All published articles of this journal are available on ScienceDirect.
The Influence of the Use of Face Masks During the COVID-19 Pandemic on the Human Microbiome – A Mini-Review
Abstract
The aim of this study was to draw attention to the possible consequences of improper, unhygienic use of mouth and nose covers in the context of prophylaxis against the spread of COVID-19 from the point of view of a family physician and focus on the risk of respiratory infections and skin lesions in patients, in different age groups. The use of protective masks may reduce the likelihood of infection but will not eliminate the risk of infection. However, it should be remembered that any mask, no matter how effective the filtration is or how well it seals, will have little effect if not used in conjunction with other preventive measures, including isolation of infected people, immunization, proper respiratory culture, regular, frequent replacement of masks, and hand hygiene. Additionally, certain risks associated with this form of prophylaxis should be taken into account, which, unfortunately, may also aggravate or even constitute a source of serious respiratory infections and lead to the development and aggravation of skin problems. Moreover, educating society not only on hand hygiene but also on the topic of the value of nose and mouth covers, as well as the frequency of their replacement and/or disinfection, is becoming a significant issue.
1. INTRODUCTION
The outbreak of the Severe Acute Respiratory Syndrome coronavirus 2 (SARS-CoV-2) in China in late 2019 has led to a pandemic and a serious global problem of both public health and the economy. Initially, the virus was named the novel 2019 coronavirus, and the disease it causes was called 2019-nCoV. The World Health Organization identified the disease as COVID-19 on March 11th, 2020. Since the first recorded local outbreak of the SARS-CoV-2 coronavirus in Wuhan, China, in December 2019, the virus and the disease it causes have spread rapidly. Due to the possibility of travel and migration of people, the SARS-CoV-2 virus has spread not only to China but also to many countries on all continents [1]. Hence, on January 30, 2020, WHO announced the outbreak of COVID-19 as the sixth public health emergency of international concern beyond H1N1 (2009), polio (2014), Ebola in West Africa (2014), Zika (2016), and Ebola in the Democratic Republic of the Congo (2019). By June 2021, the WHO had reported more than 172 mln confirmed cases and almost 3,7 mln of deaths worldwide [2]. Based on numerous reports at present, a rapidly increasing incidence of infections has been demonstrated and the possibility of virus transmission by asymptomatic carriers. Due to the constant search for targeted therapy and the development of vaccinations, the greatest emphasis is placed on COVID-19 preventative measures. Therefore, to stop the spread of the virus, social regulations and global cooperation are expected not only from health professionals but from governments and societies. In the face of the global spread of the virus and the lack of effective treatment and vaccine, the main goal is to prevent the still not fully understood effects of COVID-19. Therefore, measures of public health control and prevention of infection are essential. WHO recommends avoiding close contact with people suffering from acute respiratory infections, hand hygiene (frequent washing of hands, especially after direct contact with patients or their surroundings), and avoiding unprotected contact with livestock or wild animals. In addition, in the case of people with symptoms of acute respiratory infection, the so-called cough etiquette (distance, covering the mouth and nose when coughing and sneezing, and washing hands) and improved standard PPE (personal protective equipment), as well as strict infection prevention and control procedures in hospitals (especially in emergency departments), need to be practiced [1].
Due to the need to use prophylactic measures in the form of nose and mouth covers, such as protective masks, attention should be paid to the length of their use in the context of possible secondary infections with own and opportunistic microflora of the mouth and upper respiratory tract and the development of skin infections or their exacerbation. Microorganisms are present on the surface of many human tissues, for example, on the skin, in the mouth (oral cavity microbiome), and the respiratory tract. Extensive characterization developed in the Human Microbiome Project (HMP1) is an interdisciplinary effort of centers, such as Broad Institute, Baylor College of Medicine, Washington University School of Medicine, and the J. Craig Venter Institute, a Data Analysis and Coordination Center (DACC), and several independent investigators. HMP1 identified the diverse habitats of microorganisms in the oral cavity, including the back of the tongue, hard palate, sub- and supragingival plaque, cheek mucosa, palatine tonsils, keratinized gingival tissues, and saliva [3]. The number and variety of microbes in the oral cavity are influenced by many parameters, including age, diet, genetic factors, and individual hygiene. The human microbiota, the so-called physiological flora, is subject to changes as a result of exposure to a number of factors, including antibiotic therapy, the use of vaccines, and even active or passive smoking [4]. The mouth can be divided into three similarly colonized habitats: gingiva and hard palate, tongue and pharynx, and dental plaque [5]. Based on the analysis of the composition of the oral microbiome, its relative stability over time is suggested [6]. Oral microbiome growth is slow, and critical parameters include nutrients and the sensitivity of the microbes to oxygen availability. The oral cavity and mouth, in particular, are exposed to close contact with the surrounding environment through continuity and thus are susceptible to numerous airborne and foodborne microorganisms through food. This can lead to the development of pathobionts from other niches of the upper respiratory tract [3].
