All published articles of this journal are available on ScienceDirect.
EDL933 Strains of Escherichia coli O157 can Demonstrate Genetic Diversity and Differential Adherence to Bovine Recto-Anal Junction Squamous Epithelial Cells
Abstract
Background:
Differences between Escherichia coli O157 (O157) strains are well-established with some of these strains being associated with major outbreaks in the US. EDL933 is one such O157 strain that caused a multistate outbreak in 1982 and has since been used as a prototype in various O157-related experiments.
Objective:
As O157 can readily acquire genetic mutations, we sought to determine if the genetic and phenotypic profiles of EDL933 strains from different sources would be consistent.
Methods:
We evaluated wild-type O157 strains stocked as EDL933 from three different laboratories, in the strain typing Polymorphic Amplified Typing Sequence (PATS) and the bovine rectal-anal junction squamous epithelial (RSE) cell- and HEp-2 cell- adherence assays. In addition, we also verified if Shiga toxins (Stx), the Locus of Enterocyte Effacement (LEE) or curli fimbriae contributed to the adherence phenotypes observed using mutant and wild-type EDL933 isolates.
Results:
Our results showed differences in PATS profiles and RSE cell-adherence phenotype, with no influence from the Stx or LEE genes, between EDL933 from different sources. Interestingly, the EDL933 strain that demonstrated the most contrasting diffuse adherence phenotype on RSE cells, EDL933-T, had decreased curli production that may have contributed to this phenotype.
Conclusion:
Our observations suggest that a comprehensive characterization of bacterial isolates, even if assigned to the same strain type prior to use in experiments, is warranted to ensure consistency and reproducibility of results.
1. INTRODUCTION
Enterohemorrhagic Escherichia coli (EHEC) O157:H7 (O157) is a well-known Shiga toxin-producing E. coli (STEC) expressing different variants of the bacteriophage-encoded Shiga toxins or Stx (ex. Stx1, Stx2, Stx2c) which act to inhibit protein synthesis in human renal endothelial cells [1-4]. The resulting effects of this pathogenesis can include non-bloody to bloody diarrhea or hemorrhagic colitis (HC), and hemolytic uremic syndrome (HUS), in which Stx2 particularly plays a major role [1, 3, 5]. O157 also expresses a locus of enterocyte effacement (LEE)-encoded adhesin, intimin- γ, that contributes to the tissue tropism for Peyer’s patches and ascending colon enterocytes in humans and the HC pathogenesis through effacement of the intestinal attachment sites [6, 7]. Global estimates of STEC infection in humans report approximately 2.8 million acute illnesses, 4,000 cases of HUS, 270 cases of end-stage renal failure, and 230 deaths, annually [8].
Worldwide outbreaks of E. coli O157:H7 (O157) [2] have been reoccurring ever-since the O157 strain EDL933, banked with the American Type Culture Collection (ATCC 43895), was first isolated from a 1982 food-borne outbreak in the US [9-11]. In fact, O157 along with Campylobacter, Salmonella, and Shigella, comprise the four most commonly isolated pathogens from meat and produce [12, 13]. Cattle are the major reservoirs for O157, which tend to persist at the bovine intestinal recto-anal junction (RAJ), especially at the follicle-associated-epithelial (FAE) cells of the RAJ using the LEE-encoded intimin [6, 14-19]. O157 also binds the squamous epithelial cells (RSE) at the RAJ in an intimin-independent manner suggesting a role for alternate adhesins [20-22]. Given the harmful effects of O157 in humans, its widespread geographic distribution, persistence in the gastrointestinal tracts of cattle and environmental dissemination through cattle feces, O157 poses a significant food safety threat in both developed and developing nations [2, 8, 17, 23].
It is apparent that extensive research is needed to control the persistence of these pathogens; however, a problem becoming more marked is the incessant evolution of the O157 genome [24-27]. In a recent genome announcement, Fellner et al. reported large deletions, including the OI#48 pathogenicity island, and inversions in two of three European O157 strain EDL933 derivatives [28]. Manning et al., using SNP analysis at 96 loci in over 500 O157 strains, were able to demonstrate that an emergent O157 subgroup is acquiring new factors contributing to more severe disease in humans [29]. The study also stated that this subgroup characterized by its ability to cause HUS, increased in prevalence from 10% in the year 2002 to 46% in 2006 [29, 30]. These findings suggest that derivatives of the same strain may have drastic differences in genomic structure and as a consequence, exhibit multiple phenotypes. Thus, when picking strains for experimental work, one must consider this a possibility.
