All published articles of this journal are available on ScienceDirect.
Occurrence and Characteristics of the Extended-spectrum Beta-lactamase-producing Enterobacterale in a Hospital Setting
Abstract
Objectives:
Infectious diseases caused by ESBL-producing Enterobacterale are an emerging problem worldwide, which increase hospital costs, empirical treatment failure, together with rates of morbidity and mortality. The aims of this study were to determine the antibiotic-resistant patterns and the frequency of blaTEM, blaCTX-M and blaSHV genes among Enterobacterale.
Methods:
A total of 239 non-repeated clinical isolates of the Enterobacterale family, including 202 (84.5%) Escherichia coli and Klebsiella pneumoniae 25 (12.3%) were collected. Identification and susceptibility tests were carried out on 60 (25.9%) ESBL phenotypes using biomérieux VITEK® 2 compact system. Isolates showing ESBL positivity by the phenotypic method were all screened for blaTEM, blaCTX-M and blaSHV genes by PCR.
Results:
The prevalence of ESBL-producing Enterobacterale isolates was found to be 25.9%. These ESBL-producing isolates displayed an increasing rate of resistance for aminopenicillins (ampicillin) (96.3%), followed by piperacillin-tazobactam (54.2%), amoxicillin/clavulanic acid (45.8), the cephalosporin groups, ceftriaxone (52.4%), ceftazidime (46.7%), and fluoroquinolone (ciprofloxacin) (42.7%). Both the organisms showed a higher susceptibility to the carbapenems (Imipenem) and aminoglycosides (Amikacin). Out of the 62 ESBL positive isolates, 11 (17.7%) carried the TEM gene, 22 (35.5%) carried the CTX-M gene alone, 8 (12.9%) carried both TEM and CTX-M genes, 1(1.6%) carried both SHV and CTX-M genes, and 20 (32.3%) carried the TEM, SHV, and CTX-M genes. SHV gene alone was not reported in any of the isolates.
Conclusion:
The study indicated a moderate occurrence of ESBL-producing Enterobacterale with CTX-M being the most dominant gene. The co-existence of all three genes on many occasions suggested the carriage of multiple plasmids with three resistance genes that might pose a serious epidemiological, clinical and public health threat.
1. INTRODUCTION
Extended-Spectrum Beta-Lactamases (ESBLs) are rapidly evolving plasmid-mediated enzymes produced by certain bacteria that can hydrolyze extended-spectrum cephalosporin, thus making them effective against beta-lactam antibiotics [1, 2]. ESBLs are found in gram-negative bacteria, especially in Enterobacterale species [1], the main bacterial family associated with ESBL production, of which Escherichia coli and klebsiella pneumoniae are the most important. These organisms are of medical importance, causing infections in the various body systems. The dissemination of ESBL-producing multi-drug resistant Enterobacterale has resulted in increased bacterial resistance to antibiotics in healthcare settings and in the community worldwide [3].
The plasmid-mediated lactamase-producing isolates of the family Enterobacterale mainly possess the blaTEM gene which can hydrolyze penicillin and first-generation cephalosporins and the blaCTX-M gene that preferentially hydrolyze cefotaxime [4]. Other types of ESBLs genes include SHV, which is responsible for plasmid-mediated ampicillin resistance. The majority of the ESBLs are derivatives of TEM or SHV enzymes, which are most often found in the enterobacteria E. coli and K. pneumonia [5].
Because of the global spread of CTX-M enzyme, ESBL are the cause of increasing numbers of both nosocomial and community infections [6]. Rates of ESBL production among Enterobacterale range between 6-38.5% in the Kingdom of Saudi Arabia [7, 8]. ESBL producing Enterobacterale in Saudi Arabia is dominated by E. coli where both hospital and community-based studies have shown that more than 70% harbored the gene blaCTM-X [9].
Detection of these genes in ESBL-producing bacteria by the most sensitive molecular methods can give valuable information about its epidemiology and help in the reasonable treatment of infections [10].