2. HUMAN MICROBIOME
2.1. Oral Microbiome
Due to the diverse nature of the oral cavity (oropharyngeal) microenvironment, it is estimated that this area is inhabited by approximately 700 different species of aerobic and anaerobic biofilm-forming microorganisms. The physiological microbiota of the oral cavity consists mainly of microorganisms able to adhere to the surfaces of the gums and teeth, making them more resistant to removal. Non-adherent microorganisms are eliminated by mechanical removal from the mouth with tongue movement, chewing, or speaking, and they end up in the stomach where they are destroyed [7]. Moreover, many microorganisms, due to adherence, can form a biofilm. The structure of the biofilm is an adjacent, well-organized community of microorganisms surrounded by a polymeric, carbohydrate-rich extracellular matrix (ECM) [8].
Due to the contact of the mouth with bacteria of exogenous origin, like from food, water, or air, and even as a result of social contact (e.g., kissing), it is difficult to clearly determine the universal composition of the oral cavity microbiota [9]. As the human dentition develops, microbial populations also change, initially including mainly aerobic and strictly anaerobic species. These include the following genera: Streptococcus, Actinomyces, Veilionella, Neisseria, and some yeast species. Over time, the dominant anaerobes include Prevotella sp. and Fusarium sp. Due to various adhesive factors, which are produced, among others, by Streptococcus spp., such as S. parasanguis and S. mutans, the enamel, and sometimes the gingival epithelial surface and saliva are colonized. Additionally, species from the genus Streptococcus (oralis / mitis / peroris) are also mentioned in the literature, namely Veillonella, Selenomonas, Gemella, Fusobacterium, Prevotella, Lactobacillus, and Neisseria spp [3, 10, 11].
Usually, the relationship between the physiological microbiota and human tissues is a mutual interaction or symbiosis. The homeostasis of the microbial community is influenced by many factors, such as interactions with the external environment, community relations, and internal interactions between different organs, etc. Many oral microbes may be associated with the development of a wide variety of diseases, including periodontal disease, caries, or endodontic infections in the oral cavity, but also a number of other infections develop away from this environment. For many microorganisms, the oral cavity is a gate, due to which they enter various organs of the body and additionally constitute a reservoir of microorganisms, leading under certain conditions to the development of diseases. Research by Raghavendran et al. [12] showed a relationship between the development of various lung diseases and oral diseases, including periodontal diseases. The dentition of hospitalized ICU patients was colonized by Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus aureus [13]. These patients developed a lower respiratory tract infection in the form of pneumonia as the most common secondary infection. The division into two categories of nosocomial pneumonia is based on the use of ventilation; therefore, there are Ventilation Associated Pneumonia (VAP) and non-ventilation-associated pneumonia. In patients with acute respiratory failure or severe diseases, ventilators are used in clinical practice to sustain life. The use of artificial ventilation remains a necessity in the event of a life-threatening situation, but this form of therapy is also not indifferent to the patient's body. There are still unclear aspects of the maximum time during which the system can be safely used continuously. Research on ventilator system contamination is still limited but is very important due to the potential for the development of Ventilation Associated Pneumonia (VAP), heated or unheated humidifiers and their types, water replenishment, and condensate removal systems. In research carried out based on samples taken from ventilator condensates of circuit systems, after 24 hours of their use for patients, in eighty percent of the samples, patient contamination was found with a median bacterial concentration of 2x105 organisms/ml. Moreover, a direct correlation between the cultured bacteria isolated from circuit condensates was found with microbes cultured from patients' saliva, suggesting that the patients' oral cavity and throat microbiome were the primary sources of the circuit colonization [14]. A high level of bacterial contamination was demonstrated in the study of samples taken from various locations of reused and disposable ventilators. This research draws attention to the importance of the sterilization process, especially when reusing ventilator systems, and the avoidance of unnecessary manipulation during the operation of ventilation systems to promote health and safety also in the healthcare staff [15].
Microorganisms (pathogens, pathobionts, or the microbiome in general) inhabiting biofilms in the oral cavity can enter oral secretions and colonize the surfaces of ventilators in intensive care units, being a source of infections [16]. Another subtype of lower respiratory tract infections is aspiration pneumonia, which develops as a result of inhalation of pathogenic bacteria or pathobionts that colonize oropharyngeal biofilms [17].
