All published articles of this journal are available on ScienceDirect.
An Investigation of Honey Bee Viruses Prevalence in Managed Honey Bees (Apis mellifera and Apis cerana) Undergone Colony Decline
Abstract
Objective:
In the absence of known clinical symptoms, viruses were considered to be the most probable key pathogens of honey bee. Therefore, the aim of this study was to investigate the prevalence and distribution of honey bee viruses in managed Apis mellifera and Apis cerana in China.
Methods:
We conducted a screening of 8 honey bee viruses on A. mellifera and A. cerana samples collected from 54 apiaries from 13 provinces in China using RT-PCR.
Results:
We found that the types and numbers of viral species significantly differed between A. mellifera and A. cerana. Black Queen Cell Virus (BQCV), Chronic Bee Paralysis Virus (CBPV), Apis mellifera filamentous virus (AmFV), and Kakugo virus (DWV-A/KV) were the primary viruses found in A. mellifera colonies, whereas Chinese Sacbrood Bee Virus (CSBV) and Sacbrood Bee Virus (SBV) were the primary viruses found in A. cerana. The percentage infection of BQCV and CSBV were 84.6% and 61.6% in all detected samples. We first detected the occurrences of Varroa destructor virus-1 (VDV-1 or DWV-B) and DWV-A/KV in China but not ABPV in both A. mellifera and A. cerana.
Conclusion:
This study showed that BQCV and CSBV are the major threat to investigated A. mellifera and A. cerana colonies.
1. INTRODUCTION
Honey bees are regarded as the most important pollinators for crops, fruits, and vegetables [1]. Wild plants and flowers are pollinated on a large scale in the mountain regions of China [2]. The demand for pollination is increasing in greenhouses owing to the rapid development of modern agriculture, such as strawberries in the winter [3]. However, the number of honey bees available for pollination is not sufficient owing to the large losses suffered by honey bee colonies worldwide [4], of which China is no exception.
Viruses, bacteria, fungi, and parasites were all confirmed to be capable of inducing reductions in colony populations [5-8]. Among these pathogens, more than 20 viruses were identified to infect honey bee. Most of them belonged to the family Picornaviridae and possessed a single-strand positive-sense RNA genome. These viruses were demonstrated to not only have severe impacts on individual bees by damaging their physiology [6], interfering with their productivity [9], and contributing to behavioral disorders [10] but also on the colony growth rate and colony fitness (i.e., a decrease of foraging activity at the colony level [11]).
However, most honey bee viruses cause covert infections without significant clinical symptoms, which is a barrier to their study. A few viruses can be recognized during the late stage of infection by their appearances, such as Chronic Bee Paralysis Virus (CBPV), Black Queen Cell Virus (BQCV), Deformed Wing Virus (DWV), Sacbrood Virus (SBV), and Israeli Acute Paralysis Virus (IAPV). CBPV was found to induce typical characteristics such as trembling, crawling, flightlessness, and even a hairless black abdomen [12]. The typical symptom of DWV were wing deformities, which were often reported to be more pathogenic to honey bee colonies owing to co-infection with Varroa destructor virus-1 (DWV-B) [13]. CSBV and SBV are the only viruses that are pathogenic to larvae (Apis cerana or Apis mellifera); infections cause the larvae to turn pale yellow or dark brown and demonstrate an inability to develop into pupae [14, 15]. A few other viruses have been unintentionally found in diseased honey bee colonies, such as Acute Bee Paralysis Virus (ABPV) and Kashmir Bee Virus (KBV) [16], which are difficult to distinguish by their appearance. Kakugo Virus (DWV-A/KV) is the only virus shown to influence the behavior of honey bees by impacting the brains of aggressive guard worker bees, thereby harming their nervous functions and making the honey bees more aggressive [17]. Because IAPV was initially thought to be closely associated with colony losses [18], the characteristics of IAPV and its infection mechanism were described in detail [7, 19-23].
Nevertheless, little is known about the DNA viruses of honey bees compared to RNA viruses. Recently, the DNA virus Apis mellifera filamentous virus (AmFV) was reported to be associated with crawling bees. The AmFV viral loads were reported to be higher in the bees in spring in Switzerland and demonstrated a significant correlation with RNA viruses in infected honey bees [24]. The full genome of this virus has been published [25].