In this study, we set out to explore the idea that variants of the same strain can show genetic and phenotypic differences by testing EDL 933 strains of O157, acquired from three different sources, in assays standardized in our laboratory, namely, the strain typing Polymorphic Amplified Typing Sequence (PATS) and the bovine rectal-anal junction squamous epithelial (RSE) cell and HEp-2 cell adherence assays. We specifically chose O157 strain EDL933 since this was the first O157 to be sequenced [31] and is commonly used as a control in O157-related experiments [32-38]. As shown in Table 1, we also evaluated mutants of O157 strain EDL933 demonstrating loss of the toxin genes or decreased LEE protein expression in the adherence assays to verify the role of Stx and LEE in the observed phenotype [39-42].
2. MATERIALS AND METHODS
2.1. Bacterial Strains
O157 strain EDL933 WT ATCC 43895 (ATCC® 43895™; ATCC, Manassas, VA) was included as the comparative control in both cell-adherence assays and the PATS assay (Table 1). E. coli strain K-12 MG1655 and O157 strain 86-24 (86-24-wild-type or 86-24WT) [43, 44] from the NADC stocks were also included as controls only for PATS (Table 1). The test strains evaluated in this study are shown in Table 1 and included O157 EDL933 strains, with or without antibiotic resistance, obtained from three different sources. In addition, O157 strain EDL933 with mutations resulting in the loss of the toxin genes by targeted mutagenesis [39] or with restrictive mutations, in spontaneous streptomycin-resistant mutants [42], in the ribosomal protein S12 (rpsL) conferring resistance to streptomycin and decreasing LEE protein expression were acquired to evaluate the role of Stx and/or LEE proteins in the adherence phenotype (Table 1) [39-42]. All strains were confirmed to be O157 by latex agglutination tests (E. coli O157 latex, Oxoid Diagnostic Reagents, Oxoid Ltd., Hampshire, UK).
2.2. Adherence Assays
Assays were performed with two different cell types using previously standardized protocols [21, 45-47]. As per the protocols, all bacterial strains were cultured overnight in Dulbecco’s modified Eagle’s medium with low glucose [DMEM-LG (Invitrogen, Carlsbad, CA)] at 37°C without aeration, washed, and re-suspended in DMEM with no glucose (DMEM-NG; Invitrogen) [20, 47, 48].
| Bacteria | Source |
|---|---|
| EDL933 WT ATCC 43895 = E. coli O157:H7 strain EDL933; Stx1+, Stx2+, Intimin+; ATCC® 43895™ | American Type Culture Collection (ATCC), Manassas, VA |
|
Source A: EDL933- T = Stx1+, Stx2+, Intimin+ TUV 933-1 = cured of the phage carrying stx2 TUV 933-2 = isogenic stx1 deletion mutant TUV 933-2c = cured of the phage carrying stx1 |
Obtained from Dr. Saul Tzipori, Tufts University, North Grafton, MA [39, 40]. |
|
Source B: EDL933-R = Rifampicin resistant; Stx1+, Stx2+, Intimin+ EDL933-R Δstx1 EDL933-R Δstx2 EDL933-R Δstx1, Δstx2 |
Obtained from Dr. Ching-Hong Yang, University of Wisconsin, Milwaukee, WI [41]. |
|
Source C: EDL933-E = Stx1+, Stx2+, Intimin+ Spontaneous, streptomycin resistant mutants of EDL933-E: EDL933-E strR1 = non-restrictive K42R mutation in RpsL EDL933-E strR2 = restrictive K42T mutation in RpsL EDL933-E strR3 = restrictive K42N mutation in RpsL |
Obtained from Dr. Edward Dudley, Pennsylvania State University, University Park, PA [42]. |
2.2.1. Recto-anal Junction Squamous Epithelial (RSE) Cell Assay
The RSE adherence assay was repeated twice, with 4 technical replicates per bacterial strain each time. As previously described, RSE cells, collected from RAJ tissue at necropsies, were suspended in DMEM-NG to a final concentration of 105 cells/ml and mixed with bacteria at a bacteria:cell ratio of 10:1 [20, 45-47]. The mixture was incubated at 37°C with aeration (110 rpm) for 4 h, pelleted, washed, and reconstituted in 100 µl of double-distilled water (dH2O). Drops of the suspension (2 µl) were placed on Polysine slides (Thermo Scientific/Pierce, Rockford, IL), dried, fixed, and stained with fluorescent-tagged antibodies specific to the O157 antigen and cytokeratins of the RSE cells as previously described [20]. The fluorescein isothiocyanate (FITC; green)–labelled goat anti-O157 (KPL, Gaithersburg, MD, USA) antibody was used to target the O157 antigen and the mouse anti-(PAN) cytokeratins (AbD Serotec, Raleigh, NC, USA) in combination with Alexa Fluor 594 (red)–labelled goat anti-mouse IgG (H+L; F(ab´)2 fragment) (Invitrogen) was used to target the RSE cell cytokeratin [20].