It has been suggested that the incidence of, inter alia, infectious lung diseases in patients could be reduced by proper oral hygiene, also in patients without teeth (using dentures) [10]. Currently, efforts are being made to develop an accessible database of all known species of oral bacteria based on the use of molecular biology techniques targeting the 16S rRNA gene and next-generation sequencing methods. Thus, the HOMD (Human Oral Microbiome Database) was created, enabling the identification and characterization of bacteria using online tools. In addition, sequencing techniques have also enabled the characterization of the fungal microbiota and a range of oral viruses and bacteriophages. Literature reports suggest that the oral microbiome plays a key role in shaping the host's health profile. There is a complicated relationship between the oral microbiome and the occurrence of diseases outside this environment, affecting many other systems and organs, including diseases of the heart, liver, or respiratory system. The development of microbiomics, metagenomics, and metatranscriptomics facilitates the demonstration of the relationship of complex communities of oral microbes with various diseases of the human body [18].
2.2. Upper Respiratory Tract Microbiome
Extensive research on the complex relationships of the human body's microbiome and its overall well-being, especially on the intestinal microbiome, provide the basis for diagnosis, e.g., of the microbiome of the upper respiratory tract. Although research in this area is conducted less intensively, it is worth noting that the microbiome of the upper respiratory tract is a strong determinant of the health of the respiratory system. Disturbances of the microbiome, e.g., after antibiotic therapy, may lead to the development of infections with opportunistic, potentially pathogenic microorganisms that a human may be a carrier of (pathobionts). Such microorganisms include, among others, Streptococcus pneumoniae, and its overgrowth and spread in the body can lead to the development of both acute local and diffused severe respiratory infections, including acute otitis media (AOM), pneumonia, meningitis, and sepsis [3].
The upper respiratory tract has many important physiological functions, including filtering, moisturizing, and heating the inhaled air, which enters the system through the anterior nostrils, nasal cavity, nasopharynx, sinuses, eustachian tube, middle ear cavity, the oral cavity, the oropharynx, and the larynx. A wide range of microorganisms, mostly bacteria, such as Firmicutes, Actinobacteria, Bacteroidetes, Proteobacteria, and Fusobacteria, inhabit the mucous membranes of these microenvironments and constitute the microbiome of the upper respiratory tract. The nostrils and the nasal vestibule are closest to the external environment and the surface of the skin. A nasal cavity is also a place where secretions from the frontal, maxillary, ethmoid, and sphenoid sinuses accumulate and connect with the nasopharyngeal cavity located further. Currently, it is believed that the organization of microbial ecosystems is based on their selectivity and random processes, including selection, ecological drift, speciation, and dispersion, i.e., the dispersion of organisms in the space of initial bacterial colonization. The shape of the microhabitats of the upper respiratory tract is also influenced by differences in oxygen content, pH, humidity, immunological factors, nutrients, and the type of epithelial cells.
Additionally, the already existing microbial communities are subject to the constant influence of new “colonizers” from exogenous (including air) and endogenous sources (e.g., middle ear and sinus drainage, lung sputum). The bacterial microbiome of the upper respiratory tract varies depending on the ecological niche, and the oropharynx and the mouth are the most diverse. On the other hand, the anterior nostrils are inhabited by a microbiome of low biodiversity, comparable to areas covered with skin-like epithelium. In the case of the anterior nostrils (nasal vestibule), the microbial ecosystem is usually enriched with representatives of the genus Actinobacteria (i.e., Corynebacterium and Propionibacterium spp.) and Firmicutes (i.e., Streptococcus spp. in children and Staphylococcus spp. in adults).
