma hominis were notable.
Conclusion:
Thirty-four bacterial species may be associated with eubiosis or healthy states and 11 bacterial species could be associated with dysbiosis or pathogenic state, OSCC.
1. INTRODUCTION
Oral squamous cell carcinoma (OSCC) is the sixth most frequent cancer worldwide, representing more than half a million cases per year [1]. It is one of the main causes of morbidity and mortality, with more than 275,000 cases and 128,000 deaths per year and survival of up to five years [2]. Although this pathology can be prevented in the early stages of carcinogenesis, mainly in pre-neoplastic lesions, its detection and diagnosis are still insufficient, in part due to nearly undetectable symptoms for long periods of time [3]. Among the multiple factors associated with OSCC are carcinogenic chemicals (components of tobacco smoke and alcohol), physical agents (mainly radiation), and biological agents (virus, bacteria, and others) [4, 5].
Among the biological agents, viruses are the most studied. The human papillomavirus (HPV) has been related to OSCC [6, 7], principally due to the action of two viral oncoproteins known as E6 and E7. These proteins induce p53 degradation and pRB inactivation, respectively, leading to inhibition of apoptotic processes and an increase in cell proliferation [8]. Another important biological agent is the oral bacterial community or microbiota, where hundreds of bacteria co-exist in different microenvironments or niches, such as saliva, dental plaque, and soft tissues, among others. Under healthy conditions, the microbiota is in symbiosis, conferring benefits to the host. However, dysbiosis can be associated with pathological conditions, such as dental caries, periodontal disease, and even OSCC, given the capacity to potentiate colonization and proliferation of highly pathogenic organisms [9, 10].
Recent research has suggested that microbial dysbiosis could contribute to oral cancer development. The presence of some bacteria, such as Fusobacterium nucleatum, Fusobacterium periodonticum, Pseudomonas aeruginosa, Prevotella intermedia, Aggregatibacter segnis, Peptostreptococcus stomatis, Catonellamorbi, might have a pathogenic contribution to OSCC development [11, 12].
Possible mechanisms of microbiota influence on OSCC development could be the induction of chronic inflammatory processes, by which a hostile microenvironment is generated, resulting in increased cell proliferation, mutagenesis, oncogene activation, and angiogenesis [13]. Therefore, there is a need to identify the mechanisms by which bacteria may be involved in OSCC development and define these events as effective biomarkers that could help in designing prevention, control, and prognosis strategies with an impact on oral cancer incidence and prevalence [14].
It has been proposed that a comprehensive understanding of the microbiome and its change during healthy and pathological states may reveal that microorganisms are differentially associated with OSCC, which could be useful as biomarkers [15]. To achieve this aim, it is necessary to overcome some difficulties: (i) the human microbiome is large and diverse [16], (ii) conventional microbiological techniques for bacterial isolation and identification are of low-spectrum sensitivity and, in addition, a large number of microorganisms cannot be cultured in vitro [17], (iii) a great number of external factors (food, hygiene, population, etc.) that affect the qualitative and quantitative composition of the microbiome [18].
Bacteriome studies on OSCC patients based on 16S rRNA amplicon gene sequencing have reported significant differences in diversity and abundance of certain microorganisms in tumor samples, mainly related to the phyla Actinobacteria, Bacteroidetes, Firmicutes, Proteobacteria, Spirochetes, Synergistetes, Tenericutes, Gracilibacteria, Absconditabacteria, and Saccharibacteria [11, 19, 20]. However, there has been no consensus in the interpretation of those findings. Thus, there is a need for sequencing studies with greater and more in-depth resolution [21, 22] that might shed light on the composition and interaction of the microbial community of the OSCC microenvironment in a more precise manner at the genus and species level.
To date, limited research has been undertaken to study bacterial species associated with OSCC [23]. In this study, we have identified by next-generation sequencing the bacterial microbiome present in different oral environments in ten patients with OSCC (tumor tissue, dental plaque, and saliva) and in ten matched controls without OSCC (saliva and dental plaque), suggesting bacterial species associated with eubiosis and dysbiosis processes.
2. MATERIALS AND METHODS
2.1. Sample Collection
Thirty oral samples were collected (10 of tumor tissues, 10 of saliva, and 10 of bacterial plaque) from10 patients diagnosed with OSCC and 20 oral samples (10 of saliva and 10 of bacterial plaque) from 10 patients without OSCC (controls) (it was not possible to collect healthy contralateral tissue because some patients did not approve of it) with established clinical parameters. It was assured that all patients and healthy donors included in the study had not taken antibiotics in the preceding 18 months. All samples were collected following the established protocol of the Human Microbiome Project. For saliva sample collection and processing, the recommendation given on the website (https://hmpdacc.org/hmp/resources/) were followed. These included collection of 5 mL of non-stimulated saliva in tubes containing RNAlatter® (Thermo Fisher Scientific) buffer that were stored at -70ºC. In addition, for bacterial plaque sample collection, the Human Oral Microbiome recommendations were followed. Briefly, the supragingival bacterial plaque was collected from four tooth surfaces (palate, tongue, vestibular and distal). The following criteria were used to score the samples: 0 when the bacterial plaque was absent, 1 when the bacterial plaque was thin, and 2 when the bacterial plaque was macroscopic. Samples were collected in tubes with RNA latter® buffer (Thermo Fisher Scientific) and stored at -70 ° C. Tumor tissue was collected after surgery; each sample was divided into two pieces. To preserve nucleic acids, one fragment was placed in RNA latter® buffer (Thermo Fisher Scientific) and stored at -70°C, until molecular analysis was carried out. The second fragment was sent to the pathology department for diagnosis confirmation.
2.2. DNA Extraction
After samples were individually thawed, DNA was isolated using the purification DNA Master Pure TM kit (Epicentre Biotechnologies® Madison, WI) following the manufacturer’s instructions, allowing for a high-quality DNA extraction through filtering and enzymatic action (Lysozyme). DNA was quantified by fluorescence using QuantiTdsDNA BR Assay Kit® (Invitrogen, Carlsbad, CA, USA) (≥1ug). Moreover, the integrity of DNA was verified in a 1% agarose gel (one high-weight band without bands in agarose gel). Samples that met good quantity and quality criteria were sent for next-generation sequencing with Illumina technology MiSeq™.