Adherence patterns on RSE cells were qualitatively recorded as diffuse, aggregative, or nonadherent, and quantitatively as the percentages of RSE cells with or without adhering bacteria [21, 46, 47]; adherence was recorded as strongly adherent when more than 50% of RSE cells had 10 adherent bacteria, moderately adherent when 50% or less of the RSE cells had 1 to 10 adherent bacteria, and nonadherent when less than 50% of the RSE cells had only 1 to 5 adherent bacteria. RSE cells with no added bacteria were subjected to the assay procedure and used as negative controls to confirm the absence of pre-existing O157 bacteria. Quantitative data was evaluated for statistical significance against data from O157 strain EDL 933WT ATCC 43895 using one-way ANOVA with Dunnett’s test; p < 0.05 was considered significant (GraphPad Prism version 8.0.0, GraphPad Software, San Diego, CA).
2.2.2. HEp-2 Cell Adherence Assay
Adherence patterns displayed by bacterial strains on HEp-2 cells (human epidermoid carcinoma of the larynx cells with HeLa contamination) (ATCC® CCL-23™) were determined using the same growth conditions as those used for the RSE adherence assay and as described previously [21, 47, 48]. The HEp-2 adherence assay was performed twice, with four technical replicates per bacterial strain each time. Slides were stained with the FITC–labelled goat anti-O157 antibody (KPL) targeting the O157 antigen and the Alexa Fluor 594-labelled Phalloidin (Thermo Scientific) targeting the HEp-2 cell actin filaments [20, 21, 47]. The adherence was qualitatively and quantitatively evaluated as described for the RSE adherence assay.
2.3. Polymorphic Amplified Typing Sequence (PATS)
Colony lysates, prepared from each bacterial strain cultured on Luria Bertani agar plates, were each tested in triplicate to confirm the PATS profiles as described previously [49-52]. Briefly, primer pairs targeting the 8 polymorphic XbaI-, 7 polymorphic AvrII- restriction enzyme sites, and the four virulence genes encoding Shiga toxin 1 and 2 (Stx1 and Stx2), Intimin-γ (eae), and hemolysin-A (hlyA) were used to generate amplicons from the colony lysates in a hot start, touchdown PCR reaction [47, 49, 50, 52, 53]. PCR reactions amplifying the AvrII- restriction enzyme site were purified using the QIAquick PCR purification kit (Qiagen, Valencia, CA), and digested with the AvrII restriction enzyme (New England Biolabs, Beverly, MA) to confirm the presence of the restriction site. All reactions were analyzed by electrophoresis on 3% agarose gels stained with ethidium bromide. The presence or absence of amplicons for XbaI and the virulence genes was recorded using “1” and “0”, respectively. The absence of an AvrII amplicon was recorded as “0”, and the presence of restriction site with a SNP as “1”, “2” for an intact restriction site, and “3” for a restriction site duplication [49, 50, 53].
2.4. Curli Production Assays
As described previously, congo red binding and biofilm formation assays were set up to determine levels of curli fimbriae formation and are described briefly below [46]. E. coli strain K-12 MG1655, and O157 strains 86-24 WT and EDL933 WT ATCC 43895 that are established curli fimbriae producers were included as positive controls while E. coli O111 strain MB1068W that lacks the curli fimbriae was included as a negative control in these assays [46, 54].