In Saudi Arabia, few studies have been performed to determine the prevalence of ESBL-producing Enterobacterale and little is known about their molecular characteristics. However, the few studies conducted on ESBL phenotypes isolated from clinical samples previously indicated a predominance of blaCTX-M-like genes [8, 9, 11, 12]. Therefore, the aims of this study were to determine the prevalence of ESBL-producing Enterobacterale in a hospital setting from various clinical specimens taken from patients diagnosed with various infections and to detect the frequency of TEM, SHV, and CTX-M genes in an ever-changing prevalence and resistant pattern of ESBL-producing enterobacteria.
2. MATERIALS AND METHODS
The ethical committee of Prince Sultan Military College of Health Science approved this study. This retrospective study included 239 isolates diagnosed with culture-positive Enterobacterale at the Medical Laboratory of King Fahad Military Medical Complex, Dhahran, from various clinical specimens from March 2014 to June 2014. Isolates were then received at the Microbiology Laboratory of Prince Sultan Military College of Health Sciences (PSMCHS) where identification and susceptibility tests were carried out using biomérieux VITEK® 2 compact system (bioMérieux, Marcy l’Etoile, France) according to the manufacturer’s instructions (2009 FDA breakpoints).
Gram-negative (GN) cards and antibiotic susceptibility cards (AST) were used for the identification and the antibiotic sensitivity profile of the isolates, respectively. Antimicrobial susceptibility testing was performed using software version 06.01 and the AST-GN291 and AST-XN06 cards, according to the manufacturer’s instructions. Susceptibility cards used included aminoglycosides (amikacin, gentamicin); aminopenicillins (ampicillin); cephalosporin (cefepime, ceftazidime, ceftriaxone, cefuroxime); carbapenems (imipenem); amoxicillin/clavulanate (augmentin); trimethoprim/sulfamethoxazole (Bactrim); fluoroquinolone (ciprofloxacin); and piperacillin-tazobactam. Identification and susceptibility cards were inoculated and interpreted according to the manufacturer’s instructions. The quality control strains tested with each run included the ESBL-producing strains E. coli ATCC 25922 and K. pneumoniae ATCC 700603 for the antimicrobial identification and susceptibility testing.
Out of 239 non-repeated isolates, 62 were ESBL producers, which were further tested for the detection of ESBL genes in the PSMCHS Molecular Biology Laboratory. DNA was extracted using the method previously described [13]. The PCR mixture was made using QIAGEN Fast Cycling PCR Kit and the primers, which were designed and procured by the Eurofins Company (Eurofins Advantar Inc, San Diego USA). PCR was performed using CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad Laboratories, Inc. USA). SHV, TEM, and CTX-M primers were used on all of the 62 ESBL-producing Enterobacterale isolates (Table 1). The following PCR protocol was used for all primers as previously described [11]: initial denaturation (hot start) at 95°C for 5 minutes, followed by 30 cycles of denaturation, annealing, and extension at 92°C, 61°C, and 72°C, respectively, for 30 seconds each. The mixture was then incubated for final elongation at 72°C for 5 minutes.
PCR products were then separated on 2% agarose electrophoresis gel and visualized using Syngene U Genius Gel Imaging System (Cambridge Scientific, Watertown, Massachusetts) for the presence of 403 bp, 293 bp, and 569 bp bands representing the TEM, SHV, and CTX-M primers, respectively.
3. RESULTS
A total of 239 non-repeated multi-resistant clinical isolates of the Enterobacterale family, including Escherichia coli 202 (84.5%) and Klebsiella pneumoniae 25 (10.5%), while 12 (5%) were identified as Proteus mirabilis and Citrobacter diversus. Out of the total Enterobacterale isolates recovered, 62 or 25.9% were ESBL phenotypes. E. coli, and K. pneumoniae represented by 48 (77.4%), and 12 (19.4%), respectively. Most of the isolates were recovered from urine specimen 198 (70.3%), followed by wound swabs being 23 (9.6%), vaginal swab being 10 (4.2%), sputum being 4 (1.7%), and tissue being 4 (1.7%). (Table 2).