Owing to the constant discovery of emerging viruses, many investigations have reported the occurrence and prevalence of honey bee viruses as well as other viruses that may infect honey bees, especially since IAPV was found to be associated with Colony Collapse Disorder (CCD) [18]. DWV is the most prevalent virus worldwide. Samples collected from 1999 to 2004 in Hungary were screened for the presence of 6 viruses, and DWV was identified in 72% of the samples, followed by BQCV in 54% [26]. Similarly, DWV was found to be the most common virus in southwest England, followed by ABPV and SBV [27]. DWV was also the most prevalent virus in Austria [28] and France [29]. Several new viruses were also found during the investigation [30]. Runckel et al. [31] performed a large scale investigation on honey bee pathogens that could pose infectious or potential threats to honey bees in the USA and found 4 viruses (ALPV, Big Sioux River virus, LSV 1, and LSV 2) that had not been previously found in honey bee colonies. Metagenomic analysis of viral pathogens of honey bees from Spain showed that the viruses were not limited to ALPV and IAPV but also included turnip ringspot virus and turnip yellow mosaic virus, which originated from plants [32]. Similarly, the queen, which is the other important individual in the honey bee colony, was found to have higher DWV and BQCV titers in the gut than the other tissues during an investigation of the prevalence of 6 viruses in different tissues and feces; in contrast, the SBV and CBPV titers were higher in the hemolymph than in the other tissues [29]. Other bee subspecies and bumblebees were also investigated for the presence of honey bee viruses. For example, BQCV was the most prevalent virus in 23% of savannah honey bees (Apis mellifera scutellata) in South Africa, whereas ABPV, CBPV, DWV, and SBV were not detected [33]. An investigation of hymenopteran RNA viruses in bumblebees found that DWV, BQCV, SBV, and IAPV were the main threats [34]. DWV was recently confirmed to be the most prevalent virus in bumblebees in both laboratory and field experiments [35]. In China, Ai et al. [36] investigated the occurrence of 7 honey bee viruses in 2011 and found that DWV was the most prevalent of the viruses described above. Subsequently, a report that demonstrated the prevalence of 6 viruses in A. cerana in China found that BQCV had an infection rate of 98% in A. cerana [37]. In contrast, Yang et al. [38] found that DWV was the most prevalent among the 7 viruses in China regardless of whether A. mellifera or A. cerana was investigated.
Despite the prevalence of honey bee viruses that have been extensively investigated around the world, including China, several neglected and new honey bee viruses have not been described in China. Because most viruses are persistent and cause harm, it is necessary to widely screen for their presence. These studies will help us understand the prevalence characterization and other factors and their possible associations with colony decline in China. To obtain a broader spectrum of information about honey bee viruses and to elucidate their potential threats to honey bees, we investigated the presence of viruses in A. mellifera and A. cerana bee samples from different provinces that reported a large number of bees crawling in front of the hives. We screened bee samples from 56 apiaries of 13 provinces to identify the presence and distribution of 8 honey bee viruses in China. We found that BQCV and CSBV were the most prevalent in A. mellifera and A. cerana, respectively. Additionally, we discovered that a high percentage of colonies were infected by AmFV and DWV-A/KV in many regions in China for the first time.
2. MATERIALS AND METHODS
2.1. Bee Samples
Honey bees were sampled from 56 apiaries in 13 provinces from March to November, 2015. Worker samples of A. mellifera were collected from 38 apiaries in 8 provinces, and samples of A. cerana were obtained from 18 apiaries in 10 provinces (Table S1). Ten colonies were randomly selected in each apiary per province. Clinical signs of virus infection were not found in these colonies; thus, these colonies were thought to be capable of honey production, although the colonies were reported to have undergone large decline or were identified with crawling bees. All of the above colonies were maintained consistent with guidelines for beekeeping practice manipulation and regularly monitored and treated for mites (only for A. mellifera); no damage due to mite infestation was observed during the previous year. Living bee samples were placed in a small iron wire cage for direct transport or in 50 mL sterile tubes with dry ice for transport to the laboratory at -80°C prior to use.