2.3. Sequencing and Data Curation
All samples were processed using the IlluminaNextera XT DNA library preparation kit following the manufacturer’s instructions. The libraries were quantified and grouped at equimolar concentrations. Automated cluster generation and final double-reads matching were performed according to the manufacturer's instructions.
All samples were sequenced using the MiSeq ™ system (Illumina, San Diego, CA). Briefly, as for the first step, the library was prepared through DNA sample random fragmentation followed by the addition of 5'and 3' adaptors that acted as reference points. Subsequently, clusters were generated using the libraries’ fragments to anchor them through oligos attached to a surface. Each fragment was then amplified in different clonal groups using bridge amplification. Sequencing was performed using the reversible terminated chemistry method, detecting individual nucleotides as they incorporated into the DNA template strand. The four dNTPs attached to reversible terminators were present during each sequencing cycle (www.illumina.com/technology/next-gene ration-sequencing.html).
2.4. Bioinformatic Analyses
Trimmomatic software was used to pre-process fastq files containing sequences and to select high-quality sequences. A minimum threshold of 20 was used, where low-quality sequences from the 3’ end with less than 100 bp were not considered. Nextera adapters were eliminated from the sequences, and a filter was applied to eliminate low-quality nucleotide reads. To visualize data quality, FastQc was used (Available in www.bioinformatics.babraham.ac.uk/projects /fastqc/).To identify bacteriome, Kraken Aligner V1.0 and Kraken DB database were utilized for each sample taxonomic diversity analysis (https://ccb.jhu.edu/software/kraken/MAN UAL.html#kraken-databases). Alignments were performed with complete bacteria RefSeq genomes, and a 99% similarity threshold between sequences was used to obtain bacteria classification from Phylum to species for each sample.
For identification of the most representative bacterial genera and species in each microenvironment to be analyzed, comparisons between groups and within groups were made, complemented with statistical analysis of bacterial frequency (number of times the bacteria occur in studied samples, expressed as a percentage), average, standard deviation, using Excel V.1017 and Rstudio. (www.rstudio.com/products/ rstudio/). Alpha and beta diversity were calculated based on read counts by species. Due to the high density of bacteria to evaluate the significance of the identified bacteria, bidirectional analysis of variance (p <0.05) and principal component analysis were performed to reduce dimensionality. To identify bacteria compatible with eubiosis and dysbiosis, screening was carried out between saliva, dental plaque, and tumor tissue samples from patients with and without OSCC. In addition, a difference in frequency from 40% to 60% was taken into account as a parameter, complemented with statistical analysis of frequency, mean, standard deviation, Chi-square, and Mann Whitney (U test).
3. RESULTS
3.1. Demographic, Clinical, and Health Characteristics of Patients with OSCC and Controls
A total of 10 patients with OSCC and 10 controls were included. The average ages (mean ± SD) for both groups were 56.4 ± 18.36 and 56.5 ± 15.59 years, respectively. Females represented 90% of the total sample population. The most common site of tumor location was the tongue (50%), followed by the hard palate (30%) and lips (20%). A total of 10% of the patients with OSCC and controls smoked, 50% of the controls and 10% of the patients with OSCC consumed alcohol, and 90% of the sample population were medicated to treat systemic diseases, such as high blood pressure and dyslipidemias. None of the individuals received treatment for cancer or infections. The state of oral health (good, regular, or bad) was calculated using the Löe-Silness index. In patients with OSCC, it was bad (40%), while in healthy individuals, it was well (50%). Taking into account that it was a paired-design study, the group without OSCC (controls) presented clinical and health characteristics similar to OSCC patients (no significant differences were observed for the studied parameters (p=0.14) (Table 1).
3.2. High-Quality Sequences for Alignments
The average of reads initially obtained for OSCC patients was 646,225.8 bp in plaque, 566,467.5 bp in saliva, and 483,722.4 bp in the tumor, with an average length of 300 bp. After data curation, average reads were 276,801.6 bp in plaque, 468,862.5 bp in saliva, and 227,289.9 bp in tumor with lengths between 100 bp and 291 bp. On average, 60, 25, and 15% of the reads were assigned to dental plaque, saliva, and tumor, respectively. In patients without OSCC, the average reads were initially 1,258,362.8 bp in dental plaque and 638,546.1 bp in saliva, with an average length between 250 bp and 300 bp. After data curation, average reads were 317,082 bp in dental plaque and 370,980.1bp in saliva with lengths between 100 bp and 291 bp. On average, 80 and 20% of the reads were assigned to dental plaque and saliva, respectively (Supplementary 1).
