All published articles of this journal are available on ScienceDirect.
Preparation of Specific Polyclonal Antibody Against the Recombinant Mutacin Produced by sfGFP Fusion Protein Technology
Abstract
Mutacin I, a bacteriocin produced by streptococcus mutans, displays an antimicrobial activity against many gram positive and some gram negative bacteria. Because of its medical importance, production of this short peptide in large scale for future applications is a significant challenge. This work described the improvement of a novel system to produce the recombinant mutacin using fusion protein technology.
The short peptide was expressed directly as a fusion protein with a superfolder form of the green florescent protein (sfGFP), resulting in a high yield expression of soluble sfGFP-mutacin fusion protein (30 kDa) in the cytoplasm of E. coli. Mutacin was released from the fusion by enzymatic cleavage at the tobacco etch virus (TEV) protease recognition site and separated from the carrier sfGFP by nickel affinity and gel filtration chromatography. An additional advantage of this fusion system was tested in the generation of mutacin-specific polyclonal antibodies. Specific anti-mutacin IgGs were affinity purified, and were able to recognize the mutacin-sfGFP fusion protein or the cleaved forms of mutacin.
Even though it was efficiently produced (25 mg/L) by this method, pure mutacin was devoid of antibiotic activity. Fourier transform infrared spectroscopy (FTIR) analysis revealed the absence of thioether bonds in the purified mutacin, which are critical for final structure and function of this antibiotic. Determining whether the activity of pure mutacin could be recovered by the reformation of such structures by chemical reaction needs more investigations. The development of this system will provide large quantities of mutacin for future studies and applications as broad spectrum antibacterial peptide.
1. INTRODUCTION
Bacteriocins are proteinaceous toxins produced by streptococcus mutans, which is an indigenous bacterial inhabitant of the human oral cavity [1]. The diversity of bacteriocin from S. mutans has been recently highlighted, as they represent 40% of the listed streptococcal antimicrobial peptides [2]. Historically, the importance of mutacins was based on the role they play in the colonization of the oral cavity and their usefulness as possible anti-caries agents [3]. New applications are developing for mutacins as potential food preservatives [4] and as new effective antibiotics [5]. Mutacin I is produced by S. mutans CH43 and UA140 [6, 7]. The mature peptide is composed of 24 amino acids with a molecular mass of 2364(Da. Mutacin I belongs to the lantibiotic type IA and epidermin group. The biosynthetic operon which is located in the bacterial chromosome was reported to encompass 14 ORFs [7]. Mutacin I could be used in different conditions due to its pH, thermostability and hydrophobic nature, which can be advantageous for coating the surface of catheters and preventing adhesion of bacteria to the surface [8].
Many lantibiotics have not been extensively tested for their potential usefulness in treating infectious diseases even though they are known to be both potent and have a broad spectrum of activity, notably against gram positive species [9]. The principle reason for this is the difficulty in obtaining these molecules in high purity and in sufficient, cost effective amounts to enable their testing and commerciali-zation. For production of these peptides to be valuable in industry, they must be produced in an environmentally safe and cost effective manner [10]. Recombinant DNA technology provides means for an economical protein production, thus many antimicrobial peptides have been successfully produced through recombinant expression in various heterologous hosts [10]. E. coli and yeast are the two major systems used to produce recombinant antimicrobial peptides, which together account for over 95% of all reported cases [10, 11]. Antimicrobial peptides produced in E. coli are often expressed as fusion with carrier proteins, a strategy necessary to mask these peptides lethal effect towards the host and protect them from proteolytic degradation [12].
Recently, green fluorescent protein (GFP) from Aequorea jellyfish has become a common label for many in vivo and in vitro applications due to GFP’s ability to fold and form a visually fluorescent chromophore through autocatalytic cyclization and dehydration/oxidation reactions [13]. GFP is exceptionally stable in solution, even at high temperature, in the presence of organic solvents or alkaline pH condition. [14]. GFP has been used as a reporter in protein folding [15, 16], protein-protein interactions [17, 18] and gene translation [19]. GFP is a 238 amino acids polypeptide with a molecular weight of 27(kDa. The cDNA encoding GFP was cloned and sequenced in 1992 [20]. Lately, Waldo and coworkers reported the engineering of a superfolder GFP (sfGFP) that showed improved tolerance to circular permutation, increased resistance to denaturation, improved folding kinetics, and increased resistance to aggregation during refolding [21, 22]. Furthermore, sfGFP fusions are more soluble than conventional GFP fusions [23]. sfGFP has proven to be very useful as a scaffold for improved protein detection and tagging both in vivo and in vitro using self-assembled sfGFP fragments [18, 24]. In addition, a protein of interest can be studied in the form of GFP fusion protein, since GFP folding does not affected by the folding status or solubility of its fusion partner in E. coli.