2.2. DNA and RNA Extraction
All samples were collected, immediately placed in liquid nitrogen, and ground into a fine powder using a pestle and mortar with the corresponding extraction reagent. Briefly, for the detection of Apis mellifera filamentous virus (AmFV), total DNA was isolated from pooled honey bee samples (~50 bees) with the TissuePrep homogenizer (Gening Scientific, Beijing, China). Then, genomic DNA was extracted from the filtrate using the DNA Purification Kit (Promega Corp., Madison, WI, USA), following the manufacturer’s instructions. The resulting DNA was eluted into 20 μL of nuclease-free water and stored at -20°C.
For the other 7 RNA viruses (IAPV, SBV, ABPV, BQCV, CBPV, DWV-B(DWV-B), DWV, CSBV, KBV, and KV(DWV-A/KV), total RNA was extracted from approximately 50 pooled honey bee samples using the TRIzol Kit (Invitrogen, USA) according to the manufacturer’s protocol in the TissuePrep homogenizer (Gening Scientific, Beijing, China). The obtained RNA was dissolved in 20 µL of sterile water and stored at -80 °C prior to analysis. The quantity and purity of the RNA were measured using a Nanodrop spectrophotometer (Thermo Scientific, Beijing, China). The cDNA was synthesized using the M-MLV reverse transcriptase with the oligo dT primer according to the manufacturer’s instructions. The cDNA samples were used as templates for PCR amplification with specific primers corresponding to viral genes.
2.3. PCR Detection
The primer sequences, orientation, and references are provided in Table S2. For the 7 RNA viruses, the initial cycle for reverse transcription was 80°C for 30 min, followed by the PCR cycles. The PCR reaction for AmFV and other 7 RNA viruses consisted of a total 20 µl volume containing 2×GoTaq reaction buffer (Promega, USA), 1 µM of the sense and antisense primers, 1 µL of cDNA or DNA, and nuclease-free water. The cycling conditions were as follows: 1 min at 95°C; 33 cycles of 30 s at 94 °C, 30 s at 55 °C and 72 °C for 1 min; a final extension of 10 min at 72 °C; and cooling to 4°C. The PCR amplification products were separated in a 2% agarose gel stained with GV II (Biomec, China) and photographed with an FR-200A luminescent and fluorescent biological image analysis system (Furi, China). The product size was determined using a 100-bp molecular size ladder.
2.4. Statistical Analysis
The results were expressed as the presence or absence of a virus in an apiary. We considered the virus to be present in an apiary if samples from any colony were positive for any virus. Differences in the viral infection percentages of different provinces and virus co-occurrence were estimated using the Chi-square test with the SPSS 21 software.
3. RESULTS
3.1. Virus Prevalence and Occurrence
As shown in Table 1, we screened for the presence of 8 viruses in 56 apiaries from 13 provinces. Different types of viruses were detected in all provinces with the exception of Jiangsu province, where no virus was detected. All of the viruses were present in the different provinces with the exception of KBV and ABPV, which were not detected. BQCV was the most prevalent virus and was present in 18% of the A. mellifera apiaries and in seven provinces. CSBV was found in 38.9% of the A. cerana apiaries and in seven provinces. In contrast, BQCV was detected in 22% of the A. cerana apiaries, and CSBV was found in 2.6% of the A. mellifera apiaries. The following frequencies were obtained for the remainder of the viruses: IAPV was detected in 7.9% of the A. mellifera apiaries and 5.6% of the A. cerana apiaries; CBPV was found in 15.8% of the A. mellifera apiaries and was not detected in A. cerana; SBV was identified in 5.3% of the A. mellifera apiaries and 16.7% of the A. cerana apiaries; DWV was found in 13.2% of the A. mellifera apiaries and 5.6% of the A. cerana apiaries; DWV-B was found in 10.5% of the A. mellifera apiaries and was not detected in the A. cerana apiaries; AmFV was found in 13.2% of the A. mellifera apiaries and 11.1% of the A. cerana apiaries; and DWV-A/KV was found in 10.5% of the A. mellifera apiaries and 11.1% of the A. cerana apiaries.