Moreover, some studies indicate high numbers of Moraxellaceae in children and Proteobacteria in adult ICU patients (i.e., Enterobacterales and Pseudomonadales) and a low number of Gammaproteobacteria in healthy adults. Additionally, a certain difference in the microbiome of children and adults is indicated. Despite the similarities in the colonization of the upper respiratory tract niches in adults and children, in the case of the anterior nostrils, a greater abundance of Streptococcaceae, Moraxellaceae, and Neisseriaceae were noted in healthy children than in healthy adults. The diversification of the microbial profile in the nasopharynx is also observed in the course of human development from birth. Initially, the microbiome is characterized by a predominance of species of the genera Moraxella, Corynebacterium, Dolosigranulum, Streptococcus, or Staphylococcus spp. as well as microbiome possibly coming from maternal skin (Staphylococcus and Corynebacterium spp.) and the vaginal area (Staphylococcus, Streptococcus, and Dolosigranoccus) in the peri-birth period. During the first weeks, months, and even years of life, the nasopharyngeal microbiome changes and includes Moraxella, Corynebacterium / Dolosigranulum spp., and Streptococcus spp. Gradual acquisition of Haemophilus influenzae was also observed, the presence of which is also associated with pathogenicity. In the case of adults, the microbiome is characterized by a very similar composition, although clearly reduced by the genus Moraxella. In addition, the so-called pathobionts, including S. pneumoniae, Moraxella catarrhalis, H. influenzae, S. aureus, and beta-hemolytic streptococci, were identified. However, their interactions with commensals inhabiting ecological niches are still not fully understood.
For the oropharynx, which connects the mouth, nasopharynx, larynx, lower respiratory tract, and digestive tract, the potential for exposure to a diverse range of exogenous and endogenous microorganisms was indicated. It is also an ecological niche for potentially pathogenic bacteria that can lead to local (pharyngitis, angina) or diffused infections (inflammation of the lower respiratory tract, pneumonia). The spread of microorganisms from the oropharynx to the lower respiratory tract via (micro-) aspiration or inhalation, due to the significant overlap of the oropharyngeal microbiome and bacterial habitats in the healthy lung, is probably one of the common events in both periods of health and disease. On the basis of metagenome sequencing techniques, it has been shown that the oropharynx in healthy adults is colonized, among others, by pathobionts which include Streptococcus, Haemophilus, and Neisseria spp. and Gram-negative anaerobic commensal genera Veillonella, Prevotella, Leptotrichia, and Fusobacterium spp. The composition of the microbiome of the oral cavity and pharynx in children is similar to that of adults but enriched mainly with Neisseria, Granulicatella, Prevotella, Porphyromonas, Fusobacteriaceae, and some Prevotella spp. Moreover, the oropharynx is home to potentially pathogenic representatives of the genus Streptococcus, including S. pneumoniae, S. pyogenes (group A of beta-hemolytic Streptococcus sp.), Streptococcus dysgalactiae subsp. equisimilis (beta-hemolytic streptococci from groups C and G) [3]. Most of the mentioned microorganisms can lead to various infections both in the upper respiratory tract and in other systems, e.g., S.pyogenes, and complications after the infection (angina) in the form of rheumatic fever and glomerulonephritis.
2.3. Skin Microbiome
Areas with a high density of sebaceous glands, including facial skin, favor the development of lipophilic microorganisms (e.g., Propionibacterium spp., Malassezia spp.). In the case of the skin surface with increased humidity, e.g., when wearing masks that cover the area of the mouth and nose for many hours, they are a habitat for a larger number of microorganisms. Regions with higher temperature and humidity favor the growth of microorganisms, such as Gram-negative rods and Gram-positive cocci as S. aureus [19]. In studies on the survival of some microorganisms, Majchrzycka et al. has shown that in the currently used masks with biological filters protecting the respiratory tract, S. aureus showed the highest survival rate on the filter cloth of the masks [20, 21]. Many common skin conditions are associated with a specific stage of life (including hormonal changes), a specific topographic location, and/or specific microorganisms. In the case of skin diseases, the etiology takes into account the relationship with the cutaneous microbiota, as well as the participation of commensals, which can opportunistically become invasive and cause infection. One should remember the possibility of the presence of previously unidentified microorganisms [19]. Due to the difficulties in unambiguous diagnosis of the etiological factor of skin diseases, the current problem is the influence of various face covers, shielding especially the mouth and nose and adhering to the skin surface. Materials and fabrics, including cloths of various compositions (e.g., cotton, synthetic, mixed), which are also used in the production of masks, have close contact with microorganisms from the exhaled air, skin, and the environment. These materials create a warm and often moist environment in the area being in contact with the upper respiratory tract (nasal cavity and mouth) and on the skin of the face, which is favorable for the growth of certain bacteria and fungi [22].
2.4. Mycobiome
The oral microbiome consists of complex, interactive communities of microorganisms, mainly bacteria and nutrients, that together make up the biofilm. Biofilm can have a variety of effects on the oral cavity modulating the immune system, protecting microorganisms against environmental factors or the oral cavity against potential invading pathogens, or increasing the virulence of certain microorganisms and reducing the effect of antimicrobials. Most often, diseases in the oral cavity are caused by bacterial etiological factors, residents of the oral microbiota. Although bacteria are the dominant etiological factors, it is also worth paying attention to the fungal microbiota, so-called mycobiota, which can lead to certain oral diseases [23].