The fusion protein could be easily recovered, purified and cleaved with an appropriate protease to release the target protein in its active form. To achieve this end, specific sequences are included between the tag(s) and native protein and then cleaved with specific reagents. Although both chemical and enzymatic reagents have been used to remove tags from recombinant proteins [25], only proteases exhibit enough specificity to be generally useful for this purpose. Common proteases include enterokinase, thrombin [26], factor Xa [26] and SUMO protease [27]. Recently it has become clear that certain viral proteases have much greater stringency, making them particularly useful for this application [28]. Tobacco etch virus (TEV) is a cysteine protease, which is active on a variety of substrates and cleaves efficiently at low temperatures [29, 30]. It recognizes the amino acid sequence ENLYFQ(G with high efficiency and cleaves between Q and G [31].
The development of efficient detection and purification procedures for bacteriocins could greatly facilitate their use as food preservatives or alternative antibiotics [32]. Antibodies are an important tool used by many investigators in the research and have led to many advances. In biochemical and biological researches, Antibodies are routinely used as ligands for the preparation of immunoaffinity columns [33] and as coating or labeling reagents for the detection of bioactive molecules in a variety of assays, such as the enzyme linked immunosorbent assay (ELISA) and immunoblotting [34]. The generation of antibodies against bacteriocins may provide sensitive and specific methods for the identification and detection of mutacin I producing strains and for the quantification of bacteriocins cleaving in different substrates by the use of immunochemical assays [35].
We present here a bacterial expression system that uses sfGFP as a fusion partner with the short peptide that was released by enzymatic cleavage using the TEV protease recognition site and purified by means of the internal 6×His tag using nickel affinity chromatography. An additional advantage of this fusion system was in the generation of mutacin−specific polyclonal antibodies. Antibodies were produced after rabbit immunization by the pure fusion protein in order to increase the immune response against the displayed short peptide.
2. MATERIALS AND METHODS
2.1.. Bacterial Strains, Growth Conditions and Plasmid
E. coli strains TOP10 (Invitrogen) and BL21 (DE3) Rosetta (Novagen) were used in cloning and protein expression after transformation by electroporation with the plasmid constructs pT7-His-sfGFP-mutA and pT7-His (kindly provided by Prof. Yu Ding, Fudan University, China). For general maintenance and protein expression, E. coli were grown in Luria Broth (LB; 1% Tryptone, 0.5% yeast extract, 171mM NaCl) (Bio Basic INC) with the required antibiotic at 100mg/ml (Ampicillin; Applichem) at 37°C.
2.2.. Construction of the Plasmid pT7−His−sfGFP−mutA
The coding sequence of sfGFP in the plasmid pT7−sfGFP(mutA [36] was replaced by the DNA fragment His−sfGFP from the plasmid pRSET−sfGFP [37]. This was achieved by subcloning using a double digestion with NdeI/BamHI (Fermentas) of both plasmids, and the long linearized DNA fragment from the first plasmid was ligated with the small insert from the second using T4 DNA ligase (Fermentas). Freshly prepared electro−competent E. coli TOP10 cells were transformed with the new plasmid construct pT7-His-sfGFP-mutA by electroporation. Colony PCR screening for positive His-sfGFP clones was performed using specific primers (T7F/T7R), and the successful cloning was confirmed by digestion with restriction enzymes and by sequencing.
