All published articles of this journal are available on ScienceDirect.
Biofilm Targeting Strategy in the Eradication of Burkholderia Infections: A Mini-Review
Abstract
Burkholderia are intracellular pathogenic bacteria which can produce biofilm. This biofilm protects the intracellular pathogenic bacteria from antibiotic treatment and the immunological system of the host. Therefore, this review aims to describe the capacity of Burkholderia to form a biofilm, the regulation of its biofilm formation, the efficacy of antibiotics to eradicate biofilm, and the novel therapy which targets its biofilm. Burkholderia's biofilm is characterized by its lipopolysaccharides, exopolysaccharides (EPSs), biofilm-associated proteins, and eDNA. Its regulation is made by quorum sensing, c-di-AMP, sRNA, and two component systems. Many antibiotics have been used as sole or mixture agents; however, they are not always effective in eradicating the biofilm-forming Burkholderia. Inhibitors of quorum sensing and other non-conventional antibiotic approaches are promising to discover effective treatment of Burkholderia infections.
1. INTRODUCTION
Burkholderia is a gram-negative bacilli group and living as saprophyte primarily in soil and water [1]. However, several Burkholderia species can cause infectious diseases in human beings, such as Burkholderia pseudomallei, which cause Melioidosis and B. cepacia complex, which cause pneumonia, bacteremia, UTIs, and septic arthritis in patients with cystic fibrosis [2, 3]. Melioidosis is endemic in Southeast Asia and Northern Australia. From 2014 - 2017, a study reported that there were increasing Melioidosis cases (145 cases) in Indonesia, with a mortality percentage that reached 43% more than the previous year [4]. The virulence factors such as hydroxytetradecanoic acid, biofilm formation, flagella expression, and ultrastructure adapt to the host tissue and probably impact it by causing Melioidosis [2].
Biofilm formation by Burkholderia has an important role in its pathogenesis process. As a virulence factor, Biofilm is defined as sessile microbial communities that attached irreversibly to organic or inorganic surfaces of each other. Burkholderia produces matrixes of extracellular polymeric substances (EPS), which is embedded in them, and exhibit phenotype changes in line with the growth rate and gene transcription [5]. Biofilm increases the ability of B. pseudomallei to survive in the host cell and escape from the immune system [6]. Compared to other bacterial infections, Burkholderia infections cause a very high relapse rate associated with biofilm formation [7]. Moreover, previous studies reported that Burkholderia is more resistant to antibiotics when growing as a biofilm than their free-living [planktonic] counterparts [6, 8].
Resistance to many antimicrobial agents, including first and second generations of cephalosporins penicillins, macrolides, colistin, rifamycins, and aminoglycosides have been found in Burkholderia [9]. Several novel antibiotic combination therapies for Burkholderia infections have been reported [10]. However, biofilm-targeted therapies among Burkholderia have not been reported. Therefore, this review aims to describe the capacity of Burkholderia to form biofilm, the regulation of Burkholderia biofilm formation, antibiotic efficacy in the eradication of biofilm, and the biofilm targeting therapy in Burkholderia infections. The Burkholderia discussed in this review are the human pathogens, namely B. pseudomalei, B. mallei, and Bcc (Burkholderiacepaia complex), including B. cepacia, B. multivorans, B. cenocepacia, and B. stabilis.
2. BIOFILM FORMATION IN Burkholderia
Clinical signs of Melioidosis vary from acute septicemia to chronic inflammation or subclinical infections. These signs are determined by the capacity of B. pseudomallei to produce biofilm. B. pseudomallei biofilm plays a role during bacterial-host interaction. Biofilm determines the adhesion to epithelial cells of the human lungs, interactive intercellular processes, apoptosis/necrosis, proinflammatory responses, and cellular pathogenesis [11]. Biofilm formation begins with biofilm surface attachment by flagella or pili, then makes cell-to-cell interactions or micro-colony formations, and finishes with exopolysaccharide secretion or a biofilm matrix [12]. A biofilm matrix glues cells together on a surface and eventually releases them to make new colonization of other surfaces [13]. The capacity of Burkholderiato form biofilm is affected by several factors, such as the cell wall structure (lipopolysaccharides), biofilm proteins, and extracellular DNA (eDNA).