There are several types of physical and metabolic interactions between fungi and bacteria, which influence a healthy oral microbiome. Metabolic interactions include carbon, lactate, and oxygen metabolism [24]. Candida spp. has been found to be the most common fungal species that colonize the oral cavity and digestive tract, even in infants. Moreover, the in vitro influence of C. albicans on the bacterial composition of early oral biofilms was demonstrated, especially in terms of the presence of the obligate anaerobes. Currently, the knowledge about the fungal colonization of the oral cavity in children is quite limited; however, as in adults and the older group of children aged 10-19, the highest percentage of Candida yeasts indicated that there is an association of interactions between C. albicans and streptococci and synergistic colonization of the two microorganism types in the digestive tract. Candida can lead to oral candidiasis spread, causing candidiasis of the esophagus or other parts of the gastrointestinal tract. Currently, attention is also paid to other Candida spp. such as C. auris. This microorganism is often multi-drug resistant, invasive, and rather difficult to identify. It can colonize, e.g., the oropharynx and mouth. The implications of the C. auris presence are unclear; however, the importance of appropriate antifungal treatment is emphasized, even in the case of minor diseases of the oral cavity, due to the possibility of developing drug resistance. The oral mycobiome is complex, interactive, and is associated with the formation of biofilm. In many studies on the oral mycobiome, Candida yeast's dominant share is emphasized. However, currently, other fungi have also been suggested to be potential members of the oral mycobiome, which is less common, but can also cause oral diseases, e.g., Cryptococcus neoformans, Aspergillus spp., saprophytic Mucoraceae or Geotrichum, Cladosporium, Aureobasidium, Saccharomycetales, and Fusarium.
More research is required to get a complete picture of the role of fungi in a healthy oral microbiome, which will also focus on metabolic interactions between bacteria and other fungi. In the prevention of fungal infections, the importance of maintaining proper oral (and denture) hygiene (e.g., flossing, brushing the teeth and tongue, regular dental care, treatment of xerostomia), sugarless lozenge intake, control of diabetes, avoiding unnecessary corticosteroids, and using rational antibiotic therapy are emphasized [23, 24]. Good oral health is not only limited to dental health but includes the gums, supporting tissues, palate, and mouth lining, and also the throat, tongue, mouth, salivary glands, masticatory muscles, nerves, and jaws. It is widely believed that oral health is an integral part of the functioning and health of the entire human body [24, 25].
3. PREVENTIVE FACE COVERS AND THE CONSEQUENCES OF THEIR USE
The announced COVID-19 pandemic caused by Sars-CoV-2 is a public emergency worldwide (WHO, 2019). Social distance, hand hygiene, surface disinfection, and mouth and nose coverage are paramount practices for infection control during the COVID-19 pandemic. There are global discussions on the legitimacy of the general use of non-medical masks to limit the transmission of SARS-CoV-2 [26]. Thus, both global and local organizations recommend, especially in the case of health care workers and people who may be infected, the use of prophylaxis in the form of protective masks covering the nose and mouth [27].
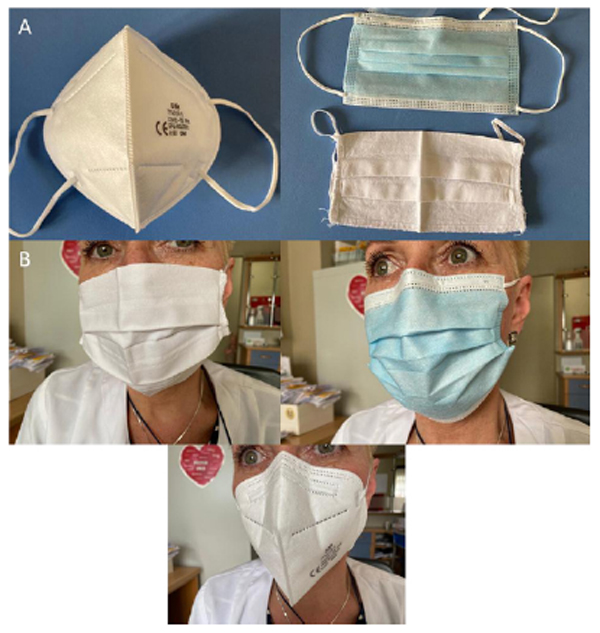
In the context of the current situation, many studies performed to date, conducted not only in recent months but also in recent decades [26], indicate the dependence of filtration efficiency on the materials of the protective masks. Since the 1970s, experiments have been carried out using various techniques (including sedimentation on agar plates) and materials (including medical, cotton, muslin, flannel, and admixture fabrics) for the production of protective masks, in which similar results were obtained. Examples of the masks used most commonly in the COVID-19 pandemic are presented in Fig. (1). In the case of bacteria, the filtration