BQCV was the most common virus and accounted for 84.6% of the tested samples in each province (Fig. 1). The second highest frequency virus was CSBV (61.6%), followed by another 3 viruses at an equal frequency (53.8%; DWV, AmFV, and DWV-A/KV). The least frequent viruses were IAPV and DWV-B with 30.8%.
| Apiary | Region | IAPVb | BQCV | CBPV | ABPV | SBV | CSBV | DWV | DWV-B | AmFV | KBV | DWV-A/KV | |||||||||||
| A. m | A.c | A. m | A.c | A. m | A.c | A. m | A.c | A. m | A.c | A. m | A.c | A.m | A.c | A. m | A.c | A. m | A.c | A. m | A.c | A. m | A.c | ||
| Zhejiang | JHa | - | - | + | + | - | - | - | - | - | - | - | + | - | - | - | - | + | + | - | - | + | - |
| Henan | PY/ZK | - | - | + | + | + | - | - | - | - | + | - | + | + | - | + | - | - | + | - | - | + | - |
| Hubei | SNJ/ES | × | - | × | - | × | - | × | - | × | - | × | + | × | - | × | - | × | - | × | - | × | - |
| Anhui | HF | - | - | + | - | + | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| Guangdong | GZ | × | - | × | + | × | - | × | - | × | - | × | - | × | - | × | - | × | - | × | - | - | - |
| Liaoning | HLD/CY | + | - | + | - | + | - | - | - | + | - | + | + | + | - | + | - | + | - | - | - | + | - |
| Hunan | YY | × | - | × | - | × | - | × | - | × | + | × | - | × | - | × | - | × | - | × | - | - | - |
| Beijing | HD | - | - | + | + | + | - | - | - | - | - | - | + | + | - | + | - | + | - | - | - | + | - |
| InnMon | AH | + | × | + | × | + | × | - | × | + | × | - | × | + | × | - | × | + | × | - | × | + | × |
| Chongqing | NC | × | - | × | - | × | - | × | - | × | - | × | + | × | + | × | - | × | - | × | - | × | - |
| Jiangsu | XZ | - | × | - | × | - | × | - | × | - | × | - | × | - | × | - | × | - | × | × | - | × | |
| Heilongjiang | YC | + | × | + | × | + | × | - | × | - | × | - | × | + | × | + | × | + | × | × | - | + | × |
| Yunnan | WX | × | + | × | - | × | - | × | - | × | + | × | + | × | - | × | - | × | - | × | - | × | + |
| No. (%) of positive apiaries | 7.9 | 5.6 | 18.4 | 22.2 | 15.8 | 0 | 0 | 0 | 5.3 | 16.7 | 2.6 | 38.9 | 13.2 | 5.6 | 10.5 | 0 | 13.2 | 11.1 | 0 | 0 | 15.8 | 5.6 | |
| 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | 38 | 18 | ||
b +, positive; -, negative; ×, samples not available.
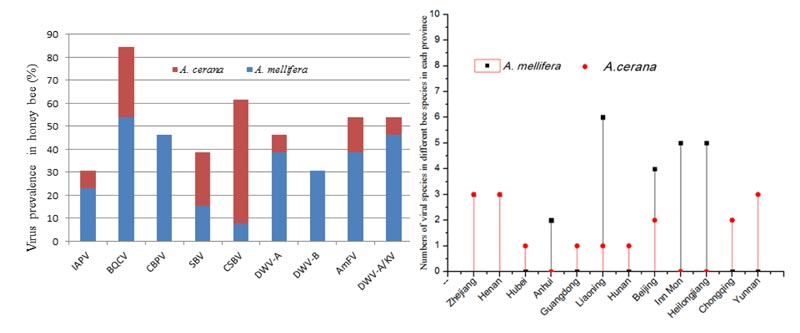
To investigate the prevalence and distribution of honey bee viruses, we screened for the 8 viral species in the samples from these two bee species (A. mellifera and A. cerana). The number of honey bee viruses in A. mellifera was higher than the number in A. cerana in the same province (Fig. 1B) with the exception of Chongqing and Yunnan, which gave priority to keeping A. cerana. The number of viral species in A. mellifera in Zhejiang and Henan was the same as the number of viruses in A. cerana. Liaoning province had the highest number of viruses, with 6 viruses in A. mellifera and only one in A. cerana. Similarly, there were 5 viruses in A. mellifera in Inner Mongolia and Heilongjiang, and no viruses were detected in A. cerana. These results demonstrated that the virus host choices were significantly different in the different bee species (A. mellifera and A. cerana).