2.1. Lipopolysaccharides (LPS)
Lipopolysaccharides (LPS) are common structural components in a Gram-negative bacteria cell wall, including Burkholderia. Generally, LPS has a role as endotoxin and triggers the defense-related immune responses of the host [14, 15]. Moreover, LPS also able to determine the resistance to antimicrobial peptides and also have the other function that was listed in Table 1. Structurally, the LPS component of Burkholderia species is varied, but mostly it has three regions such as Lipid A, core oligosaccharide, and O-polysaccharides (OPS) [16].
| Type of LPS | Function |
|---|---|
| BP-LPS | Endotoxin, tumour necrosis factor, interleukin-6, and nitric oxide |
| LPS B.cenocepacia |
cytokine induction |
| LPS B.cenosepacia |
Determine resistance to antimicrobial peptides Prevent binding of the peptide to the bacterial cell envelope |
| LPS Bcc |
Induce a strong immune response that can contribute to host cell damage. Lower the anionic charge of the Bcc cell surface, which inhibits the binding and subsequent effects of cationic antibiotics |
| LPS B.mallei |
Play a significant role in the pathogenesis of human disease. Produce high levels of TNF-alpha, IL-6 and RANTES |
| CPS, B. mallei and B.pseudomallei | Antigen |
Lipid A is composed of hexa-acylated diglucosamine that binds LPS to the outer membrane [15, 16]. Lipid A is potentially immunogenic and can interact with receptor proteins of the innate immune system such as Toll-like receptor 4; such interactions are the first line of defense against bacterial infections and can trigger caspase-11 mediated cell death [17]. O-polysaccharides (OPS) are attached to the core of oligosaccharides and are comprised of repeating units of three to five sugars, each of which may bear a range of post-glycosylation modifications [16, 17].
LPS in different species of Burkholderia is varied. LPS of B. multivorans has two O-polysaccharide chains. The B. cenocepacia LPS inner core oligosaccharide determines the in vitro resistance to antimicrobial peptides and B. pseudomallei. LPS has three types that are serologically different [type A, type rough, and type B smooth] [18]. A study conducted by Narisara et al. [18] showed that the type A LPS of B. pseudomallei produced the lowest amount of biofilm. Therefore, it means that the different types of LPS impact the capacity of Burkholderiato from a biofilm.
2.2. Exopolysaccharides (EPSs)
Burkholderia can produce extracellular polysaccharides or exopolysaccharides (EPSs), the high-molecular-weight sugar-based polymers that are synthesized and secreted by multi microorganisms.EPSs have the function as a scaffold of biofilms to cross-link the bacterial cells together and make bacterial adaptation to different stress conditions [19, 20]. Pellizzoni et al.reported that B. cenocepacia in a non-mucoid state could form biofilm containing EPSs [21].
2.3. Biofilm-Associated Proteins
Biofilm-associated proteins in Burkholderia are unclearly identified. However, previous studies reported that some proteins and encoded genes had a role in biofilm formation.
Pinweha reported the bpsl1039-1040 ATP-binding cassette transporter's inactivation reduced the biofilm formation of B. pseudomallei [22], and BcpA protein in Burkholderia plays an important role in biofilm development. [23] However, the BcpA activity is an independent contact-dependent growth inhibition (CDI) system. CDI is the toxic C-terminus of a surface exoprotein used to inhibit the growth of susceptible bacteria on cell contact. CDI proteins are important in cooperative behaviors to build biofilm communities and prevent non-self-bacteria from entering the community [23]. Encoded genes for surface proteins involved in the biogenesis and maintenance of an integral outer membrane. Encoded genes for regulatory factors are required for biofilm maturation [24].
2.4. Extracellular DNA
Extracellular DNA (eDNA) is an essential component of biofilm. eDNA is released from the autolysis processes. The lysis of a bacterial subpopulation generates eDNA under the control of the quorum-sensing system or proceeded in a fratricide mode/suicide similar to eukaryotic cells' necrosis. The eDNA is an important substance in biofilm formation, especially in DNA damage repair, gene transfer, and nutrient source. Other functions of eDNA are to stabilize biofilm, bind and shield biofilm from antibiotics and aminoglycosides, as well as to induce antimicrobial peptide resistance. The acidification of biofilm by eDNA increases aminoglycoside resistance [25]. Pakkulnan et al. (2019) reported that eDNA is important during bacterial cell attachment and biofilm formation [26].