| Group | Age (Years) | Sex M=Male F=Female |
Smoker | Alcohol Consumption | Tumor Localization | Medicaments Consumption |
|---|---|---|---|---|---|---|
| Patient 1 | 45 | M | No | No | Tongue | Yes |
| Patient 2 | 65 | F | No | No | Tongue | Yes |
| Patient 3 | 62 | F | No | No | Cheek | Yes |
| Patient 4 | 62 | F | No | No | Tongue | Yes |
| Patient 5 | 30 | F | No | No | Tongue | Yes |
| Patient 6 | 44 | F | Yes | Yes | Cheek | Yes |
| Patient 7 | 85 | F | No | No | Cheek | Yes |
| Patient 8 | 69 | F | No | No | Lip | Yes |
| Patient 9 | 30 | F | No | No | Tongue | No |
| Patient 10 | 72 | F | No | No | Lip | Yes |
| Control 1 | 48 | M | No | No | NA | No |
| Control 2 | 60 | F | No | No | NA | Yes |
| Control 3 | 62 | F | No | No | NA | Yes |
| Control 4 | 60 | F | No | Yes | NA | No |
| Control 5 | 36 | F | No | No | NA | No |
| Control 6 | 45 | F | Yes | Yes | NA | Yes |
| Control 7 | 80 | F | No | Yes | NA | No |
| Control 8 | 68 | F | No | Yes | NA | Yes |
| Control 9 | 33 | F | No | Yes | NA | No |
| Control 10 | 73 | F | No | No | NA | Yes |
| Patients | Mean 56,4 SD±18,36 | F=9 (90%) M=1 (10%) | Smoker= 1 (10%) No Smoker=9 (90%) | Yes: 1 (10%) No = 9 (90%) | Tongue=5 (50%) Cheek=3 (30%) Lip=2 (20%) | Yes =9 (90%) NO=1 (10%) |
| Controls | Mean 56,5 SD±15,59 | F=9 (90%) M=1 (10%) | Smoker= 1 (10%) No Smoker=9 (90%) |
Yes: 1 (10%) No= 9 (90%) | None | Yes =5(50%) NO=5 (50%) |
3.3. Dental Plaque Bacterial Microbiota Present in OSCC Patients and Controls: Intergroup Comparisons
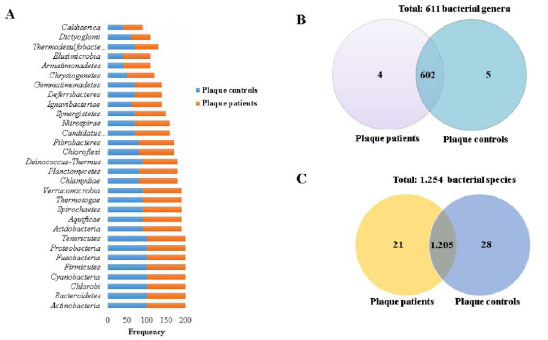
Comparisons of dental plaque bacteriome were made between patients with OSCC and controls. A total of 30 bacterial phyla were identified; the most frequent (100%) and abundant (2,000 to 15,000 reads) were Actinobacteria, Bacteroidetes, Firmicutes, Fusobacteria, and Proteobacteria (Fig. 1a and Supplementary 2A). In total, for both groups, 611 genera were identified, of which 602 were shared among patients and controls. The most frequent (100%) and abundant (500 to 8,000 reads) were Aggregatibacter, Bacteroides, Capnocytophaga, Corynebacterium, Fusobacterium, Haemophilus, Neisseria, Porphyromonas, Prevotella, Rothia, Selenomonas, Streptococcus, Tannerella, and Veionella. Out of the nine genera that were not shared, five were only observed in controls and corresponded to Neorickettsia, Desulfocapsa, Candidatus Riesia, Aquifex, and Acetomicrobium. The remaining four were only identified in plaque in patients with OSCC and were Wigglesworthia, Parageobacillus, Coprothermobacter, and Candidatus Profftella (Fig. 1b and Supplementary 2B).
In regard to bacterial species, 1,254 species in total were identified for both groups, out of which 1,205 were common for both groups. The species identified with 100% frequency and abundance (150 to 4,000 reads) were Aggregatibacter aphrophilus, Capnocytophaga ochracea, Fusobacterium nucleatum, Haemophilus parainfluenzae, Leptotrichia buccalis, Neisseria gonorrhoeae, Neisseria meningitidis, Porphyromonas gingivalis, Prevotella dentalis, Prevotella denticola, Prevotella intermedia, Prevotella melaninogenica, Prevotella sp oral taxon 299, Selenomonas sputigena, Streptococcus cristatus, Streptococcus gordonii, Streptococcus mitis, Streptococcus oralis, Streptococcus parasanguinis, Streptococcus pneumoniae, Streptococcus pseudopneumoniae, Streptococcus sanguinis, Tannerella forsythia, and Veillonella parvula. Moreover, 21 bacterial species were exclusively observed in dental plaque of patients with OSCC and the more frequent (30 to 50%) were Pseudomonas monteilii, Coprothermobacter proteolyticus, Bordetella parapertussis, and Klebsiella variicola. Last, the remaining 28 bacterial species were presented only in controls, among which the most frequent (30 to 40%) and abundant were Edwardsiella tarda, Sulfurihydrogenibium azorense, Desulfocapsasulfexigens, Ehrlichia ruminantium, and Rickettsia africae (Fig. 1c and Supplementary 2C).
3.4. Bacterial Microbiota Present in the Saliva of OSCC Patients and Controls: Intergroup Comparisons
A comparison of saliva bacteriome was made between patients with OSCC and controls. Total 30 phyla were identified in patients with OSCC and controls. The most frequent (90 to 100%) and abundant (180 to 6,300 reads) were Bacteroidetes, Fusobacteria, Spirochaetes, Proteobacteria, Actinobacteria, Tenericutes, and Firmicutes (Fig. 1a and Supplementary 3A). Out of the 30 phyla, both populations shared 28. Only the Chrysiogenetes and Armatimonadetes phyla were present in the saliva from patients with OSCC. At the genus level, a total of 596 were identified, including both patients with OSCC and controls, where 535 were present in both groups (Fig. 2b). The most frequent (90 to 100%) and abundant (200 to 3,800 reads) were Fusobacterium, Haemophilus, Porphyromonas, Prevotella, Selenomonas, Streptococcus, Veillonella and Rothia. In regard to the 61 genera that were not shared, 41 of them were only present in saliva from controls, among which the most frequent and abundant were Acidothermus, Candidatus Desulforudis, Candidatus Solibacter, Coraliomargarita, Frateuria, Mesoplasma, Parvularcula, and Simiduia. The remaining 20 genera were present only in the saliva from patients with OSCC, among which the most frequent and abundant were Chthonomonas and Desulfobacula (Supplementary 3B).