2.4.1. Congo Red Binding Assays
Congo red indicator (CRI) plates were prepared with Luria-Bertani broth with no salt (LBNS) agar, mixed with 40 µg/ml Congo red dye and 10 µg/ml Coomassie brilliant blue. Following subculture, the CRI plates were incubated at 26°C for 2 to 3 days before reading for dye assimilation by curli-producing bacteria [46]. On CRI, curli-producing (C) bacteria form red (R; C+) or super-red (SR; C++) colonies while curli-deficient form white colonies (W; C-).
2.4.2. Biofilm Formation Assays
Bacteria were grown in LBNS broth at 26°C overnight. Each culture was diluted in fresh LBNS broth at a concentration of 105 cells/ml, and 5 ml of diluted culture was dispensed into a borosilicate glass tube and incubated statically at 26°C aerobically for 5 days. After incubation, the planktonic cells were removed gently, and the tubes rinsed and stained with 0.1% crystal violet at room temperature for 30 min. Post-washes the crystal violet bound to the glass tube was solubilized in 1.0 ml of 33% acetic acid, and the absorbance was determined at 570 nm using a microplate reader (SpectraMax 340; Molecular Devices, Sunnyvale, CA). An average of three biological replicates were used to represent the data.
3. RESULTS AND DISCUSSION
3.1. Polymorphic Amplified Typing Sequences (PATS) Identified Genetic Differences, and Targeted Mutations, in the EDL 933 Strains Tested
The O157 strain EDL933 WT ATCC 43895 with the four virulence genes stx1, stx2, eae, and hlyA, the O157 strain 86-24 lacking the stx1 gene, and the avirulent E. coli K-12 lacking all the virulence genes along with some of the genetic regions in the O157 genome were included as comparative controls in the PATS assay (Table 2) [53]. Latex agglutination confirmed the serogroup of the O157 control and test strains. Using PATS, we were able to identify distinct genetic differences in the EDL933-T and TUV 933-1 strains, compared to the prototypical EDL933 WT ATCC 43895 (Table 2) [49, 50, 53]. All other EDL933 strains shared the same PATS profile with EDL933 WT ATCC 43895 (Table 2). In addition, PATS profiles of the Stx deletion mutants accurately determined the Stx gene deletions, thereby demonstrating that these strains were isogenic derivatives of the EDL933 WT ATCC 43895 strain (Table 2).
The profile of EDL933-T differed from the EDL933 WT ATCC 43895 profile in 5 of the 8 XbaI- and 4 of the 7 AvrII- polymorphic sites tested (Table 2). However, the TUV 933-1 strain only lacked only one of the 8 XbaI- polymorphic sites besides the stx2 gene (Table 2). Deletions and single nucleotide polymorphisms contributing to the variant PATS profile in EDL933-T and TUV 933-1 were in unknown regions (IK114, IKNR16), cryptic prophages (IK118, IK123, IK127, IKNR3), bacteriophage encoding Stx2 (IKB3), region encoding anaerobic respiration genes (IKNR27) or integrase, helicase (IKNR33) [51, 53]. Since the differences were in these known polymorphic regions, it is possible that the mutations were acquired over time in the process of serial passage and/or storage [28, 30, 55-58].
3.2. The EDL 933 Strains Demonstrated Different Adherence Profiles on the Bovine RSE Cells without Any Influence from the Stx and LEE Genes
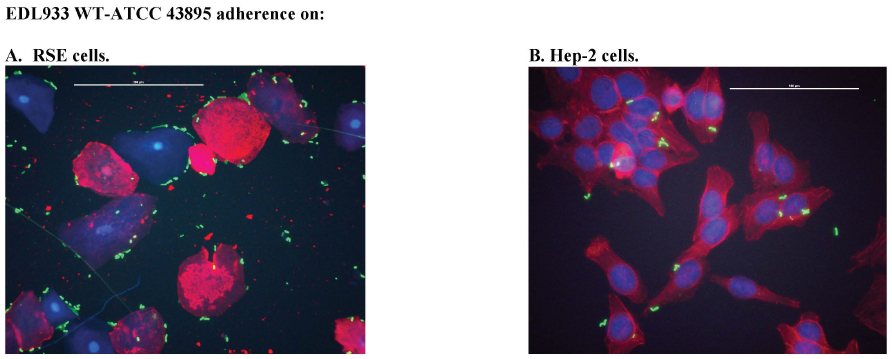
Phenotypic variations could result in colonization/ persistence differences of O157 strains in animal experiments. For instance, super-shed O157 have been consistently shown to have an aggregative, strong adherence on RSE cells [47] that may contribute to their persistence at the bovine rectoanal junction. In this study, the parent, wild-type EDL933 strains, from different sources, demonstrated varied adherence patterns on bovine RSE cells. The EDL933-R and EDL933-E strains demonstrated an aggregative, strong adherence pattern on RSE cells compared to the aggregative, moderate adherence of the control EDL933 WT ATCC 43895 (Table S1, Figs. 1 and 2) strain. In sharp contrast, EDL933-T exhibited a diffuse, moderate adherence pattern (Table S1, Figs. 1 and 2) similar to the O157 strain 86-24WT in previous studies [20, 46].