2.3.. Expression and Purification of sfGFP−mutacin Fusion Protein
The confirmed plasmid construct was used to transform by electroporation E. coli BL21 (DE3) Rosetta cells. Protein expression of sfGFP-mutacin was performed in 250ml shake flasks by growing the bacteria in LB medium untill an optical density of 0.5 to 0.7 was reached, and then expression was induced with 0.5mM Isopropyl β−D−thiogaldctoside (IPTG; Promega) for 16h at 19°C. After pelleting the cells, the pellet was resuspended in PBS 1×, then lysed by sonication (Lab Sonic) on ice and the lysate was cleared by 20min centrifugation at 8,000(rpm and 4°C. Using fast protein liquid chromatography (FPLC) with AKTAprime plus system (GE Healthcare), recombinant sfGFP-mutacin was purified from this cytoplasmic extract using a 5ml column of nickel charged Nitrilotriacetic acid (NTA) superflow Sepharose (Qiagen). After washing, the bound proteins were eluted from the column with a 500mM imidazole buffer. The eluted fraction was concentrated on Vivaspin concentrators with a molecular mass cutoff of 10(kDa (Vivascience). Protein samples were separated by SDS-PAGE using a Bio-Rad mini-Protean II system following the manufacturer’s instructions. Gels were prepared using stacking gel 5% and running gel 12%. After electrophoresis, the gel was stained in coomassie blue for 2h then distained.
2.4.. Cleavage of sfGFP-mutacin Fusion Protein by TEV Protease
Recombinant sfGFP−TEV protease was expressed from pT7−His plasmid and purified by nickel affinity chromatography using the standard method [23]. The purified protease, at a protein concentration of ~1(µg/µl in 50% glycerol, was used in the cleaving reaction of the substrate sfGFP-mutacin (1 mg), which contained a TEV cleavage site at the N−terminal of mutacin. Protease digestion was performed in a mass ratio of substrate to enzyme 100:4 (w:w) in a reaction buffer (75mM NaCl, 0.5mM EDTA, 25mM Tris pH8, 10% glycerol and 100mM DTT) at 30°C for 16h. The extent of cleavage of the samples was determined or compared by SDS−PAGE analysis. The cleavage product was purified by nickel affinity chromatography to eliminate His tagged proteins (sfGFP and TEV protease).
2.5.. Purification of Short Peptide by Gel Filtration
The short peptide resulted from TEV protease digestion was purified using Superdex 200 10/300 GL column (GE Healthcare). After washing, the bound proteins were eluted from the column with sodium phosphate buffer 1× (30.5mM Na2HPO4, 19.5mM NaH2PO4, 0.15M NaCl, pH(7). The purity of the short peptide was evaluated in SDS−PAGE stained in cold silver staining buffer (0.2% silver nitrate AgNO3 and 0.03% formaldehyde) for 20min, then destaining was done with the stop solution (50% methanol, 12% glacial acetic acid).
2.6.. Production of Polyclonal Antibody Against the Recombinant sfGFP−mutacin Protein
For the first immunization, purified sfGFP−mutacin protein (500µg) in PBS (1ml) was mixed with an equal volume of Freund’s complete adjuvant (Bio Basic Inc.) to form a stable emulsion. One white rabbit was injected subcutaneously at 2 to 4 different sites. Three booster injections were given with 250µg recombinant protein mixed with incomplete Freund’s adjuvant at 15 day intervals. To evaluate the immune response, blood samples (0.5−1ml) were collected before each injection and bleeding (30ml) was done 10 days after the last boost. Polyclonal antibodies (IgGs) were purified from rabbit serum by affinity chromatography on a 5ml HiTrap Protein A column (GE Healthcare) according to the manufacturer's instructions. Binding was performed in 0.02M sodium phosphate, pH7.0, and elution was in 0.1M citric acid, pH3.0. Eluted IgGs were collected and immediately neutralized to physiological pH by adding 1M Tris−HCl buffer, pH9 and then concentrated to 1mg/ml using 30kDa−molecular mass cutoff concentrators.
2.7.. Affinity Purification of Anti−mutacin Specific IgG
Two N−hydroxysuccinimide (NHS) activated Sepharose Columns (GE Healthcare) were used for affinity purification of mutacin−specific IgGs from the rabbit total antibodies. Concentrated sfGFP and sfGFP-mutacin (20mg) in coupling buffer (0.2M NaHCO3, 0.5M NaCl, pH(8.3) were applied onto two NHS−activated columns, respectively, and allowed to bind for 30min at 20°C. Any excess of active NHS groups that have not been coupled were deactivated by sequential washing with (0.5(M ethanolamine, 0.5(M NaCl, pH(8.3) and (0.1M sodium acetate, 0.5M NaCl, pH(4). Protein A purified IgGs from rabbit serum were first passed through the sfGFP−column. Purified sfGFP(specific IgG sample was concentrated and stored at -20°C. Unbound IgG, which was empty of sfGFP(specific IgGs, was washed away with the washing buffer (0.05(M sodium phosphate, 0.15(M NaCl, pH(7), and then it was re-purified by passing it through the sfGFP-mutacin column in order to obtain pure mutacin(specific IgGs which were eluted with 3–5 column volumes of 0.1M citric acid buffer and then neutralized with 1M Tris−HCl buffer, pH(9.0. Purified mutacin−specific IgG sample was buffer exchanged with 0.05M sodium phosphate buffer pH7, and then concentrated down to 1mg/ml.