3.2. Co-occurrence of Viruses in Each Province
The simultaneous infection of more than one virus in one province was fairly common in both the A. mellifera and A. cerana apiaries (Tables 2 and 3). In A. mellifera, virus co-occurrence varied from 2 to 5 viral species. Co-occurrence with 2 viruses represented 39.46%, 3 viruses represented 35.83%, 4 viruses represented 10.52% and 5 viruses represented 7.89%. Among the multiple co-occurrences, BQCV and CBPV simultaneous infection was the most common 2 virus co-occurrence and accounted for 10.53%. IAPV, SBV, and CBPV were the highest percentage 3 virus co-occurrence with 7.89%, followed by BQCV, CBPV, and AmFV. IAPV, CBPV, AmFV, DWV, and DWV-B were 4 virus co-occurrences equal to IAPV, BQCV, CBPV, DWV, DWV-B, and AmFV with 5.26%. Surprisingly, we detected 5 viruses co-occurrence in A. mellifera with IAPV, BQCV, CBPV, DWV, DWV-B, and AmFV. In contrast, fewer co-occurrences were detected in A. cerana compared to A. mellifera (Table 3). Virus co-occurrence in A. cerana primarily involved 2 and 3 viruses. Simultaneous infections with 2 viruses were detected in 44.45% and with 3 viruses in 11.12% of the apiaries. The most prevalent co-occurrence was BQCV and AmFV in A. cerana, accounting for 11.11% and equal to BQCV and CSBV.
Co-occurrence with 3 viruses (IAPV, BQCV, DWV, and DWV-B) was significantly higher than the other co-occurrence in A. mellifera, as shown in Table 1 (χ2=12.61, P=0.00038, df=1). (Table S3), whereas in A. cerana co-occurrence with 2 viruses was significantly more common than the other co-occurrence types (e.g., BQCV and DWV; χ2<10-5, P=0.004, df=1) (Table S4). Similarly, the numbers of viral species significantly differed between the different provinces. As shown in Table 1, Liaoning province was the highest with 9 viruses, followed by Henan province with 8 viruses. Another 3 provinces (Beijing, Inn Mon, and Heilongjiang) hosted 7 viruses. The lowest number of viruses was detected in Hubei and Yunnan provinces, with one virus each.
| Number of Viruses | Type of Co-occurrence | Number of Apiaries | Percentage (%) |
|---|---|---|---|
| 2 | BQCV; DWV-A/KV | 1 | 2.63 |
| BQCV; CBPV | 4 | 10.53 | |
| CBPV; DWV | 2 | 5.26 | |
| BQCV; DWV | 1 | 2.63 | |
| BQCV; AmFV | 2 | 5.26 | |
| IAPV; DWV | 1 | 2.63 | |
| IAPV;SBV;CSBV | 2 | 5.26 | |
| IAPV;DWV;DWV-B | 2 | 5.26 | |
| 3 | BQCV; AmFV; DWV-A/KV | 1 | 2.63 |
| BQCV; CBPV; DWV | 1 | 2.63 | |
| BQCV; CBPV; AmFV | 2 | 5.26 | |
| BQCV; CSBV; CBPV | 1 | 2.63 | |
| IAPV; SBV; CBPV | 3 | 7.89 | |
| IAPV;SBV;CSBV;DWV | 1 | 2.63 | |
| IAPV; BQCV; DWV; DWV-B | 5 | 13.16 | |
| 4 | IAPV; DWV; DWV-B; CBPV; AmFV | 2 | 5.26 |
| IAPV; SBV; BQCV; DWV; CSBV | 2 | 5.26 | |
| 5 | IAPV; BQCV; CBPV; DWV; DWV-B; AmFV | 3 | 7.89 |
| Number of Viruses | Type of Co-occurrence | Number of Apiaries | Percentage (%) |
|---|---|---|---|
| 2 | BQCV; AmFV | 2 | 11.11 |
| BQCV; CSBV | 2 | 11.11 | |
| IAPV; SBV;CSBV | 1 | 5.56 | |
| BQCV; DWV | 1 | 5.56 | |
| BQCV; CBPV | 2 | 11.11 | |
| 3 | BQCV; CSBV; DWV-A/KV | 1 | 5.56 |
| SBV; CSBV; AmFV | 1 | 5.56 |
4. DISCUSSION
This study aimed to elucidate the prevalence and distribution of honey bee viruses in China and provided deep insight into the distribution and prevalence of 8 viral species in two honey bee species (A. mellifera and A. cerana) in China. The results showed that several viruses with a high prevalence and wide distribution in China were not previously reported, such as DWV-B and DWV-A/KV. Our surveys indicated that DWV-A/KV and AmFV were widely distributed in China in both A. mellifera and A. cerana. Although a previous study investigated the viruses across several provinces, the viruses were not detected [37, 38]. Additionally, neither ABPV nor KBV was found in any province. This primary investigation helps to narrow future efforts to identify the cause of colony decline.