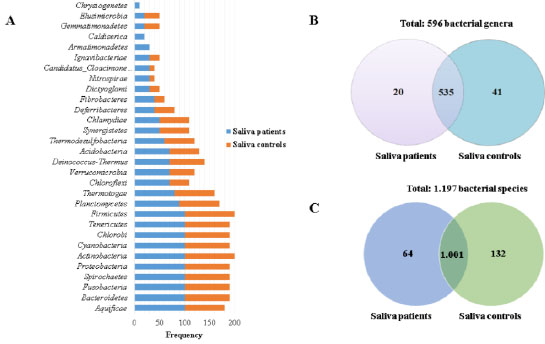
Concerning species of bacteria, a total of 1,197 were identified in the saliva from patients with OSCC and controls, of which 1,001 were determined in both groups (controls and patients with OSCC). Among the most frequent (90 to 100%) and abundant (240 to 1,430 reads) species were Streptococcus oralis, Streptococcus pneumoniae, Veillonella parvula, Streptococcus parasanguinis, Streptococcus pseudopneumoniae, Neisseria meningitidis, Porphyromonas gingivalis, Prevotella melaninogenica, Streptococcus gordonii, Streptococcus mitis, Streptococcus salivarius, Streptococcus sanguinis, Streptococcus sp I-P16, Rothia dentocariosa, Rothia mucilaginosa. Out of the 196 species, which were not common for both groups, 132 were present in saliva from controls, where the most frequent (40 to 50%) were Enterobacter asburiae, Marinobacter hydrocarbonoclasticus, Mycobacterium chubuense, Mycoplasma penetrans, Acidovorax avenae, Listeria ivanovii, and Mesoplasma florum. The remaining 64 bacteria species were identified only in saliva from patients with SCC, and the most frequent (30 to 40%) and abundant were Dehalococcoides mccartyi, Pseudoalteromonas atlantica, Chthonomonascalidi rosea, Desulfobaculatoluolica, Nostoc punctiforme, and Secondary endosymbiont of Heteropsylla cubana (Fig. 2c and Supplementary 3C).
3.5. Bacterial Microbiota in Tumor Tissue, Dental Plaque, and Saliva of Patients with OSCC: Intragroup Comparisons
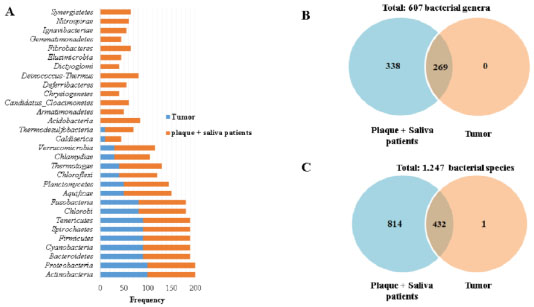
Furthermore, to complement the bacterial microbiome study, intragroup comparisons were made to identify specific differences and similarities; 17 phyla were observed in tumor tissue, dental plaque, and saliva from patients with OSCC, and the most frequent (90 to 100%) were (Actinobacteria, Proteobacteria, Bacteroidetes, Firmicutes, Spirochaetes, and Tenericutes). In addition, 13 phyla were exclusively observed in plaque and saliva from patients with OSCC (Acidobacteria, Armatimonadetes, Candidatus Cloacimonetes, Chrysiogenetes, Deferribacteres, Deinococcus-Thermus, Dictyoglomi, Elusimicrobia, Fibrobacteres, Gemmatimonadetes, Ignavibacteriae, Nitrospirae, and Synergistetes) with frequencies ranging from 40 to 85% (Fig. 3a and Supplementary 4A)
In regard to genus, for all three microenvironments, 607 genera were identified in total, tumor tissue did not present an exclusive genus, and out of the 607 genera, 269 were common to all three microenvironments (Fig. 3b). The most frequent (80 to 100%) were Bacteroides, Mycoplasma, Streptococcus, Fusobacterium, Haemophilus, Prevotella, and Veillonella. On the other hand, 338 genera were only present in plaque and saliva samples, principally by their frequency (>70%), the genera were Clostridioides, Finegoldia, Halanaerobium, Pasteurella, Arcobacter, Paludibacter, Basfia, Bernardetia, Desulfitobacterium, Desulfovibrio, Haliscomenobacter, Rhodococcus, Ruminiclostridium, Akkermansia, Carnobacterium, Desulfotomaculum, Geobacter, Mageeibacillus, Agrobacterium, Arcanobacterium, Clavibacter, Deinococcus, Eggerthella, Enterobacter, Erysipelothrix, Ferrimonas, Gordonia, Laribacter, Leifsonia, Leptothrix, Methylovorus, Micrococcus, Paraburkholderia, Propionibacterium, Pusillimonas, Sideroxydans, Thiomonas, and Xylanimonas (Fig. 3b and Supplementary 4B).
In the three microenvironments, a total of 1,247 bacterial species were identified. In tumor tissue, 433 bacterial species were found, 432 of which were also present in dental plaque and saliva in patients with OSCC. Only one species Rickettsia slovaca was exclusively present with a 10% frequency in tumor tissue (Fig. 3c and Supplementary 4C). Out of the 432 common species, the ones identified with higher frequency (80 to 90%) were Bacteroides helcogenes, Chamaesiphon minutus, Verminephrobacter eiseniae, Mycoplasma conjunctivae, Anabaena sp9, Bacteroides fragilis, Candidatus Azobacteroides pseudotrichonymphae, Cardinium endosymbiont of Encarsia pergandiella, Chlorobium phaeobacteroides, Escherichia coli, Flavobacterium psychrophilum, Fusobacterium nucleatum, Mycoplasma hyopneumoniae, Riemerella anatipestifer, Veillonella parvula, Yersinia enterocolitica, Lawsonia intracellularis, Oscillatoria acuminata, Photobacterium profundum, and Borrelia crocidurae (Supplementary 4C).
On the other hand, out of the 1,247 species common to the three microenvironments, 814 bacterial species were present only in dental plaque and saliva of patients with OSCC. Thirteen species were observed with higher frequency (80 to 95%) and abundance (39 to 75 reads (Bacteroides salanitronis, Streptococcus salivarius, Streptococcus spI-P16, Streptococcus spI-G2) (Fig. 3c and Supplementary 4C).