efficiency of medical and fabric masks was assessed up to 99%, and for aerosols (<3.3 μm) up to 89%. The ability of the material to block the transmission of microorganisms is determined by assessing the filtration efficiency expressed as a percentage using surrogate indicators, e.g., biological aerosols, included by the ASTM (American Society of Testing and Materials) in standards for masks. The filtration efficiency depends on the physical retention of particles of varying sizes, regardless of the microorganisms they contain [26]. The efficiency of blocking droplets and aerosols of masks made of cloths may increase with the number of layers [27, 28]. In the analysis of the filtration efficiency of single-layer cotton fabric masks in bioaerosol (0.2 μm), it was estimated at 43% to 94%, and in the case of disposable surgical masks, it was estimated at 98% to 99% [26, 29]. Experience suggests that many (but not all) cloth masks reduce droplet and aerosol penetration. Hence, although there is still a lack of conclusive and unequivocal evidence due to difficulties in developing control groups, nose and mouth protection with protective masks may reduce the risk of contamination of the environment by any virus, including SARS-CoV-2. In the course of the current COVID-19 pandemic and the difficulties in controlling it, the potential advantage of benefits over possible risks is emphasized [26, 30].
The disadvantages of wearing masks, sometimes for a long time, which should be kept in mind, are possible exposure and the risk of developing respiratory tract infections (especially lower respiratory tract can be affected). The source of infection is the secondary acquisition of the patient's own microbiome of the upper respiratory tract, including pathobionts from the patient's exhaled air, depositing, e.g., in the form of droplets on the surface of the mask fabrics. Hence, it is worth paying attention to its use, considering the time, especially long-term (e.g., health care workers). Persistent contact with contaminated and moist cloth/mask material (also with the lowest degree of filtration), apart from respiratory system infections, may lead to extensive skin lesions of the face and exacerbation of existing lesions.
Synthetic fibers have a poor adsorption capacity due to their molecular structure. Cotton is a natural fiber composed almost entirely of cellulose with a high adsorption capacity [31]. Some materials additionally promote the development of certain microorganisms, including Micrococcus spp., Staphylococcus spp., and Propionibacterium spp. Confirmed by in vitro studies, a high affinity of Staphylococcus spp. for cotton and polyester was found [32, 33]. In studies by Callewaert et al., colonization of various materials (depending on composition) by microorganisms belonging to the human microbiome that had contact with them was shown [22].
The influence of the microbiome on the possibility of developing both upper and lower respiratory tract infections is increasingly emphasized. In both cases, a significant role of the so-called pathobionts, the carrier of which is usually asymptomatic, has been indicated in recent studies. Uncomplicated infections of the upper respiratory tract are common worldwide and are characterized by a large incidence. Lower respiratory tract infections are estimated to be less common but are associated with a high mortality rate. Pathobionts that can potentially lead to the development of a number of respiratory diseases, such as S. pneumoniae, M. catarrhalis, H. influenzae, and S. aureus, are often considered to be residents of the upper respiratory microbiome, especially in children under 2 years of age. It has also been shown that the carrier status of these potentially pathogenic microorganisms is not synonymous with the development of respiratory diseases but still poses a risk of their occurrence. There is also a correlation in which the presence of pathobionts enhances the acquisition and replication of viruses in humans. An example is S. pneumoniae, the preincubation of which with human bronchial epithelial cells leads to increased susceptibility to human metapneumovirus infections [3].
