All published articles of this journal are available on ScienceDirect.
The Influence of Lactobacillus Acidophilus on MUC1, GAL-3, IL-1β and IL-17 Gene Expression in BALB/c Mice Stomach
Abstract
Background and Objective:
Lactobacillus acidophilus has been widely used for the management of gastrointestinal carcinoma owing to its immunomodulation effect; however, the role of L. acidophilus and its specific mechanism of action in the stomach is not fully comprehended. The present study evaluated the expression profile of MUC-1, GAL-3, IL -1β, and IL-17 in the L. acidophilus treated mice stomach.
Methods:
The study was conducted utilizing three groups of mice, 6 mice for each group, administered with different doses of L. acidophilus and a control group treated with normal saline. The results were analyzed with the Mann-Whitney Test.
Results:
The results demonstrated that L. acidophilus elevated IL-1β insignificantly and inhibited the expression of IL-17. The MUC-1 expression is influenced by L. acidophilus and inversely proportional to GAL-3 expression.
Conclusion:
Lactobacillus acidophilus plays a prominent role against inflammatory responses and has a potential in the treatment of gastrointestinal cancer.
1. INTRODUCTION
Probiotics are beneficial microorganisms that, when administered in adequate doses, provide a health benefit to humans [1]. Probiotics modulate the gastrointestinal tract by boosting the immune response and have a positive impact on gastrointestinal microflora [2]. Probiotics are known for their potential immunomodulatory properties and are often used for the therapeutic management of cancer [3].
Lactobacillus spp. are gram-positive, rod or coccobacilli shaped, non-spore forming and catalase-negative [4]. The bacteria adhere to the mucosal layer and epithelial surface of the upper gastrointestinal tract [4]. The in-vitro and in-vivo analyses revealed that Lactobacillus spp. reduced the concentration of pro-inflammatory cytokines [3]. Lactobacillus spp. inhibit mutagenic compounds, which are associated with carcinogenesis and harmful enzymes [3]. Lactobacillus spp. interact with the epithelial cells through several mechanistic pathways that could modulate the host immune system, improve barrier integrity, and stimulate anti-inflammatory immune response [5]. An in-vivo study using the rat model demonstrated that L. acidophilus downregulated not only tumor incidence and tumor volume but also tumor multiplicity [3]. However, the specific mechanism of action of L. acidophilus has not been completely elucidated.
Gastric cancer is the second leading cause of cancer-related death because of its high incidence, poor prognosis, and high mortality rate, particularly in developing countries [6]. Environmental condition plays an essential role in the manifestation of gastric cancer [6]. The development of gastric carcinoma is closely associated with chronic inflammation [7]. Dietary factors and H. pylori infection act collectively and are believed to cause gastric carcinogenesis [6, 8]. H. pylori induce chronic inflammation that contributes to the development of carcinogenesis [9, 10].
MUC1 is membrane-bound mucin expressed on gastric mucosa and is known to limit pathogen infection and colonization [11]. MUC1 plays a role in counter regulate gastric inflammatory response to the pathogenic invasion of microbes [11]. MUC1 regulates inflammation during infection by modulating the NLRP3 inflammasome complex and as a negative regulator of TLR signaling.11 Gastric MUC1 forms a protective barrier that restricts the acute and chronic inflammation caused by colonization of H. pylori thereby impeding the infection [12].
Galectin-3 (Gal-3) is a β-galactoside-binding-lectin protein that is widely expressed in epithelial mucosa [13]. It is known to have pro-inflammatory activity and is involved in the recruitment of the immune cell [13]. Galectin-3 contributes to the adhesion of bacterial and tumor cells, inflammation, wound healing process, and immunity [14, 15]. Galectin-3, which can recognize PAMP (pathogen-associated molecular patterns) allows this glycan-binding protein to recognize pathogenic microorganisms and induces pro-inflammatory activity [16]. An increased concentration of Gal-3 expression is associated with the occurrence of gastric adenocarcinoma and other malignancies [17]. Little is known about Gal-3 interaction with commensal bacteria.13
Interleukin-17 is a pro-inflammatory cytokine and is responsible for immune cell recruitment that helps in microbial elimination during infection. IL-17 stimulates cytokine produc- tion that induced inflammation response [18]. A previous study revealed that a high level of IL-17 is associated with tumor progression [19]. Therefore, it is believed that IL-17 plays a major role in the progression of gastric carcinogenesis.