3.6. Intergroup Comparison between Tumor Tissue Bacterial Microbiota and Microbiota from Dental Plaque and Saliva in Individuals Without OSCC
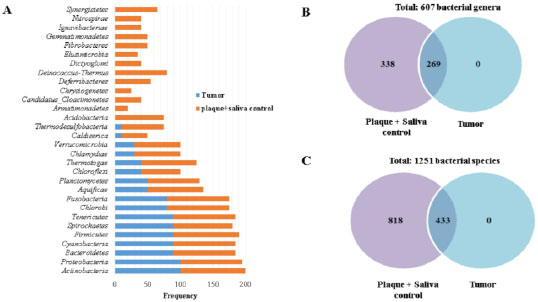
As previously described in tumor tissue, 17 phyla were identified, among which the following had the highest frequency (90 to 100%): Actinobacteria, Proteobacteria, Bacteroidetes, Cyanobacteria, Firmicutes, Spirochaetes, Tenericutes, Chlorobi, and Fusobacteria (Fig. 4a and Supplementary 5A). These 17 phyla from tumor tissue were also observed in isolates from dental plaque and saliva in controls. Additionally, the 9 phyla more frequently (from 90 to 100%) found in dental plaque and saliva in controls coincide with the 9 phyla more frequently (from 80 to 100%) found in tumor tissue (Actinobacteria, Proteobacteria, Bacteroidetes, Firmicutes, Spirochaetes, Tenericutes, Chlorobi, and Fusobacteria). However, in addition to the 17 phyla common to tumor tissue, in dental plaque and saliva group in controls, the following phyla were exclusively found (Acidobacteria, Armatimonadetes, Candidatus Cloacimonetes, Chrysiogenetes, Deferribacteres, Deinococcus-Thermus, Dictyoglomi, Elusimicrobia, Fibrobacteres, Gemmatimonadetes, Ignavibacteriae, Nitrospirae, and Synergistetes) with frequencies ranging from 20 to 80% (Supplementary 5A).
Tumor tissue did not present any exclusive genus, and 607 genera in total were present in all three microenvironments. Out of the 607 genera, 269 were common to all three microenvironments (Fig. 4b and Supplementary 5B). The genera most frequently observed (80% to 90%) were Bacillus, Bacteroides, Borrelia, Mycoplasma, Streptococcus, Verminephrobacter, Acinetobacter, Anabaena, Candidatusazo bacteroides, Candidatus cardinium, Chlorobium, Escherichia, Flavobacterium, Fusobacterium, Haemophilus, Lawsonia, Occillatoria, Photobacterium, Prevotella, Rimerella, Veillonella, and Yersinia (Supplementary 5B).
On the other hand, 338 genera were present only in the plaque and saliva of controls. The most frequently (>80%) and abundantly observed were Acidovorax, Basfia, Beutenbergia, Carnobacterium, Chromobacterium, Clavibacter, Clostridioides, Desulfovibrio, Enterobacter, Geobacter, Intrasporangium, Isoptericola, Kytococcus, Leifsonia, Marinobacter, Nocardioides, Pasteurella, and Rhodococcus (Fig. 4b and Supplementary 5B). In the three microenvironments, 1,251 bacterial species were identified. In tumor tissue, 433 species were observed, all common to dental plaque and saliva from controls. The most frequent (80 to 90%) species were Mycoplasma fermentans, Bacteroides vulgatus, Chlamydia caviae, Francisella noatunensis, Vibrio anguillarum, Anabaena sp90, Roseburiahominis, Vibrio alginolyticus, Mycoplasma mobile, Carnobacterium sp17-4, Bacteroides thetaiotaomicron, Eubacterium limosum, Chloroherpeton thalassium, Borrelia recurrentis, Legionella pneumophila, Candidatus Blochmannia chromaiodes, gamma proteobacterium HdN1, Owenweeksia hongkongensis, Yersinia pseudotuberculosis, Phycisphaera mikurensis, Vibrio anguillarum, Borrelia recurrentis, and Carnobacterium sp 17-4 (Fig. 4c and Supplementary 5C). In addition, out of the 1,251 identified for the three microenvironments, 818 bacteria were only observed in plaque and saliva from controls (Fig. 4c and Supplementary 5C). Furthermore, the 45 bacterial species more frequently (from 80 to 100%) and also 15 abundantly found (mean reads 200 to 4,000) were Streptococcus oralis, Streptococcus parasanguinis Streptococcus pseudopneumoniae, Streptococcus sanguinis, Prevotella denticola, Streptococcus pneumoniae, Streptococcus gordonii, Streptococcus mitis, Porphyromonas gingivalis, Streptococcus macedonicus, Prevotella melaninogenica, Aggregatibacter aphrophilus, Streptococcus equi, Porphyromonas asaccharolytica, Starkeya novella, Streptococcus agalactiae, Streptococcus salivarius, Leptospira interrogans, Haemophilus parainfluenzae, and Roseburia hominis (Supplementary 5C).
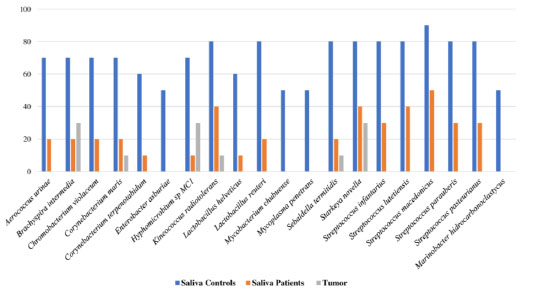
3.7. Bacterial Microbiota Profile Associated with the Processes of Eubiosis and Dysbiosis in Saliva and Dental Plaque Samples
After establishing bacterial profiles in patients with OSCC and controls and determining inter-and intra-group comparisons, we decided to identify bacterial species that were differentially expressed in both populations and that we could associate with eubiosis and dysbiosis states. It is noteworthy to clarify that the terms eubiosis (natural microbiota present and reproducibly detectable in a healthy individual) and dysbiosis (change or imbalance of the natural microbiota and associated disease) have been analyzed extensively and are key terms in human microbiome research, especially when associated with disease states, such as cancer [23].
Regarding species compatible with eubiosis, a screening in saliva and dental plaque of patients with OSCC and controls was performed to search for bacteria that were differential for each group in frequency values from 40 to 60%. Furthermore, frequencies were also compared to those present in tumor tissue. Hence, at least 20 bacterial species were identified in saliva samples that could be compatible with eubiotic processes (p < 0.05) (Aerococcus urinae, Brachyspira intermedia, Chromobacterium violaceum, Corynebacterium maris, Corynebacterium terpenotabidum, Enterobacter asburiae, Hyphomicrobium sp MC1, Kineococcus radiotolerans, Lactobacillus helveticus, Lactobacillus reuteri, Mycobacterium chubuense, Mycoplasma penetrans, Sebaldella termitidis, Starkeya novella, Streptococcus infantarius, Streptococcus lutetiensis, Streptococcus macedonicus, Streptococcus parauberis, Streptococcus pasteurianus, and Marinobacter hydrocarbonoclastycus) (Fig. 5c and Supplementary 6).