Since it has been reported that Stx may play a role in enhancing O157 adherence to HEp-2 cells and mouse intestinal cells [59] by possibly increasing expression of the alternate receptor for intimin, namely nucleolin, we evaluated Stx mutants of the EDL933-T and EDL933-R strains in the eukaryotic cell adherence assays [39-41, 47]. In the assay with the bovine RSE cells, contrasting results were obtained in that the deletion of Stx genes increased O157 adherence to RSE cells rather than decreasing the same (Table S1, Figs. 1 and 2). Even when the parent isolate exhibited an aggregative, strong adherence on RSE cells, as with EDL933-R, the Stx mutants had slightly greater numbers of bacteria attached per RSE cell contributing to the same phenotype (Fig. 2). Hence, Stx appeared not to have any direct role, rather a modulating role, in O157 adherence to bovine RSE cells (Table S1).
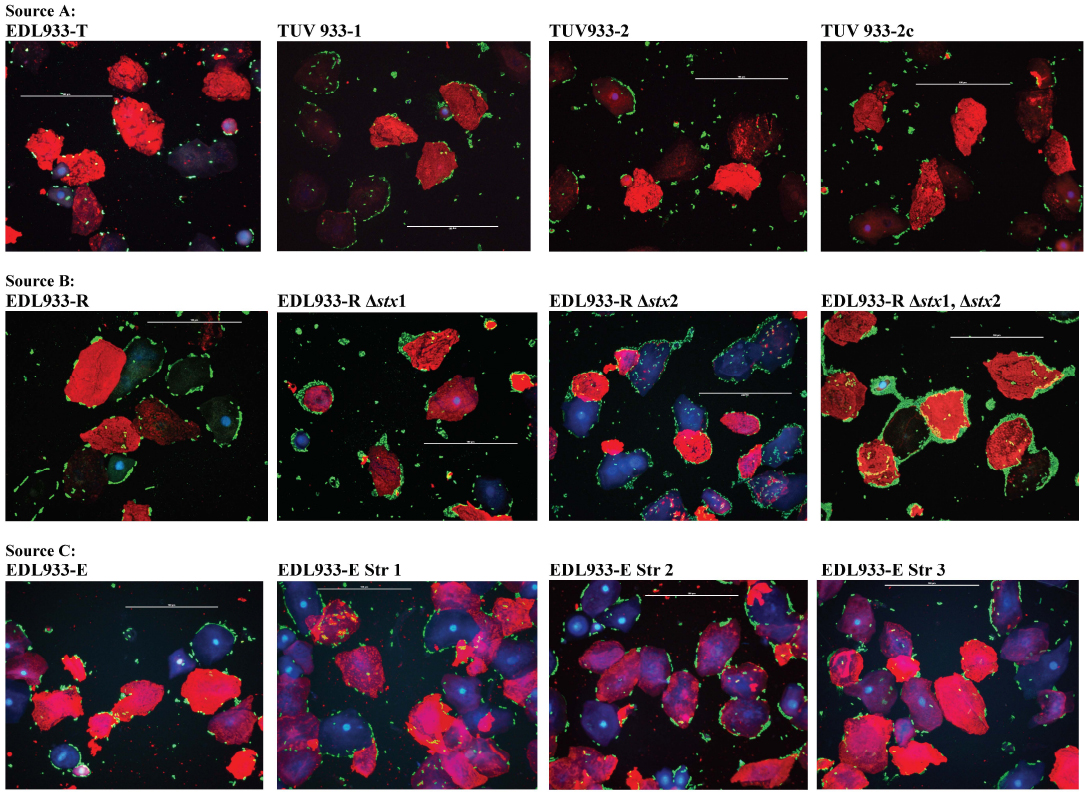
Previously, we had determined that LEE-encoded proteins, although critical for O157 adherence to human HEp-2 cells and FAE cells of the bovine rectoanal junction, have no effect on adherence to bovine RSE cells [21]. In this study, the spontaneous streptomycin mutants (EDL933-E -str1, -str2, -str3), with reduced LEE expression, did not differ from the parent strain EDL933-E in their RSE cell adherence phenotype [42] (Table S1, Figs. 1 and 2). This observation further supports a role for adhesins other than LEE-encoded proteins in RSE cell adherence as previously reported [21].