The aim of the present study was to determine the MUC1, Gal-3, IL-1β, and IL-17 gene expression in the L. acidophilus treated mice stomach.
2. MATERIALS AND METHODS
2.1. Bacterial Strain and Culture Conditions
L. acidophilus (ATCC 4356) was purchased from MicrobiologicsTM Fisher Scientific, USA and is abbreviated as LA. Bacterial cultivation was carried out using MRS broth (Oxoid, CMO359) at 37 ºC in microaerophilic condition comprising of 5% O2, 10% CO2, 85% N2 (Campygen CN0035). The confirmation test to Lactobacillus acidophilus is determined with gram stain, carbohydrate fermentation, indol test and catalase test. Further, it was incubated for 24 h on MRS Agar and was harvested in 0.85% Na Cl. Three doses were prepared with a different bacterial density which was determined through a spectrophotometer (Thermo Spectro- photometer) at 600 nm and confirmed with plate count method. The bacterial concentrations in the low dose, moderate dose, and high dose were approximately 106 CFU/mL (OD 0.3), 107 CFU/mL (OD 0.6), and 108 CFU/mL (OD 1.0), respectively. For long-term storage, L. acidophilus is maintained in MRS broth containing 20% glycerol at –80 ºC.
2.2. Animal Study
24 BALB/c male mice were used in the study who were 7 weeks old, obtained from PT BIOFARMA, Bandung, Indo- nesia. Mice were kept in normal conditions of temperature and light and had access to food and water ad libitum. After seven days of the adaptation period, mice were divided into four groups, 6 mice for each group, namely the control group administered with normal saline (LA0), low-dose LA group (LA1), moderate dose LA group (LA2), and high-dose LA group (LA3). Mice were administered for 14 days with 0.5 mL LA suspension by orogastric gavage. Normal saline was given to LA0 group. On the 15th day, all mice were anesthetized with ketamine and sacrificed. Gastric tissues were collected and stored in RNAlater® tubes (Sigma, R0901) at –80 ºC.
2.3. RNA Isolation
RNA extraction was done by homogenizing the gastric tissue using TRIzol Reagent (Invitrogen, USA). The extract was then centrifuged at 14000 ×g in a column tube following the manufacturer's protocol. The RNA pellet was collected and purified using phenol-chloroform extraction and ethanol precipitation and was further dissolved in TAE buffer pH 8.0. The quantification of RNA was done using Nanodrop spectrophotometer ND-100 (Thermo Scientific, USA).
2.4. Real-Time PCR (qRT - PCR)
One-step qRT-PCR kit (Kapa Biosystem, USA) was used to study the mRNA expression. Primers were designed using forward and reverse sequences, as shown in Table 1. Primer designs were determined by Integrated DNA Technologies (IDT) Reference Sequences on MUC1 (IDT DNA Ref No 102171764-102171765), GAL-3 (IDT DNA Ref No 102171768-102171769), IL-1β (IDT DNA Ref No 102171770-102171771), IL-17 (IDT DNA Ref No 102171772-102171773) and GAPDH (IDT DNA Ref No 102389365-102389366). The qPCR reaction was preceded by reverse transcription at 42 ºC for 5 min followed by enzyme inactivation of reverse transcriptase (RT) at 95ºC for 3 min. The PCR process was repeated for 40 cycles. The PCR requires temperature cycling for different stages of the reaction to occur. The denaturation of the dsDNA was done at 95 ºC for 30 s followed by annealing of the primer at 60 ºC for 20 s and extension was carried out at 72ºC for 20 s. The results were presented as Ct value. The average Ct value was calculated for the targeted genes and the internal control gene (GAPDH). The ΔCt was obtained from the difference in threshold cycles for targeted genes and GAPDH. Gene expression was concluded from 2- ΔΔCt [20].