Risk factors that may facilitate the development of respiratory infections include new human-acquired strains of microorganisms or the state of bacterial symbiosis. Hence, the stimuli to induce disease may be exo- or endogenous. In addition, it is believed that the composition of the upper respiratory microbiome may also influence the potential for the development of infections. Studies on the tonsil crypt microbiome in children and adults suffering from recurrent tonsillitis showed that Proteobacteria (mainly Haemophilus spp.) were associated with the disease, while high numbers of Bacteroidetes (especially Prevotella spp.) limited disease progression. In analyses of the oropharyngeal microbiome in elderly pneumonia patients, the absence of Bacteroidetes, including Prevotella spp. and other anaerobic bacteria such as Leptotrichia and Veillonella spp., was associated with pneumonia. Based on conventional microbiological tests, a viral etiology of infections is suggested in the majority of acute rhinitis and laryngitis cases. The most common bacterial etiological factors of these infections include S. pneumoniae and H. influenzae, and in the case of laryngitis, additionally S. aureus, beta-hemolytic streptococci, M. catarrhalis, and Klebsiella pneumoniae are considered. Additionally, studies suggest that early colonization of S. pneumoniae, H. influenzae, and M. catarrhalis causes a long-term increased risk of pneumonia and bronchiolitis in healthy newborns, and elderly and immunocompromised people may also cause pneumonia or acute exacerbations of COPD (chronic obstructive pulmonary disease). H. influenzae is an invasive burden on the elderly, with the frequency of occurrence increasing with age.
Due to the need to define preventive strategies for the development of severe diseases in patients >65 years of age, attention should be paid to the potential sources of the invasive microorganisms, such as upper respiratory tract infection, and also on the colonization of the nasopharynx and hygiene when using face masks [34-36]. In addition, studies on the oral microbiome in the context of the development of periodontal diseases, caries, and the cardiovascular system, draw attention to the impact of microbiota imbalance on the potential development of respiratory diseases due to physiological micro-aspiration episodes. The significant importance of the influence of the oral microbiome and microaspiration has been demonstrated in studies in nursing home patients, where neglecting oral hygiene measures led to a significant increase in the incidence and then mortality due to pneumonia. It has been suggested that imbalances in the microbiome of the upper respiratory tract play a key role in the pathogenesis of acute infections such as AOM (acute otitis media) and pharyngitis and may also play a role in the development of lower respiratory tract infections, leading to pneumonia. Composition, biodiversity, host factors, and viral infections all influence the health of the microbiome of the upper respiratory tract. Under certain conditions (e.g., wearing masks for a long time), the balance of commensal microorganisms may be upset, leading to an increased share of pathobionts, the most common of which are: S. pneumoniae, H. influenzae, S. pyogenes, M. catarrhalis, and S. aureus, resulting in respiratory diseases of various course [3].
The studies by Johnson and Morawska [37] showed that when speaking, exhaling, and especially coughing, droplets of 5–100 μm come out of the mouth, and as shown in the studies by Wei and Li [38], masks suppress and reduce the possibility of transmission of infectious agents during close contact and by air in confined spaces. Moreover, in the analyses of Rengasama et al. [39], it was found that masks made of 100% cotton or with 30% polyester admixture showed 40-60% filtration efficiency for polydispersed/dispersed sodium chloride (NaCl) aerosol particles (75 ± 20 nm diameter of the Count Median Diameter (CMD) and a Geometric Standard Deviation (GSD) not exceeding 1.86) at a frontal speed of 5.5 cm / s. In studies by Ho et al. designed for cotton and surgical masks, it was shown that at a speed of 5.5 cm / s, the filtration efficiency was rated at 86.4% and 99.9%, respectively. These results confirmed earlier analyses by van der Sande et al. [40], as well as similar studies by Davies et al. Healthy adult volunteers and children [40], in a specific procedure (identical for all of them) at the appointed time (10-15 minutes or 3 hours for adults) wore filtering masks (FFP2), surgical masks, and cloth masks (kitchen towels), and the test protocol included the measurement of particles concentration on both sides of the mask. All free-floating particles in the air were analyzed using a portable electrostatic classifier and counter that recorded particles ranging in size from 0.02 mm to 1 mm, covering most of the infectious respiratory aerosols [40]. A reduction in aerosol exposure was observed for each type of mask, although FFP-2 masks were the most effective, and towel masks were found the least effective. The studies by Davies et al. [41] were based on the analysis of masks made of cotton T-shirts, surgical masks, and people without masks. In various analyses of protective masks made of single layers of scarves, sweatshirts, T-shirts, and towels, the filtration efficiency was estimated at 10% to 40% with the use of a NaCl aerosol (0.075 μm) [39]. In experiments using a bacterial marker with aerosol-sized particles, the filtration efficiency of a single-layer towel fabric was found to be 83%, and with 2 layers 97%, compared to 96% for surgical masks [41]. The filtration efficiencies of different materials used to produce face masks (based on several studies) are summarized in Table 1.