3. REGULATION OF BIOFILM FORMATION
Biofilm formation is a complex process that needs multifactorial things to develop. The regulation of biofilm formation is controlled by a quorum-sensing system (QS), second messenger cyclic diguanosine-5′-monophosphate (c-di-GMP) as a global intracellular protein expression, small RNAs (sRNAs), and Two Component Systems (TCSs) [27, 28]. QS is responsible for regulating the eDNA release, biosurfactants production, and the expression of a large surface protein. c-di-GMP is important in the regulation of the production of EPS and surface proteins. sRNAs are able to regulate the production of EPS [27]. The last TCSs have functions in multiple mechanisms such as cross-regulate, integrate, and coordinate various input stimuli to control biofilm formation [28].
3.1. Quorum Sensing System of Burkholderia
Two main behavioral traits of Burkholderia are intracellular life and biofilm formation [29]. The expression of these two virulence factors determines antibiotic treatment failures and is associated with quorum sensing. Two recognized signaling systems of quorum sensing in Burkholderia are the diffusible signal factor cis-2-dodecenoic acid (BDSF) and N-acyl homoserine lactones (AHLs). Both control similar phenotypic traits [30].
Acyl-HSL [Acyl homoserine lactone] mediates the gene regulation that influences biofilm formation. Most of the synthesized acyl-HSL is octanoyl-HSL [31]. The synthesis of Acyl-HSL is associated with biofilm formation. Therefore, quorum sensing plays an important role in the pathogenesis of Burkholderia, even though it does not regulate biofilm formation under all growth conditions [31].
The diffusible signal factor (DSF) molecule involved in biofilm formation and pathogenesis is a medium-length chain of monounsaturated fatty acids with an unusual cis-2 double bond. Burkholderiahas DSF (named BDSF), which contains cis-2-dodecenoic acid that can inhibit the formation and cause the dispersion of biofilm. Moreover, BDSF can regulate the expression of many genes, such as chitinase, which is responsible for biofilm formation. The presence of chitinase affects the antibiofilm activity of BDSF [32].
3.2. Cyclic Diguanosine-5′-Monophosphate (c-di-GMP)
Cyclic diguanosine-5′-monophosphate (c-di-GMP) is a second messenger that plays a main role in regulating biofilm formation in many bacteria, including Burkholderia. [33] In general, high intracellular c-di-GMP levels induce the production of extracellular biofilm matrix components. In contrast, low intracellular c-di-GMP levels suppress the production of matrix components and promote single cell motility [34, 35]. To regulate these cellular functions, c-di-GMP binds to specific effectors, which could be proteins or RNA, and alters their structure [36]. The presence of higher intracellular c-di-GMP concentration correlates with the increasing cell-to-cell aggregation and EPS production, swimming motilityor absence of flagella, and abundance of biofilm [37].
3.3. Small Non-Coding RNAMolecules (sRNAs): Genetic Control for Bacterial Biofilms
Small non-coding RNA molecules (sRNAs) are an RNA fragment that is 50–500 nucleotides in size and can regulate gene expression by interacting with other RNAs or proteins [38, 39]. As a regulator, sRNAs play the main role in the regulatory network of the post-transcriptional level of biofilm formation.[40, 41] Sasset et al. reported that the biofilm matrix of B. cenocepacia had more sRNAs than planktonic cultures [42].
3.4. Two-Component Systems (TCSs)
Two-component systems (TCSs) are essential in signaling events. Adaptation to environments, cell-cell communication, and pathogenesis are associated with TCSs. TCSs are absent in humans and other mammals. Adaptive changes in cellular processes are regulated by TCSs to respond to changes in environmental conditions [42, 43]. A study reported a novel TCS in B. cenocepacia, namely RqpSR, which plays an important role in modulating QS and pathogenesis in B. cenocepacia. Mutations in rqpS and rqpR exert overlapping effects on B. cenocepacia transcriptomes and phenotypes, including motility, biofilm formation, and virulence [44].