2.5. Statistical Analysis
The experimental data were documented as mean ±SD. The data were statistically analyzed by Mann Whitney test. The differences were considered statistically significant at p < 0.05.
| Gene | Forward Primer (5’-3’) | Reverse Primer (5’-3’) |
|---|---|---|
| MUC-1 | CTT CCT GCT GCT ACT TCT AG | TGG TCG ATG CTA AGG AA |
| GAL-3 | CGC ATG CTG ATC ACA AT | GTC CTG CTT CGT ACA CA |
| IL-1β | CCA TTG ACC ATC TCT CTC TG | ACT CAG TCT CTT CTT CA |
| IL-17 | GAG CTT CAT CTG TGT CTC TG | GAG GTT GAC CTT CAC ATT CT |
| GAPDH | TCA AGA TGG TGA AGC AG | ATG TAG GCC ATG AGG TCC AC |
3. RESULTS
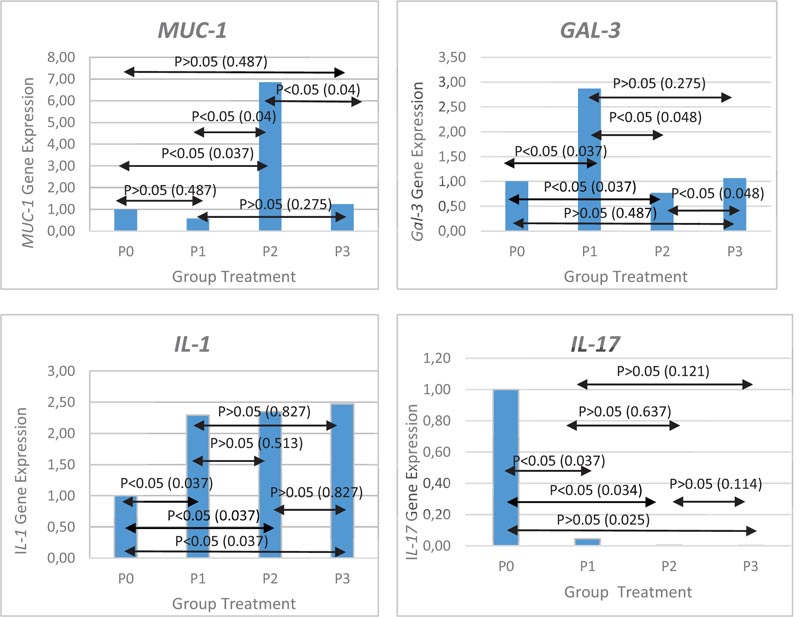
Administration of L. acidophilus at low moderate, and high concentration caused a 2-fold increase in IL-1β gene expression when compared with control (p <0.05), but the result did not show any significant difference. IL-1 gene expression were 2.29 ± 1.12 for low concentration, 2.35 ± 2,41 for moderate concentration and 2.51 ± 1.27 at high concentration. Administration of L. acidophilus at low, moderate and high concentrations decreased IL-17 gene expression compared to the control group (p <0.05) but was not found significant. IL-17 gene expression were 0.04 ± 0.05 for low concentration, 0.02 ± 0.02 for moderate concentration and 0.00 ± o.o for high concentration. MUC-1 gene expression after L. acidophilus administration revealed that it decreases at low concentrations (0.57 ± 0.63 times), gets slightly increased at high concentrations (1.24 ± 0.86 times), but an increase in the concentration of about 6.85 ± 2.65 times was noticed at moderate concentrations (p <0.05). We noticed that Gal-3 gene expression increase at low concentration (2.87 ± 1.28 times), decrease at a moderate concentration (0.77 ± 0.14 times) and slightly increased at high concentration (1.07 ± 0.04 times). The increased expression of mucin after administrating L. acidophilus at a certain concentration results in low Gal-3 expression and vice versa. Gene expression after administration of L. acidophilus is shown in Fig. (1).
4. DISCUSSION
The variation in L. acidophilus concentration was intended to determine whether the difference in the probiotic concentration causes different effects on the gene expression of MUC-1, GAL-3, IL-1β, and IL-17. The probiotic concentration that gives the best effect is expected to be used as a reference dose for the management of diseases.
From the result of IL-1β gene expression, L. acidophilus administration caused a 2-fold increase in IL-1β gene expression when compared with control (p <0.05). Jiang et al. showed that L. acidophillus induced IL-1 production [21]. In contrast to L. acidophilus and L. sakei, L johnsonni did not induce IL-1β mRNA expression in studies with Caco2-cells [22]. We noticed that probiotic strains will induce different proinflammation cytokines. Induction of pro-inflammatory cytokines in probiotic administration is likely to occur because probiotics and commensal bacteria have PAMP (pathogen-associated molecular patterns) recognized by PRRs (pattern-recognition receptors) expressed in immune cells [23]. The administration of L. acidophilus increases bacterial concen- tration, followed by the increase of IL-1β gene expression (p>0.05). The absence of a significant difference between concentration could interpret that the administration of a higher concentration of L. acidophilus will not induce the inflammatory response that will damage the gastric mucosa.