The quantitative difference in RSE cell-adherence that was observed between all wild-type and mutant EDL933 strains when compared to EDL933 WT ATCC 43895 was statistically significant (p <0.05) for the >10 bacteria/cell counts (Table S1). The exception was the EDL933-T strain which demonstrated moderate adherence similar to EDL933 WT ATCC 43895 (Table S1). Overall, no definite correlation between the PATS-based genotype and the differences in the RSE cell adherence phenotype could be established, but the experiments with the isogenic mutants established that Stx and LEE did not play a role in the adherence phenotypes. Considering that O157 strains have a plastic genome, it is likely that these EDL933 strains have other mutations in the genome, not targeted by PATS, influencing the adherence phenotype, which needs to be further evaluated [25].
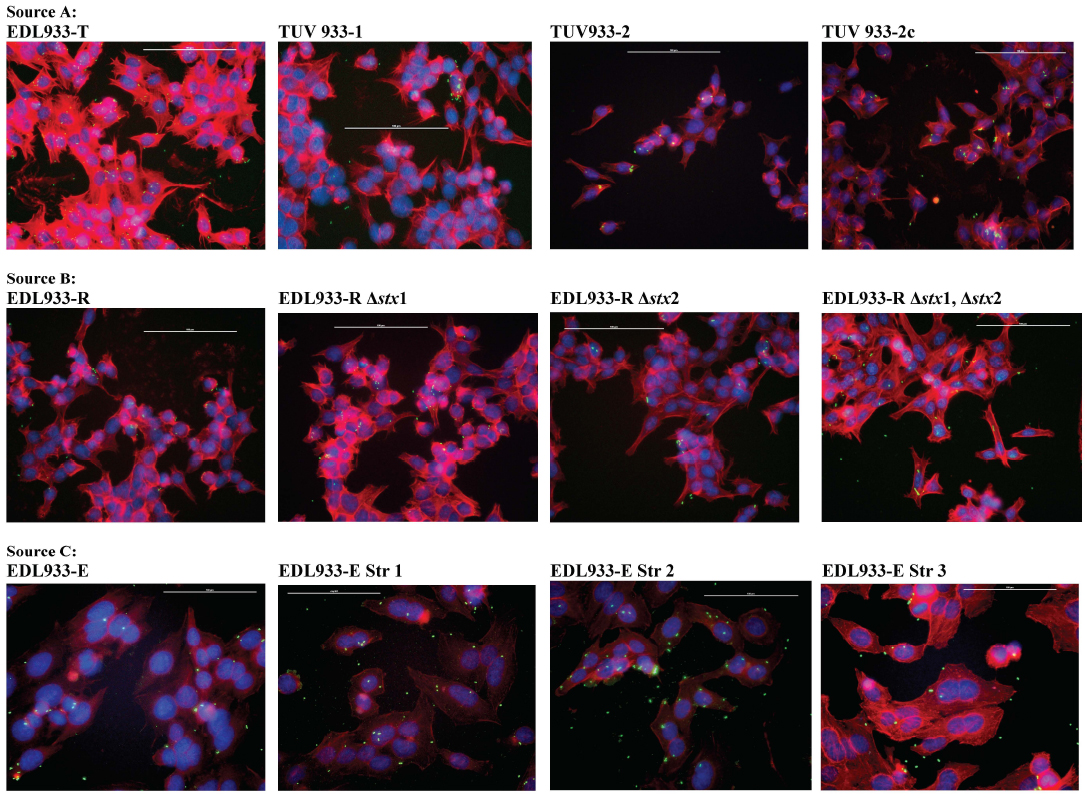
With the HEp-2 cells, none of the EDL933 parent and mutant strains exhibited any significant differences in the adherence patterns (Table S2, Figs. 1-3). Chen et al. had reported that the mutants of O157 strain EDL933-E, with restrictive mutations in the ribosomal subunit protein S12, have reduced LEE expression and hence a corresponding decrease in the attaching-effacing lesions on HeLa cells [42]. However, in this study, these mutants demonstrated no significant differences in the adherence phenotype on HEp-2 cells compared to the parent strain (Table S2, Figs. 1 and 3). Likewise, the lack of Stx genes did not alter the HEp-2 cell adherence phenotype (Table S2, Figs 1 and 3) (Table S2, Figs. 1 and 3). This observation supports our previous report that the O157 adherence phenotype varies between epithelial cells of different host origins and perhaps a complete absence of intimin, not just minimal expression, is needed to prevent O157 adherence to HEp-2 cells [20-22, 47].