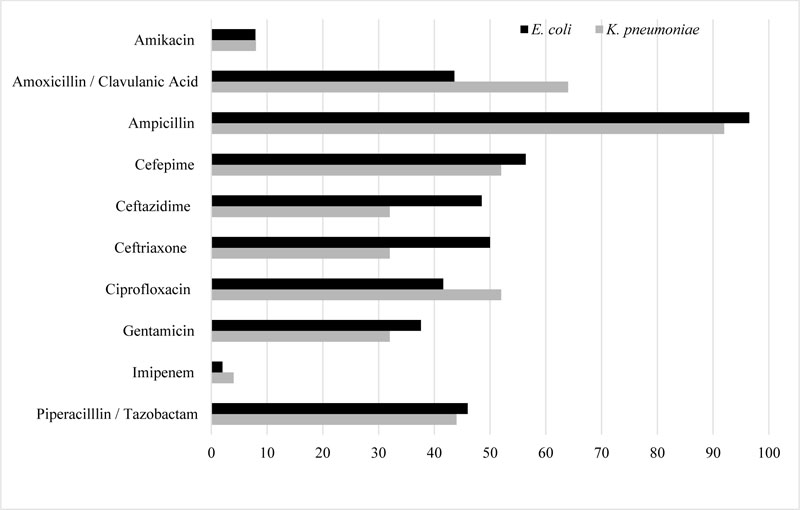
ESBL producers demonstrated the highest resistance rates to ampicillin (96.3%) followed by piperacillin/tazobactam (54.2%), ceftriaxone (52.4%), ceftazidime (46.7%), amoxicillin/clavulanic acid (45.8), and ciprofloxacin (42.7%). E. coli was the most resistance to ampicillin (96.5%), cefepime (56.4%), ceftriaxone (50.0%), ceftazidime (48.5%), piperacillin/tazobactam (46.0%), amoxicillin/Clavulanate (43.6%), and ciprofloxacin (41.6%). K. pneumoniae was resistant to ampicillin (92.0%), amoxicillin/clavulanate (64.0%), ciprofloxacin (52%), cefepime (52.0%), and piperacillin/tazobactam (44.0%). ESBL producers showed a higher susceptibility to imipenem and amikacin as the resistance rate was 2.2% and 7.9%, respectively. The resistance pattern is almost similar in both organisms. However, E. coli was more resistant to ceftriaxone and ceftazidime than K. pneumoniae. On the other hand, K. pneumoniae showed more resistance to amoxicillin/clavulanic than E. coli (Table 3, Fig. 1).
Out of the 62 ESBL positive isolates, 11 (17.7%) carried the TEM gene, 22 (35.5%) carried the CTX-M gene alone, 8 (12.9%) carried both TEM and CTX-M genes, 1(1.6%) carried both SHV and CTX-M genes, and 20 (32.3%) carried the TEM, SHV, and CTX-M genes. SHV gene alone was not reported in any of the isolates. The results indicated a Predominance of CTX-M gene among ESBL isolates (Table 4).