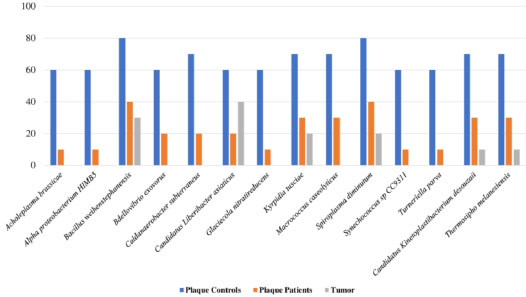
Furthermore, a total of 14 bacterial species were observed in dental plaque samples that could also be associated with eubiotic processes (p < 0.05) (Bacillus weihenstephanensis, Spiroplasma diminutum, Kyrpidia tusciae, Macrococcus caseolyticus, Caldanaerobacter subterraneus, Bdellovibrio exovorus, Candidatus Liberibacter asiaticus, Acholeplasma brassicae, Alpha proteobacterium HIMB5, Glaciecola nitratireducens, Synechococcus sp CC9311, Turneriella parva, Candidatus kinetoplastibacterium desousaii, and Thermosipho melanisiensis) (Fig. 6 and Supplementary 6).
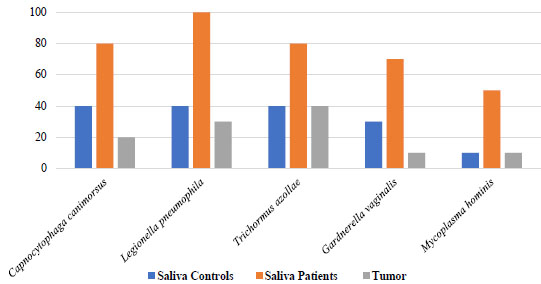
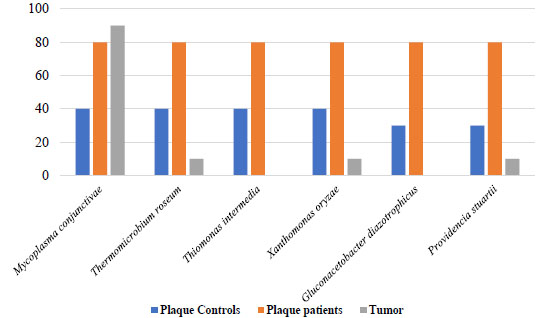
Concerning bacterial compatibility with dysbiosis, a screening was performed in saliva and dental plaque samples from OSCC patients and controls to search for bacterial species that could be differentially observed in each group in frequency values ranging from 40% to 60% compared to the frequencies present in tumor tissue. In this search, five bacterial species were identified (Legionella pneumophila, Mycoplasma hominis, Trichormus azollae, Gardnerella vaginalis, and Capnocytophaga canimorsus) (Fig. 7 and Supplementary 6 showed saliva samples compatible with dysbiotic processes and significant differences (p=0.03). In dental plaque, it was observed that six bacterial species were associated with the dysbiosis process (p < 0.05) (Mycoplasma conjunctivae, Thermomicrobium roseum, Thiomonas intermedia, Xanthomonas oryzae, Gluconacetobacter diazotrophicus, and Providencia stuartii) (Fig. 8 and Supplementary 6).
4. DISCUSSION
Describing and structuring oral microbiota is of great interest to understand its interrelationship with cancer, particularly the microbiota related to OSCC development [6, 24, 25]. Hence, microbiota identification with pathogenic potential at the species level is of great importance since it could be associated with oral eubiosis and dysbiosis states and the onset and progression of OSCC [26]. The majority of studies on the oral microbiota have focused on 16S rRNA sequencing. However, many limitations related to the sensitivity, depth, and robustness of the technique are encountered. In the past years' development and implementation of high throughput sequencing techniques to identify whole genomes has allowed for advancements in microbiome study and the identification of bacteria at the species levels in different pathogenic processes [12, 27, 28]
Our study aimed at identifying the microbiota present in different oral microenvironments of patients with OSCC and control. In both groups, great variability in microbiota was observed, represented by a total of 30 phyla (shared by both groups) and 611 genera (334 shared by both groups, 273 only in OSCC patients, and 4 only in controls). Additionally, 1,269 species were also observed (1,231 were shared by both the groups, 17 were only found in patients with OSCC and 21 only in controls).
Part of the findings herein observed coincide with those found in other studies, since Bacteroidetes, Firmicutes, Proteobacteria, Fusobacteria, and Actinobacteria phyla have been more frequently identified in patients with oral cancer [11, 29, 30], esophageal squamous cell carcinoma [31], head and neck squamous cell carcinoma [19], precursor epithelial lesions (dysplasias, hyperplasias, hyperkeratosis), tumor tissue [20, 32], and contralateral healthy tissues in patients with oral cancer [32] with the exception of Spirochaetes. Additionally, Guerrero Preston and collaborators in 2016 identified a greater frequency of Firmicutes, Bacteroidetes, and Proteobacteria in saliva from healthy individuals, whereas in head and neck squamous cell carcinoma, Bacteroidetes and Proteobacteria were decreased, and Firmicutes increased [19]. Another oral squamous cell carcinoma study performed on 16S rRNA identified 11 phyla in greater frequency Proteobacteria, Bacteroidetes, Firmicutes, Fusobacteria, and Actinobacteria, and Tenericutes, Synergistetes, Saccharibacteria, Spirochaetes, SRI, and Gracilibacteria with less frequency [33].
On the other hand, other studies have identified the phyla Saccharibacteria, Spirochaetes, Tenericutes, Cyanobacteria, Synergistetes, Absconditabacteria, Thermi, Gracilibacteria, Chloroflexi, Armatimonadetes, Omnitrophica, and Verrumicrobiota with greater frequency and abundance in saliva from patients with oropharyngeal squamous cell carcinoma [33, 34] and esophagus of healthy individuals [35]. In the present work, 13 phyla were found in plaque and saliva from patients with OSCC and controls but were completely absent in tumor tissue.