3.3. The Variant Diffuse Adherent Phenotype of EDL 933-T Strain may have been caused by Decreased Curli Fimbriae Production
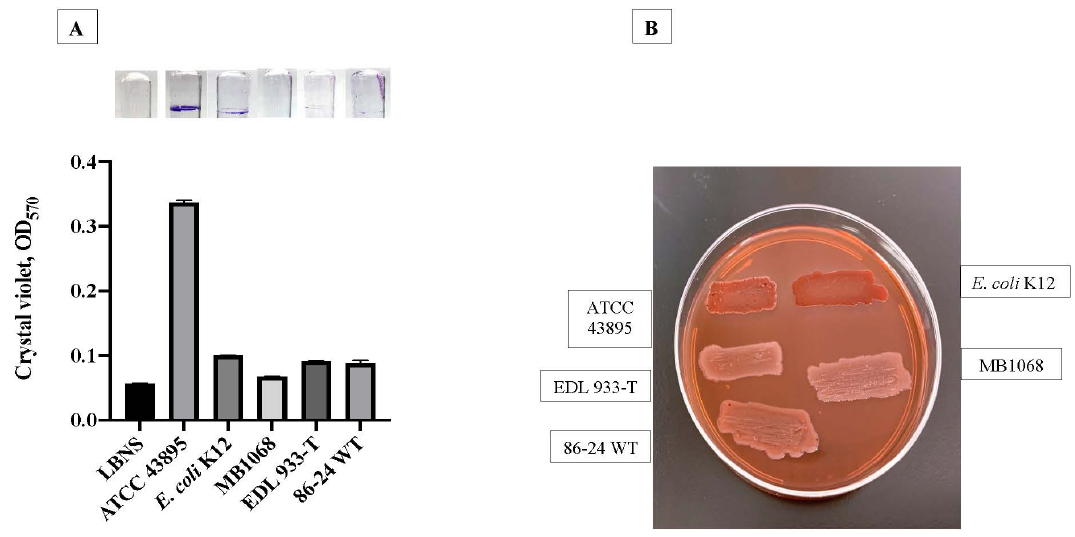
Since curli fimbriae contribute to biofilm formation by aggregation [46, 54, 60], we hypothesized a role for curli in the formation of aggregates on RSE cells. Hence, we evaluated curli production in EDL933-T, which was the only EDL933 strain to demonstrate a variant diffuse adherent phenotype on RSE cells. In comparison to the curli producing control strains, EDL933-T colonies on CRI plates were white, indicating lack of or minimal curli production (W, C-) (Fig. 4). Biofilm formation corresponded with the amount of curli produced by the tested strains with EDL933-T yielding significantly less crystal violet bound biofilms compared to positive control strains as shown in Fig. (4). Since the complete absence of curli results in increased adherence of O157 to RSE cells [46], the current observation further supports the previously reported modulatory role for curli in adherence of some O157 strains to RSE cells. Specifically, the suboptimal concentration of curli appears to alter the qualitative adherence phenotype in EDL933 strains changing it from aggregative to diffuse while optimal concentrations support the aggregative phenotype; the complete absence of curli, on the other hand permits other adherence factors to directly interact with the eukaryotic cell allowing for quantitatively increased adherence [46].