Besides the rqpSR gene, another response regulator gene of TCS is bfmR (biofilm formation-associated regulator], which regulates B. pseudomallei biofilm formation. A mutant of the bfmR gene shows suppression of assembly of fimbriae on the cell surface. It reduces biofilm formation that is led by a decrease in the expression of fimbriae chaperone-usher assembly genes. The low-iron growth condition upregulates the bfmR gene expression. Bacteria in low-medium iron with mutant bfmR show retarded growth. Therefore, bfmR is considered an important positive regulator in controlling the assembly of fimbriae and biofilm formation and is upregulated under low-iron conditions [45]. Mangalea and Borlee also investigated the role of a two-component nitrate sensing system, NarX-NarL, in the biofilm formation of B. pseudomallei. They found that the deletion of narX and narL could decrease the biofilm inhibition activity by nitrate. It means that the NarX-NarL two-component system is a global regulator of biofilm formation [46].
4. ANTIBIOTIC EFFICACY FOR Burkholderia INFECTIONS
Resistance to antibiotics is one of the characteristics of biofilm bacteria, including Burkholderia biofilm [47]. Caraher et al. [2007] reported that antimicrobial agents were active against all the Burkholderia strains when cultured planktonically; however, antimicrobial agents' activity diminished when the Burkholderia strains were grown as biofilms [48]. Following the result, the next study reported multidrug-resistance in Burkholderia following the use of beta-lactams, including meropenem, piperacillin/tazobactam, ceftazidime, and imipenem. B. cenocepacia is also reported to be resistant to antimicrobial peptides, such as polymyxin B (PmB) [49]. Burkholderia pseudomallei Bp1651 has also been found to be resistant to several classes of antibiotics that were usually effective for the treatment of Melioidosis, including tetracyclines, sulfonamides, and β-lactams such as penicillins (amoxicillin-clavulanic acid), cephalosporins (ceftazidime), and carbapenems (imipenem and meropenem) [50]. Therefore, in many cases, antibiotics were only effective when used in combination.
Combination therapy can be used to increase the efficacy of treatment. The summary of several examples of antibiotics used for Burkholderia infections is listed in Table 2. Burkholderia has been resistant to many antibiotics when used as a single therapy, but they are effective in combination, such as Burkholderia is resistant to tobramycin but susceptible when tobramycin and amiloride are used as a combination therapy (Table 2).
5. RECOMMENDATIONS FOR THE FUTURE IN THERAPEUTIC TARGETING BurkholderiaBIOFILMS
Alternative approaches are needed to tackle Burkholderia resistance issues, especially in therapies to improve cystic fibrosis' patients' life expectancy and eradicate Burkholderia infections. The main problem of the resistance is Burkholderia can form biofilm. Therefore, alternative treatment strategies need to be explored to ensure a robust pipeline of effective therapies, especially targeting Burkholderia biofilm.
Many studies have been conducted to find a new strategy therapy that can increase the eradication of Burkholderia infections. As an example, Sidrim et al. found that Promethazine, anefflux pump inhibitor, could improve the antibiotic efficacy and disrupt biofilms of Burkholderia pseudomallei [51]. The disruption of biofilm formation can be done from its eDNA, quorum sensing, etc. (Table 3).
| Antibiotic | Burkholderia | Description |
|---|---|---|
| Amikacin | B.cepacia | Resistance |
| Azithromycin-trimethoprim-sulfamethoxazole | Bcc | Inhibit |
| Azithromycin-ceftazidime | Bcc | Inhibit |
| Azithromycin-doxycycline | Bcc | Inhibit |
| Azithromycin-trimethoprim-sulfamethoxazole | Bcc | Inhibit |
| Biapenem | B.cepacia | Resistance |
| Carbenicillin | B.cepacia | Resistance |
| Cefotaxime | B.cepacia | Resistance |
| Cefuroxime | B.cepacia | Resistance |
| Chloramphenicol | B.cepacia | Resistance |
| Ciprofloxacin, imipenem | B.cepacia | Resistance |
| Gentamicin | B.cepacia | Resistance |
| Polymyxin (PmB]. | B. cenocepacia | Resistance |
| Sulphamethoxazole | B.cepacia | Resistance |
| Piperacillin/ tazobactam | B. cepacia | Resistance |
| Tobramycin | Bcc | Resistance |
| Tobramycin & amiloride | B.cepacia | Susceptible |
| Tobramycin & dichloro-isoproterenol & propranolol | B.cepacia | synergistic with dichloroisoproterenol and propranolol. |
| Trimethoprim | B.cepacia | Resistance |
| Clarithromycin-tobramycin | P. aeruginosa | Active/effective |
| Tobramycin & triclosan | Bcc | reduction of viable cells within biofilms but no antimicrobial activity |
| Target | Description |
|---|---|
| BiofilmReg operon | Need to develop a common drug which is effective for treating all these causative agents [63] |
| DNABII | Antiserum [64] |
| eDNA | Tobramycin & DNase (rhDNase) &dispersin [65] |
| Efflux pump | phenylalanine arginine beta-napthylamide (PAbetaN), a universal efflux inhibitor & CTZ and DOX [8] |
| Polysaccharides | Cellulase development resistance [8] |
| PNAG | Immunotherapy. Antisera [67] |
| LpxC LpxC-4 |
LpxC inhibitor, and LpxC-4 in combination with CAZ [68] sulphonamide derivates [69] |
| QS | Quorum sensing inhibitors (QSIs) [70] |
2) LpxC is UDP-3-O-(R-3-hydroxymyristoyl)-N-acetylglucosamine deacetylase, a metalloenzyme that catalyzes the second step in the biosynthesis of lipid A.