In contrast, most oral microbiotas focused on 16S rRNA research have reported a total of 12 bacterial phyla in oropharyngeal squamous cell carcinoma samples, dysplastic tissue, and healthy tissue, where the most frequent were Bacteroidetes, Firmicutes, Proteobacteria, Fusobacteria, and Actinobacteria [31]. It is important to highlight that the phylum Armatimonadetes was identified in a lesser proportion in saliva from patients with OSCC and healthy controls [34], and the Phylum Synergistetes was only identified in OSCC biopsies [33]. In samples isolated from the tongue, supragingival plaque, and esophagus from healthy individuals [35], 27 phyla were identified in the oral cavity and esophagus, where the most frequent were Proteobacteria, Firmicutes, Bacteroidetes, Actinobacteria, Fusobacteria, and Saccharibacteria [35]. Brian et al. in 2014 identified 12 phyla of Firmicutes, Bacteroidetes, Proteobacteria, Fusobacteria, and Actinobacteria with greater frequency and abundance in cancer samples and healthy tissue [32]. Following, Wolf A and collaborators in 2017 identified 17 phyla in saliva from patients with oral squamous cell carcinoma and healthy individuals, among which the following phyla were observed in a higher frequency: Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria. In contrast, less frequent were Fusobacteria, Spiroquetas, Saccharibacterias, Tenericutes, Cianobacterias, Sinergistetes, Absconditabacteri, Thermi, Gracilibacteria, Cloroflexi, Armatimonadetes, Omnitrophica, and Verrucomicrobia [34]. Collectively, most findings point out that the most frequent phyla in oral squamous cell carcinoma are Firmicutes, Bacteriodetes, Proteobacteria, Actinobacteria, and Fusobacteria.
In contrast to other studies [11, 19, 31, 32, 36], in the present study, no genus was exclusively identified in tumor tissue; the observed 269 genera were common to plaque and saliva in patients with OSCC. Among the genera that have been exclusively reported in tumor tissue are Bacillus in pre-cancerous lesions and oral cancer [20], Haemophilus, Lactobacillus, Veillonella, Citrobacter, Neisseraceae, and Kingella in tumor tissue [29, 37], Streptococcus, Prevotella, Haemophilus, and Veillonella in patients with OSCC [36] and saliva from patients with esophageal OSCC [22].
On the other hand, in the present study, 338 genera were found in plaque and saliva microenvironment from controls, which were not identified in tumor tissue; conversely, investigations conducted on samples of healthy patients found the genera Streptococcus, Neisseria, Actinobacillus, and Veillonella in the oral cavity and esophagus of healthy people, Flavobacteria in plaque, Bacteroides in saliva and Clostridia in the back of the tongue of healthy people [35].
Multiple studies have described microbiota at the genus level in different types of carcinomas. Cheng et al. [31] identified 44 genera in total and described that the following genera were increased in healthy tissue, oropharyngeal OSCC, and dysplasia tissue: Megasphaera, Aggregatibacter, Atopobium, Lautropia, Actinobacillus, Bulleidia, Catonella, Filifactor, Corynebacterium, TG5, Acholeplasma, Moryella, Butyrivibrio, Dialister, Peptococcus, and Cardiobacterium. Conversely, Al-Hebshi [33], in a study involving three patients with oral OSCC, identified 78 genera, where the most frequent and abundant genera were Haemophilus, Neisseria, Prevotella, Fusobacteria, Streptococcus, Porphyromonas, Leptotrichia, and Aggregatibacter [33]. Guerrero Preston et al. [19] identified 120 genera in saliva samples, where the most frequent were Streptococcus and Prevotella. The genus Lactobacillus was more frequent in head and neck squamous cell carcinoma compared to controls. On the other hand, the genus Neisseria was more abundant in oropharyngeal squamous cell carcinoma than controls, whereas Citrobacter was more frequent in oropharyngeal squamous cell carcinoma [19].
Wolf et al. [34] studied samples of saliva from patients with oropharyngeal squamous cell carcinoma, OSCC, and healthy individuals and found the genera Streptococcus, Prevotella, and Rothia with greater frequency in the general population. Furthermore, the most abundant in healthy controls were Prevotella, Haemophilus, Neisseria, Streptococcus, and Veilonella, and the most abundant in patients with cancer were Actinomyces, Schwartzia, Treponema, and Selenomonas [34]. Moreover, Dong et al. [35] in oral niches (saliva, the dorsum of the tongue, and supragingival plaque) and different locations of the esophagus identified 365 genera from the oral cavity and 594 from the esophagus. The most frequent and abundant were Streptococcus, Neisseria, Prevotella, Actinobacillus, and Veillonella [35]. All the previous studies regarding the description of genera have indicated a greater diversity in relation to phyla. Among the most frequent in squamous cell carcinoma samples were Megasphaera, Aggregatibacter, Atopobium, Lautropia, Actinobacillus, Bulleidia, Catonella, Filifactor, Corynebacterium, Synergistetes, Acholeplasma, Moryella, Butyrivibrio, Dialister, Peptococcus, Cardiobacterium, Haemophilus, Neisseria, Prevotella, Fusobacteria, Streptococcus, Porphyromonas, Leptotrichia, Actinomyces, Schwartzia, Treponema, Selenomonas, Veillonella, Lactobacillus, and Citrobacter.
In general, the consensus points out that the human oral microbiome is composed of more than 700 bacterial species, distributed in oral microenvironments, such as teeth, cheeks, tongue, hard and soft palate, gingival sulcus, and tonsils [38]. However, the qualitative and quantitative microbial species varies according to age, gender, oral health, nutritional habits, alcohol, and cigarette consumption, among others.
In the only study performed to determine bacterial species, Al–Hebshi et al. [33] identified 228 bacterial species in a total of three patients with OSCC, of which 35 were shared in all samples [33]; the most abundant were Prevotella oris, Neisseria flava, N. flavescens/subflava, Fusobacterium nucleatum ss polymorphum, Aggregatibacter segnis, Streptococcus mitis, Fusobacterium periodonticum, Neisseria elongata, Porphyromonas sp. oral taxon 279, and Alloprevotella tannerae.