DWV-A/KV was reported for the first time in China. DWV-A/KV was found in 7 out of 13 provinces and in both A. mellifera and A. cerana. Our results showed that DWV-A/KV had a fairly high infection rate and a significant geographical distribution in A. mellifera. There have been few reports on its occurrence and prevalence since DWV-A/KV was demonstrated in Japan [39] and Italy [40]. In this survey, the frequency of DWV-A/KV was equal to DWV, which was considered the most prevalent virus in A. mellifera. In a previous study, DWV-A/KV was identified as highly similar to the DWV strain [41]. This finding might reflect the interaction between DWV and DWV-A/KV, which increased the infection rate of DWV-A/KV owing to the increase in DWV by V. destructor [42]. These interactions may have caused the wide distribution described above. We also found that DWV-A/KV infected A. cerana with a high prevalence. Although we do not know why and how A. cerana is infected by DWV-A/KV, it may be a key player in the honey bee population decline.
AmFV is one of two DNA viruses that infect honey bees and have an infection prevalence equal to DWV-A/KV. As shown in the results, AmFV had a high prevalence in A. mellifera and A. cerana and was widespread in different areas in China. This finding suggested that AmFV did not represent a sudden emergence that might be ignored but instead previously existed in colonies. In most cases, V. destructor, which is an important parasite of honey bees, is considered to be an effective vector for various honey bee viruses, such as KBV [43] and DWV [44]. Recent evidence on the use of typical resistance behavior to combat Varroa showed that Varroa was rarely found in A. cerana year-round [45] and thus did not pose a deadly threat even when found in A. cerana colonies. Consequently, the presence of AmFV in A. cerana was not likely to be due to transmission from V. destructor. AmFV was initially reported to possibly correlate with crawling bees at the entrance of beehives [16]. Subsequently, there were few reports on AmFV infection for a long time worldwide. Recently, AmFV was found in crawling bees and was closely associated with Nosema apis, which also had a significant correlation with BQCV during the fall but no correlation with Nosema ceranae [24]. We found that more colonies declined in provinces where more bee colonies were infected AmFV, such as Zhejiang and Liaoning provinces. However, the mechanism underlying the crawling symptom is not clearly understood. The current study combined with previous data showed that this virus might be a new putative key player in honey bee decline, although we did not provide direct evidence correlated with the crawling symptom. We would like to investigate the presence of N. apis and identify the relationship between AmFV and N. apis. Luckily, the full AmFV genome was announced, implying that we need to pay more attention to DNA viruses that can infect honey bees.