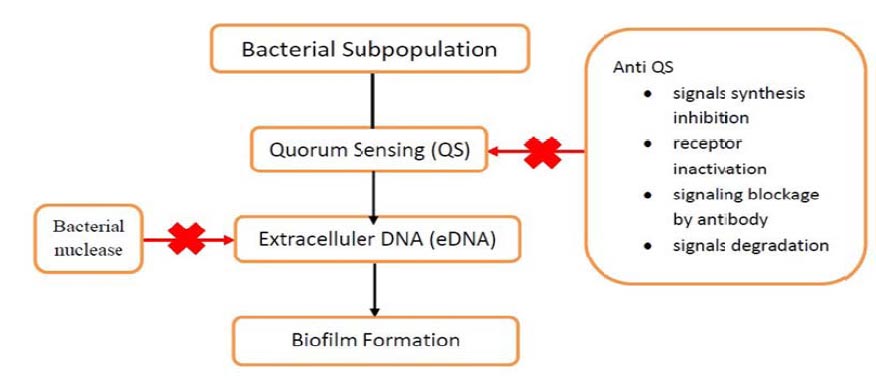
eDNA is an attractive target because it is a matrix component of biofilm. eDNA enzymatic degradation can sensitize biofilm to antimicrobials. In this case, bacterial nucleases can be applied to degrade eDNA (Fig. 1). This means that eDNA could be targeted therapy by disrupting its interactions with other matrix components. It was discovered that eDNA-binding matrix components have come to light. Therefore, targeting the biofilm matrix via eDNA is emerging and promising [52].
In fact, pathogens that colonized the host cell will produce virulence factors and then do the QS signaling to form a biofilm. This bacteria conversation is broken by using anti-QS agents, making pathogens more susceptible to host immune responses and antibiotics (Fig. 1). The QS disruption strategies can be managed by several methods, including signal synthesis inhibition, receptor inactivation, signaling blockage by antibodies, and signal degradation. This way can be used as a potential therapeutic target for bacterial diseases [53].
Carbonic anhydrase (BpsCAgamma) as crucial enzymes in B. pseudomallei can be interfered with by Carbonic anhydrase inhibitors. The most effective inhibitors of BpsCAgammaare acetazolamide, benzolamide, and metanilamide. Other sulfonamides/sulfamates such as ethoxzolamide, topiramate, sulpiride, indisulam, sulthiame, and saccharin are also active even in a higher range [54].
The migration of bacterial cells is allowed by swarming. Swarming motility might thus represent a form of social behavior and is associated with widespread antibiotic resistance [55]. One strategy to block social behaviour of bacteria is to interfere the flagella as an apparatus to motile, cell-cell interactions and decreases the presence of surfactant. Therefore swarming motility might be useful as a model and target of eradicating biofilm antibiotic resistance [56-66].
CONCLUSION
Biofilm is a complex adherence structure to the surface that has a single type of cell or different bacterial colonies. Extracellular polymeric substances (matrix) submerge these colonies. The matrix is composed of proteins, eDNA, and polysaccharides, showing high resistance to antibiotics. Biofilm formation is regulated by quorum sensing. Several approaches are used to inhibit the biological activities of Burkholderia. Inhibitors of quorum sensing need to be discovered and screened. Conventional antibiotic therapy is not effective enough to remedy Burkholderia infections. Therefore, combining other non-conventional antibiotic approaches is promising to discover effective treatments of Burkholderia infections.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