| Mask Type | Mask Efficiency - Analyzed Particles Size | References | |
|---|---|---|---|
| Droplets | Aerosol | ||
| Single-layer cotton fabric masks (efficiency increase with the number of layers) | 99% | 43% to 94% (~200nm) | Clase et al. (2020) |
| Single-layer: cotton, silk, chiffon, flannel, various synthetics, and their combinations | 5-95% (>300nm) | 5-80% (<300nm) | Konda et al. (2020) |
| Fabric hybrids (such as cotton–silk, cotton–chiffon, cotton–flannel) | >90% | >80% | Konda et al. (2020) |
| Cotton 1-layer | 98.4% ± 0.2 | 79% ± 23 | Konda et al. (2020) |
| Cotton 2-layers | 99.5% ± 0.1 | 82% ± 19 | Konda et al. (2020) |
| Surgical mask | 99.6% ± 0.1 | 76% ± 22 | Konda et al. (2020) |
| N95 (FFP2/FFP3)* | 99.9% ± 0.1 | 85% ± 15 | Konda et al. (2020) |
| Single layer of scarves, sweatshirts, T-shirts, and towels | - | 10% - 40% (NaCl aeroslos - 0.075 μm) |
Davies et al.(2013) |
| 100% cotton and 30% polyester admixture | - | 40-60% (75 ± 20nm NaCl) |
Rengasama et al. (2010) |
| Single-layer towel fabric | - | 83% | Davies et al. (2013) |
| Two-layer towel fabric | - | 97% | Davies et al. (2013) |
| Surgical mask | - | 96% | Davies et al. (2013) |
| Disposable surgical mask | 99% | 89% - 99% (<330nm) | Clase et al. (2020); Furuhasmi (1978) |
| Cotton mask | 86,4% (20-1000nm) | Ho et al. (2020) | |
| Surgical mask | 99,9% (20-1000nm) | Ho et al. (2020) | |
It is confirmed that surgical masks are three times more effective in blocking the transmission of microorganisms than cotton masks; however, each form of airway isolation significantly reduces the number of microorganisms released [27]. It should be noted that the fabric does not retain individual virions, but virus transmission is mostly via larger particles in the secretions, in the form of an aerosol (<5 μm) or droplets (> 5 μm) when talking, eating, coughing, and sneezing, and the evaporation of droplets creates aerosols. Experimental analyses of the filtration efficiency for viruses showed 72% efficiency for single-layer towel fabrics, 51% for t-shirt fabrics, and 90% for surgical masks [41].
Improper fit of the mask can result in over a 60% decrease in the filtration efficiency, implying the need for future cloth mask design studies to take into account issues of “fit” and leakage while allowing the exhaled air to vent efficiently [28].
In general, according to the newest studies, the clinical usefulness of masks, also non-medical ones, is strongly advocated because even fabrics in a single layer restrict the transmission of aerosols, and thus of microorganisms [28]. The main purpose of the use of face masks is, therefore, the ability to retain certain particles that have a limited escape route, especially beyond the mask, so that virus carriers do not penetrate the air in the form of aerosols and have no chance of settling on potentially human-touched surfaces [26]. In this way, the possibility of transmission of infectious agents in the population is reduced with the proper use of masks.
CONCLUSION
Time is needed to characterize COVID-19 and the SARS-Cov2 virus, especially since new mutations of this virus appear in a short time. Despite the vaccines developed, every effort is made to slow the spread of the disease, allowing time for a better and more effective capacity for the health system and society to prepare since no effective treatment has been developed to date [1].
Moreover, it also takes time to vaccinate large parts of the world population and observe the results, and ending the pandemic still seems a long way off. Hence, the use of the currently developed prophylaxis will find its long-term application.
The use of protective masks may reduce the likelihood of infection but will not eliminate the risk of infection. However, it should be remembered that any mask, no matter how effective the filtration is or how well it seals, will have little effect if not used in conjunction with other preventive measures, including isolation of infected people, immunization, proper respiratory culture, regular, frequent replacement of masks, and hand hygiene [41].
Cotton masks can be a potential substitute for surgical masks for people with respiratory tract infection, living in an air-conditioned microenvironment, or in human communities that do not provide social distance. Due to the multiple-use, availability of materials, and the possibility of maintaining the hygiene of this type of mask, it is an alternative to everyday protection for healthy people during a pandemic of respiratory infections [27]. However, certain risks associated with this form of prophylaxis should be taken into account, which, unfortunately, may also aggravate or even constitute a source of serious respiratory infections and lead to the development and aggravation of skin problems. Moreover, educating society not only on hand hygiene but also on the topic of the value of nose and mouth covers, as well as the frequency of their replacement and/or disinfection, is becoming a significant issue.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors confirm that this article's content has no conflict of interest.
ACKNOWLEDGEMENTS
Declared none.