| XbaI | AvrII | Virulence Genes | ||||||||||||||||||
| Isolate | IK8 | IK25 | IK114 | IK118 | IK123 | IK127 | IKB3 | IKB5 | IKNR3 | IKNR7 | IKNR10 | IKNR12 | IKNR16 | IKNR27 | IKNR33 | stx1 | stx2 | eaeA | hlyA | |
| Controls | ||||||||||||||||||||
| K-12 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 0 | 0 | |
| 86-24WT | 1 | 1 | 0 | 1 | 1 | 1 | 0 | 1 | 2 | 2 | 2 | 2 | 2 | 3 | 0 | 0 | 1 | 1 | 1 | |
| EDL933WT ATCC 43895 |
0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | |
| Source A: | ||||||||||||||||||||
| EDL933-T | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 2 | 2 | 2 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | |
| TUV 933-1 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 0 | 1 | 1 | |
| TUV 933-2 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 1 | 1 | 1 | |
| TUV 933-2c | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 1 | 1 | 1 | |
| Source B: | ||||||||||||||||||||
| EDL933-R | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | |
| EDL933-R Δstx1 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 1 | 1 | 1 | |
| EDL933-R Δstx2 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 0 | 1 | 1 | |
| EDL933-RΔstx1, Δstx2 | 0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 0 | 1 | 1 | |
| Source C: | ||||||||||||||||||||
| EDL933-E EDL933-E strR1 EDL933-E strR2 EDL933-E strR3 |
0 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | |
CONCLUSION
Overall, our results demonstrate that the EDL933 strains of O157, obtained from different sources, can exhibit varied genetic and adherence profiles on eukaryotic cells (Table 2 and Table S1). The O157 strain EDL933-T with the most variations in the PATS profile (Table 2) also differed in the qualitative adherence profile on bovine RSE cells (Table S1). This strain adhered RSE cells in a diffuse, moderate pattern unlike its Stx mutants (aggregative, strong) or EDL933 WT ATCC43895 (aggregative, moderate) (Table S1, Fig. 2). Interestingly, EDL933-R and EDL933-E, along with the corresponding isogenic mutants, also demonstrated an aggregative, strong adherence pattern on RSE cells that differed from EDL933 WT ATCC 43895. A definite correlation between the genotype and phenotype differences could not be established, but the experiments with the isogenic mutants established that Stx and LEE genes were not contributory factors in the adherence phenotypes. A unique association between decreased curli production and the variant diffuse adherence phenotype of EDL933-T on RSE was observed. Since curli fimbriae contribute to biofilm formation, these may also be involved in the aggregative adherence phenotype characteristic of EDL933 strains. However, the absence of curli can also increase adherence to RSE cells. Additional transcriptomics or proteomics experiments are being planned to identify other factors contributing to the varied interactions with RSE cells. In addition, experiments are being planned with the EDL933 WT ATCC 43895 strain to dissect out storage and continuous passage conditions that may initiate genetic mutations resulting in altered strains; these may also provide insights into other mutations that may have influenced the variations in the adherence profiles. Using a combination of PATS and RSE cell adherence assays, we were able to gather information on strain differences which would be especially relevant in adherence and colonization studies. Overall, our study highlights the importance of carefully evaluating bacterial strains acquired from different sources to ensure the reproducibility of results within and between laboratories.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
All data is included in the manuscript (main documents + supplemental files) and there is no additional data to deposit.
FUNDING
This work was supported by USDA-ARS CRIS projects 5030-32000-100-00D and 5030-32000-112-00D.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Excellent technical support provided by Bryan Wheeler is acknowledged. The Animal Resource Unit is gratefully acknowledged for the assistance provided during the collection of RSE cells from the terminal rectum of cattle. We also gratefully acknowledge: (i) Dr. Saul Tzipori, Tufts University, North Grafton, MA., (ii) Dr. Ching-Hong Yang, University of Wisconsin, Milwaukee, WI., and (iii) Dr. Edward Dudley, Pennsylvania State University, University Park, PA., for providing the EDL933 strains and mutants.
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publishers web site along with the published article.