In contrast to Al-Hebshi, in the present study, we identified 1,254 species, in which 1,230 were common to plaque, saliva, and tumor in patients with OSCC and plaque and saliva in healthy controls. Also striking is the non-coincidence observed in the bacterial composition in the 10 most frequent species in plaque and saliva of patients with OSCC in relation to the 10 most frequent in plaque and saliva of controls.
The term biomarker refers to the study and identification of a biological molecule produced by cells, tissue, bacteria, virus, or other live organisms that can be found in blood, body fluids, tissues, and multiple samples [14]. It can be used as an indicator or marker of a normal or abnormal process, condition, or disease; thus, it becomes very useful in clinical practice.
In regard to bacteria as possible biomarkers, various metagenomic studies have addressed microbial communities in humans, making it possible to suggest certain bacteria as biomarkers for clinical and microbiological diagnosis associated with health and disease states [39].
For the oral cavity, some studies have proposed a number of bacterial species associated with periodontal disease [40]. It has been suggested that oropharyngeal cancer can be predicted by changes in the oral microbiota, depending on the microenvironment found, and that differences in bacteria can be associated with cancer pathogenesis [41].
The search and identification of bacterial biomarkers could be very useful in monitoring the course of certain types of cancerand designing strategies for their prevention and control. It has been proposed that microorganisms differentially associated with oral cancer could be used as biomarkers of pathological or healthy states [15].
Concerning compatible health and bacterial disease profiles, the present study was capable of identifying a profile composed of 45 bacterial species (34 eubiotic and 11 dysbiotic). In regard to the eubiotic population, the 34 bacterial species are important not only for their role in the physiology of oral ecology but also because they are differentially separated by high frequency and statistically significant differences in saliva and plaque from healthy individuals compared to patients with OSCC. Therefore, they become possible candidates to be detected in combined or multiple PCR amplification systems. Additionally, they can serve as oral health markers by obtaining isolates that can be cultured from saliva or plaque from controls.
It is notable the 34 bacterial species compatible with eubiotic processes were found at low frequencies in saliva and plaque samples in OSCC patients, as well as tumor samples. On the other hand, the 11 bacterial species compatible with dysbiotic processes were only present in high frequencies in saliva and dental plaque samples of OSCC patients. Within this context, these results suggest that detection of bacteria compatible with eubiotic and dysbiotic processes should be carried out particularly in saliva and dental plaque samples and not tumor samples, where all bacteria were present in low frequencies. Therefore, this could become a useful tool to take into account for prevention and OSCC follow-up.
Of the group with 34 species with eubiosis, more than nine bacteria could be suggested as possible biomarkers (Aerococcusurinae, Brachyspira intermedia, Chromobacterium violaceum, Enterobacter asburiae, Lactobacillus helveticus, Mycobacterium chubuense, Mycoplasma penetrans, Streptococcus infantarius, and Streptococcus pasteurianus). They are differentially distinct by 50% in frequency between saliva and plaque samples in controls from those with OSCC.
In regard to dysbiosis, the 11 bacterial species are important because they are differentially separated by high frequency in saliva and plaque from OSCC patients from control individuals. Thus, they become possible candidates in the search for molecular and culture methods from saliva and plaque isolates in control individuals to establish oral pathology markers. It is important to specify that these groups of bacteria are in line with the present work findings, observed in high frequencies in the tumor, where they colonize and remain in saliva and plaque from OSCC patients. Additionally, they were found in high frequencies, suggesting the continuous presence of these types of samples in OSCC patients. From the 11 species associated with dysbiosis, six bacteria could be suggested as possible biomarkers (Legionella pneumophila, Mycoplasma hominis, Gardnerella vaginalis, Capnocytophaga canimorsus, Mycoplasma conjunctivae, and Providencia stuartii). They were differentially disparate in high frequencies and statistically significant differences between samples from saliva and plaque from healthy individuals without OSCC compared to samples from OSCC patients. Additionally, they have recognized importance in human ecology and pathology.
Present work data that could be used as possible bacterial biomarkers in eubiosis and dysbiosis states associated with oral cancer must be yet validated in other oral cancer studies. Hence, it is imperative to shed light on bacterial molecular, cellular, and pathological mechanisms that impact tissues of the oral cavity. In addition, because many of these bacteria have not yet been identified or studied in cancer.
Differences observed between our data and other studies could be mainly associated with ecological determinants of each microenvironment, factors associated with the host’s immunity, oral hygiene, nutritional habits, hormone changes, medication intake, alteration in the mucosa and dental tissues, among others [18]. On the other hand, the technique used to identify the microbiota also plays an important role since most researchers have used the 16S rRNA genes, in contrast to the present work, which used high throughput sequencing.
CONCLUSION
In this study, we have identified by next-generation sequencing the bacterial microbiome present in different oral environments in patients with OSCC (tumor tissue, dental plaque, and saliva) and matched controls without OSCC (saliva and dental plaque). Our data revealed a specific bacterial species associated with OSCC and identified 34 and 11 bacterial species, respectively, associated with health and disease states.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by the ethics committee of the Pontifical Xavierian University, Bogotá D.C., Colombia with approval number OD-0149.
HUMAN AND ANIMAL RIGHTS
No animals were used in this research. All human research procedures followed were in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Written and informed consent was obtained from all the participants involved.
AVAILABILITY OF DATA AND MATERIALS
The datasets used during the present study are available from the corresponding author [A.E.] on reasonable request.
FUNDING
This study was supported by the Pontificia Universidad Javeriana, Bogotá, in terms of funding the following projects; 1. “Identification of biomarkers involved in the progression or regression of HPV-associated cancer, through tissue samples for histopathological study, in the city of Bogotá ” (Grant ID 7736) and 2.“ Integrative analysis of the microbiome and transcriptome present in dental plaque, saliva and tumor tissue in oral carcinoma and its relationship with the microbial metabolome determined in saliva ” (Grant ID 7745).
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors express their gratitude to the Pontificia Universidad Javeriana, Bogotá, for funding the present study and Universidad de Los Andes for the data analysis collaboration.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publishers web site along with the published article.