Bacterial infection will increase IL-17 levels and can be found in high levels in chronic inflammation. In this study, we found that L. acidophilus administration at low, moderate and high concentrations decreased IL-17 gene expression compared to the control group (p <0.05). This may be due to the fact that L. acidophilus has the potential to inhibit IL-17 activity. Therefore, L.acidophilus is suggested to play a role in gastric tissue damage that promotes gastric carcinogenesis. Chen et al. confirmed that proinflammatory cytokine IL-17 in L. acidophilus administration may result in the downregulation of IL-23, TGFβ1 and STAT3 [24]. Research conducted by Yang Wu et al. demonstrated that in Histoplasma capsulatum infection, IL-23/17 axis in dendritic cells was negatively regulated by galectin-3.
Interleukin 17 is known to promote cytokine production to induce inflammation such as IL-1,IL-6, TNFα, chemokines and CSFs infection [18, 25]. Based on this study, we also suspected that IL-1β expression was not associated with IL-17.
MUC-1 gene expression after L. acidophilus adminis- tration revealed that it decreases at low concentrations (0.57 times), gets slightly increased at high concentrations (1.24 times), but an increase in the concentration of about 6.85 times was noticed at moderate concentrations (p <0.05). The difference in concentration of L. acidophilus seems to determine the expression of mucin in the stomach. The scientific community is still searching for possible mechanisms associated with probiotic density and mucin expression. Mucin in the gastrointestinal tract plays an important role in microbial attachment [26]. This layer forms the first physical barrier that provides protection to the inflammatory responses resulting from microbial infection, such as Helicobacter pylori [26]. In this study, we found that L. acidophilus induces MUC-1 expression in order to inhibit or reduce the adherence of pathogenic microbes to gastric epithelial cells. Decreased microbial pathogens adherence will reduce tissue damage due to activated proinflammation cytokines. L. acidophilus was known to play a role in regulating and modulating the protection action of intestinal epithelial tight junction in studies with Caco-2 cell [27]. Further research is needed to determine the specific mechanism of action of L. acidophilus in influencing the barrier function in the stomach.
In this study, it is revealed that the expression of GAL-3 seems to be associated with the concentration of L. acidophilus and the MUC-1 expression. The increased expression of mucin after administrating L. acidophilus at a certain concentration results in low Gal-3 expression and vice versa. We are still verifying whether there is a reciprocal relationship between MUC 1 and GAL-3. Piyush et al. demonstrated that galectin-3 and MUC-1 interactions cause EGFR dimerization and activation in epithelial cancer cells [28]. Research conducted by Merlin et al. showed that in pancreatic cancer cell GAL-3 regulated the MUC-1 transcription [29].
Unfortunately, because the material used in this study was gastric tissue, so it was not possible to carry out a cohort study in which the genes expression was determined prior to administration of L. acidophilus and compared their genes expression after treatment.. By knowing the initial gene expression before treatment, the effect of L. acidophilus on the expression of these genes will be more clearly described.
CONCLUSION
L. acidophilus plays an important role in MUC-1, GAL-3 and IL-17 expression. Therefore, it may be suggested that L. acidophilus can be used in the prevention and treatment of tissue damage due to gastric inflammation. A further detailed study on L. acidophilus is needed to understand its specific mechanism of action and its role in stomach immuno- modulation.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study was approved by the Research Ethics Committee of Padjadjaran University, Indonesia with Approval No. 977/ UN6.KEP/EC/2019.
HUMAN AND ANIMAL RIGHTS
No humans were part of this study. This study is performed in accordance with the "Guide for the Care and Use of Laboratory Animals" eighth edition, 2011.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The data supporting the finding of the article is available in the Zenodo Repository at zenodo.org, reference number: https://doi.org/10.5281/zenodo.5006563.
FUNDING
This research was financially supported by the Ministry of Research, Technology and Higher Education of the Republic of Indonesia.
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest.
ACKNOWLEDGEMENTS
The authors are thankful to Dr. Wahyu Widowati from Biomolecular and Biomedical Research Center for laboratory support. The authors thank Wahyu, Dewani and Nurul for techinal assistance.