Another interesting result was that KBV and ABPV were not detected in any of the samples. KBV has often been described in Europe and the USA [30, 31] and other regions [46-48] but has never been found in China, Austria, or Hungary [26]. Recently, a survey on CCD colonies found that the KBV infection rate and titer level were higher in CCD colonies than in control colonies [49]. A subsequent investigation demonstrated that KBV occurred in 1.23% of A. cerana samples in South Korea [50]. Our results were in accordance with a previous report from China [37] that did not detect KBV. These results suggested that KBV infection did not occur in China. Similarly, ABPV was not present in our samples. This result differed from a previous study [36] in which ABPV only occurred in adult bees of A. mellifera from Shanxi and Qinghai but was not detected in A. cerana; similar results were reported by Li et al. [37]. This difference might be due to the use of different sample sources or that the level of infection was below the detection limit. Likewise, ABPV was not detected in A. cerana in South Korea [50] and in savannah honeybees [33]. This lack may be related to V. destructor infection because V. destructor is a significant vector for ABPV [51]. Accordingly, these studies demonstrated that ABPV occasionally occurred in A. mellifera but not in A. cerana in China.
BQCV was the most prevalent virus in A. mellifera (18%) and the second most prevalent virus in A. cerana (22%). This finding was in accordance with the results of Li et al. [37], who reported that BQCV had the highest infection percentage in all detected viruses and reached 92%. In this study, the BQCV infection rate was lower than a previous study in A. cerana [38]. In contrast to other viruses, BQCV was very common in A. mellifera and A. cerana, suggesting that BQCV was not strictly required for host selection. As described in different bee species from other regions [27, 50, 52], its incidence was always greater than 70%, especially in Austria, where the prevalence was up to 94% [28]. Similarly, BQCV was the most prevalent virus in savannah honey bees across all seasons and could be found throughout the year [34]. Moreover, BQCV was present independent of the keeping type and was identified in 60% and 92% of local and migratory bees, respectively [53]. However, we could not confirm the cause of the high prevalence owing to the lack of data for other pathogens. For instance, BQCV was reported to be closely associated with Nosema apis [17], and occasionally, the presence of BQCV was completely dependent on the parasite [54]. BQCV has been known to be a honey bee pathogenic virus for a long time, including A. cerana cerana. These results indicated that more studies need to be performed to identify why BQCV is prevalent in most honey bee species.
CBPV was the second most prevalent virus in A. mellifera colonies. A total of 15.8% of the A. mellifera samples from 6 provinces that we screened were positive for CBPV. However, the virus was not detected in A. cerana, which was in agreement with the results of Li et al. [37]. This result was lower than the 69% reported for Belgian apiaries [52] but higher than other regions, such as the 3% in the Czech Republic [55] and Austria [28]. Ai et al. [36] found CBPV at a low level in Guangdong in A. cerana, which was similar to reports of A. cerana from South Korea [50]. This result showed that sporadic CBPV infection might occur as a covert infection at a lower level in A. cerana colonies. Thus, CBPV may prefer A. mellifera over A. cerana.
DWV and DWV-B were found in 13.2% and 10.5% of the A. mellifera samples, respectively, and occasionally occurred in A. cerana. This result significantly differed from previous studies in which DWV was thought to be the most prevalent virus in Europe; for instance, DWV was found in France at rates up to 56% in spring, 66% in summer, and 85% in autumn [29]. Similarly, DWV was found in up to 98% of bee samples and 72% of bees in migratory hives in the USA [53] owing to the spread of V. destructor. DWV and DWV-B were frequently found together owing to their close association with V. destructor, which is an important vector of honey bee viruses. This survey concerning DWV was different than the report by Li et al. [37] and agreed with the Ai [36] and Yang [38] reports. One reason for the discrepancy might be sampling differences because Yang and Ai sampled from 7 provinces and did not collect sufficient data. Another explanation may be the use of different primers because, in some cases, a combination of DWV and DWV-B infection occurs. As described by Gideon [56], the DWV complex was closely associated with overwintering colony losses and was more harmful than the individual infection. However, DWV was not found in wild A. mellifera scutellata in South Africa [25] and honey bees in Uganda [21]. Therefore, the difference in the DWV infection rate between A. mellifera and A. cerana suggested that although DWV is thought to be responsible for colony decline [31], it exhibits differences in pathogenic virulence in different regions and bee species.