| Primers | Forward Primers 5′-3′ | Reverse Primers 3′ - 5′ | Reference |
| SHV | AGGATTGACTGCCTTTTTG | CGAGTAGTCCACCAGATCCT | EF125011 |
| TEM | TTTCGTGTCGCCCTTATTCC | ATCGTTGTCAGAAGTAAGTTGG | AB282997 |
| CTX-M | CGCTGTTGTTAGGAAGTGTG | GGCTGGGTGAAGTAAGTGAC | DQ303459 |
| ESBL genotype | Number of isolates (% ) | |||||||
| E. coli | k. pneumoniae | Others | Total | |||||
| MRD | ESBL | MRD | ESBL | MRD | ESBL | MRD | ESBL | |
| Urine | 168 | 40 (23.8) | 18 | 9 (50.0) | 12 | 2 (16.7) | 198 | 51 (25.8) |
| Wound swab | 18 | 4 (22.2) | 5 | 2 (40.0) | 0 | 0 (0.0) | 23 | 6 (26.1) |
| Vaginal swab | 10 | 2 (20.0) | 0 | 0 (0.0) | 0 | 0 (0.0) | 10 | 2 (20.0) |
| Sputum | 4 | 1 (25.0) | 0 | 0 (0.0) | 0 | 0 (0.0) | 4 | 1 (25.0) |
| Tissue | 2 | 1 (50.0) | 2 | 1 (50.0) | 0 | 0 (0.0) | 4 | 2 (50.0) |
| Total | 202 | 48 (23.8) | 25 | 12 (48.0) | 12 | 2 (16.7) | 239 | 62 (25.9) |
ESBL = Extended spectrum beta-lactamases isolates.
| Antibiotic (Class) | MIC range mg/L | E. coli n =202 |
K. pneumoniae n =25 |
Total n =227 |
| Amikacin | 8 – 64 | 16 (7.9) | 2 (8.0) | 18 (7.9) |
| Amoxicillin/Clavulanic acid | 4/2 – 32/16 | 88 (43.6) | 16 (64.0) | 104 (45.8) |
| Ampicillin | 4 – 32 | 195 (96.5) | 23 (92.0) | 206 (96.3) |
| Cefepime | 2 – 32 | 114 (56.4) | 13 (52.0) | 58 (27.1) |
| Ceftazidime | 1 – 32 | 98 (48.5) | 8 (32.0) | 106 (46.7) |
| Ceftriaxone | 1 – 32 | 101 (50.0) | 8 (32.0) | 119 (52.4) |
| Ciprofloxacin | 0.5 – 4 | 84 (41.6) | 13 (52.0) | 97 (42.7) |
| Gentamicin | 4 – 32 | 76 (37.6) | 8 (32.0) | 84 (37.0) |
| Imipenem | 1 – 12 | 4 (2.0) | 1 (4.0) | 5 (2.2) |
| Piperacilllin/Tazobactam | 2/4 – 48/8 | 93 (46.0) | 11 (44.0) | 104 (54.2) |
| ESBL genotype | Number of isolates (% ) | Total | |
| E. coli | K. pneumoniae | ||
| TEM | 7 (13.0) | 4 (57.1) | 11(17.7) |
| CTX-M | 20 (37.0) | 2 (28.6) | 22 (35.5) |
| TEM and CTX-M | 8 (14.8) | 0 (00) | 8 (12.9) |
| SHV and CTX-M | 0 (00) | 1 (14.3) | 1 (1.6) |
| TEM, SHV, and CTX-M | 19 (35.2) | 1 (14.3) | 20 (32.3) |
| Total | 54 (100) | 7 (100) | 62 (100) |
4. DISCUSSION
Enterobacterale are among the most important causes of nosocomial and community-acquired infections. Infections caused by these microorganisms are mainly treated with beta-lactams and fluoroquinolones [6]. However, resistance to these compounds is being reported more frequently [14].
The current findings have detected a high occurrence of ESBLs (25.9%) among the Enterobacterale members isolated from various specimens. Previous studies have reported various ranges of ESBL prevalence of 10% to 36% [8, 15-18]. ESBL-producing Enterobacterale prevalence varies greatly and has increased over time worldwide. The pooled estimated prevalence was the highest in the West Pacific (46%), followed by southeast Asia and Africa (22%), the Eastern Mediterranean (15%) and Northern Europe (4%) [19]. Factors associated with a higher risk of colonization were antibiotic use and international travel in the previous year [19].
In the present study, ESBL-producing Enterobacterale displayed an increasing rate of resistance for aminopenicillins (ampicillin), followed by piperacillin-tazobactam, cephalosporin groups (ceftriaxone, ceftazidime, cefepime, ciprofloxacin), and fluoroquinolone (ciprofloxacin). The study has shown that both E. coli and K. pneumoniae have developed a high rate of resistance to ampicillin, piperacillin-tazobactam, amoxicillin/clavulanic acid, the cephalosporin groups, ceftriaxone, ceftazidime, and fluoroquinolone (ciprofloxacin) (42.7%). Both the organisms showed a higher susceptibility to the carbapenems (imipenem) and the aminoglycosides (Amikacin). This resistance pattern has been observed in many other studies [8, 9, 12, 15, 17, 18] and might probably be attributed to the extensive use of these antibiotics.
Our study also included the molecular aspect of resistance through the identification of genes carried by these resistant isolates. The ESBL types detected in these isolates belonged to the CTX-M and TEM, while the SHV type was not detected alone in any of the tested isolates.
The CTX-M genes are the most prevalent (35.5%) among the isolates of this study. This finding agrees with other studies conducted elsewhere in the Middle East and many parts of the world [7, 20]. Among the different ESBL families, CTX-M has become the most prevalent worldwide, both in nosocomial and community-acquired infections [20].
The TEM class alone was the next frequent class of ESBLs (11.3%), but a higher percentage of harboring TEM also had CTX genes along with it (12.9%). Isolates were not found to harbor the SHV genes alone, whereas 12.9% had CTX- M also.
Many different studies show that the distribution of ESBL genes is diverse. Some studies have shown a predominance of the TEM gene with less SHV and CTX-M [21]. In one study in Saudi Arabia, characterization of the resistance genes revealed the dominant presence of SHV, followed by TEM and CTX-M [22]. Other findings agree with ours, which indicated a predominance of CTX-M [9, 23].
Earlier reports mentioned that the most prevalent type of ESBL genes is SHV, TEM, and CTX-M. During the past decade, TEM and SHV types were reported to be the most common types of b-lactamase genes, but recently, the CTX-M type has spread worldwide compared to TEM and SHV genotypes [24].
CTX-M is predominant in many regions; several reports were made from Iran (74%) [25], Morocco, North Africa (70%) [26] and India (93.7%) [27]. However, TEM-type b-lactamase gene was predominant in Italy (45.4%) [28], Portugal (40.9%) [29], and Turkey (72.7%) [30].
The most important finding in our study was the co-existence of different ESBLs. A number of the isolates had multiple genes, namely TEM and CTX-M; TEM, SHV, and CTX; and SHV and CTX-M in their genome suggesting the carriage of multiple plasmids. The co-existence of all three bla genes was observed in 32.5% of the ESBL isolates. Several studies reported the co-existence of various ESBL genes within the same isolates [5, 24, 31]. The co-existence of all the three bla genes observed in our study was similar to that previously described in Iran [21].
CONCLUSION
In the present study, ESBL-producing Enterobacterale displayed an increasing rate of resistance for aminopenicillins (ampicillin), followed by piperacillin-tazobactam, cephalosporin groups (ceftriaxone, ceftazidime, cefepime, ciprofloxacin), and fluoroquinolone (ciprofloxacin).
The findings in this study document the threat of ESBL among Enterobacterale especially the CTX-M class in our geographical region with the occurrence of these strains as etiological agents of infection in the hospital setting. The frequency of antibiotic usage and even the dosages and period of administration vary greatly from one country to another. This has led to large differentials in the emergence of resistant patterns. Therefore, it is essential to study and report trends in antimicrobial resistance on a regular basis. Our findings represent a preliminary study for follow-up work, establishing novel features concerning the potential of multiple plasmids.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The ethical committee of Prince Sultan Military College of Health Science approved this study.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The authors confirm that the data supporting the findings of this research are available within the article.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Thanks are due to the microbiology department of the King Fahad Military Medical Complex for providing the specimens. We thank Mr. Louis Greenwood for proofreading the manuscript.