The overall incidence of IAPV reported here was lower than in other regions [34] and a previous investigation in China [37], reaching 7.9% in A. mellifera and 5.6% in A. cerana. Our results demonstrated that IAPV had a significantly different geographic distribution in A. mellifera and A. cerana. All of the IAPV detected in A. mellifera was from northern China, whereas all of the positive A. cerana samples were from southern China. These results suggested that the IAPV prevalence was closely associated with the climate or temperature of the different host ranges. In addition, we did not find that IAPV was closely associated with bee colony losses. Indeed, our results had very few incidences of IAPV, which was consistent with other studies [31].
CSBV and SBV were found to be the most prevalent viruses in A. cerana. The infection percentage of CSBV was 38.9%, which was far higher than the rest of the viruses in all detected provinces. The infection percentage of CSBV and SBV accounted for 55.6% and infected the majority of A. cerana colonies. This result was in accordance with a previous study that showed that these viruses were the most deadly threat to A. cerana [14]. Similar frequencies were previously described in asymptomatic honey bees in China [37] and Italy [31]. Compared to other regions around the world [29, 30], CSBV and SBV were more prevalent in A. cerana than A. mellifera in China. These results showed that the strain of CSBV or SBV was significantly different than the strain used in the phylogenetic analysis [39], which had a remarkably different host. Typically, SBV and CSBV only infect A. mellifera and A. cerana, respectively, although occasionally CSBV infects A. mellifera as recently described [57].
We found multiple virus co-occurrences in honey bee colonies of both A. mellifera and A. cerana. The widespread distribution of honey bee viruses in the two honey bee species suggests a stable interaction between the viruses and their hosts. Simultaneous infection with more than 3 viruses occurred in more than 12 apiaries of A. mellifera, which was more than occurred in A. cerana; in contrast, most cases of virus co-infection in A. cerana involved 2 viruses. As shown in Table 2, more than 3 viruses accounted for 55.24% of the infections in A. mellifera, suggesting that multiple infections were very common in A. mellifera. This finding was in agreement with a previous report that co-infection strengthened the harm to a healthy colony [58]. For A. cerana, mixed infections primarily involved 2 viruses, and no co-occurrence with 4 or 5 viruses was observed [27]. As described in Belgium [59], we also observed a significant correlation between the number of honey bee viruses and colony decline. This result might be correlated with multiple virus co-occurrences. For example, recombination between DWV and DWV-B will induce obvious clinical symptoms and more pathogenicity [60]. Data from experimental studies indicated that multiple virus co-occurrences would increase virulence [61]. A recent report showed that IAPV could be elevated to a higher level when several viruses were mixed to infect honey bees, although the initial IAPV infection level was not the highest [62]. Thus, colony losses might be due to a combination of multiple viruses.
CONCLUSION
To summarize, we investigated the occurrence and prevalence of the eleven most important honeybee viruses and identified remarkable differences in the distribution patterns of the viruses in 13 provinces of China. We demonstrated for the first time the presence of DWV-A/KV in China. The most prevalent viruses were BQCV, CBPV, DWV-A/KV, and AmFV in A. mellifera and CSBV and SBV in A. cerana. Although viral infections of the honey bee have been recognized for a long time, their significance is still not fully appreciated. The results of this study suggest that future studies should focus on these viruses to narrow our objectives and help us understand the differences in different areas and hosts. According to the results published in a previous study [33], the differences in the distributions and levels of different viruses among the different tissues of honey bees also require further study, although multiple virus co-occurrences were found.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Our field collections did not involve endangered or protected species, and no specific permissions were required. The protocol was approved by the Animal Research Ethics Committee of the Institute of Apicultural Research, Chinese Academy of Agricultural Sciences (Approval No. 14925-2001).
HUMAN AND ANIMAL RIGHTS
The honey bees were handled in accordance with the Good Animal Practices required by the Animal Ethics Procedures Guidelines of the People’s Republic of China.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The authors confirm that the data supporting the findings of this research are available within the article.
FUNDING
We acknowledge the support of the Fundamental Research Funds for CAAS (Y2020PT17), the Agricultural Science, Technology Innovation Program of CAAS (CAAS-ASTIP-2020-IAR), The Key Technology Research and Development Program of Guangxi province (AB16380094), The Project of Guizhou Science and Technology in 2019 (2292).
CONFLICTS OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publisher’s website along with the published article.