2.8.. Mutacin Detection by ELISA Assay
An indirect ELISA format was employed for the analysis of sera and purified IgG. The basic protocol was as follows: sfGFP, sfGFP-mutacin, and mutacin antigens were diluted in carbonate coating buffer and 100μl were added to the wells of an ELISA microplate at 4°C overnight. After washing (phosphate-buffered saline containing 0.05% Tween 20) the plate was washed 3 times to remove unbound antigen, 150μl of blocking buffer (1% bovine serum albumin (BSA), 3% skimmed milk in 1 PBS) were applied to all the wells for one hour at 37°C. After the removal of blocking buffer, 100μl of rabbit sera or IgGs diluted in 1% blocking buffer were added per well for 1h at RT. After 3 washes, goat anti rabbit IgG conjugated to horse-radish peroxidase (HRP) was diluted (1:2000) in 1% blocking buffer, added (100μl per well), and allowed to bind to captured rabbit IgG for a further hour at RT. After an additional 5 washes, bound conjugate was detected with 50μl of 3,3′,5,5′-Tetramethylbenzidine (TMB, Sigma) substrate, the reaction was stopped after 8min with the addition of 50(µl 1M H2SO4. The spectroscopic absorbance of each well was measured in an automated plate reader at a wavelength of 450nm.
2.9.. Biological Activity of Recombinant Mutacin
Antibiotic potency of recombinant mutacin was tested on three different strains; Streptococcus thermophilus, Bacillus thuringiensis, Lactobacillus bulgaricus, by turbidimetric assay. Bacterial strains were cultured in LB medium until OD600 reached 1, each strain was diluted to 0.1 in LB medium and cultured in 300(μl/well in a 48(well microplate. Thirty µl of antibiotic stocks (1mg/ml), mutacin or ampicillin, were added to the wells of each type of bacteria to obtain a final concentration of 100(μg/ml. the culture microplate was incubated at 37°C for 3h with gentle shaking. The spectroscopic absorbance of each well was measured in an automated plate reader at a wavelength of 600nm.
2.10.. FTIR Analysis
The IR spectrum of purified mutacin was determined using FTIR spectrometer (Thermo–Nicolet 6700) instrument at the Department of Chemistry in the AECS. Lyophilized sample (0.5mg) was mixed with 200mg of KBr in a clean glass pestle and mortar. The contents were dispersed in a KBr disk and then pressed into 1mm pellets. The KBr disk was placed in a special sample holder in the IR beam of the spectrometer. Sample was analyzed in the wavenumber range of 400 and 4000cm-1 with a resolution 4cm-1 using 64scans.
3. RESULTS
3.1.. Construction of the Plasmid pT7−His−sfGFP−mutacin
In our previous work, the coding sequence of the mature mutacin (mutA) was constructed by Splicing by Overlap Extension (SOE) PCR using long overlapping primers, which cover the whole gene, and the DNA adapter was cloned in the plasmid pT7−His [36]. Mutacin was produced as a part of a high solubility and productivity fusion protein (30 kDa), which was detectable by the colorimetric properties of sfGFP (Fig. 1A). However, the expressed fusion protein was mostly lost in the flow−through during nickel chromatography purification, resulting in a low yield purification. We surmised that the purification problem could be related to the inaccessibility of the internal 7×His tag, that was added between the coding sequences of sfGFP and mutacin. To overcome the purification problem in the previous study, an extra 6×His tag was added to the N−terminal of sfGFP-mutacin fusion protein, since it is impossible to eliminate this tag from mutacin if it was added to the C−terminal. To achieve this goal, the sfGFP gene in pT7−sfGFP−mutA plasmid construct was replaced with the His(sfGFP adapter from pRSET(sfGFP plasmid [37], by NdeI/BamHI double digestion and sub(cloning (Fig. 1A).
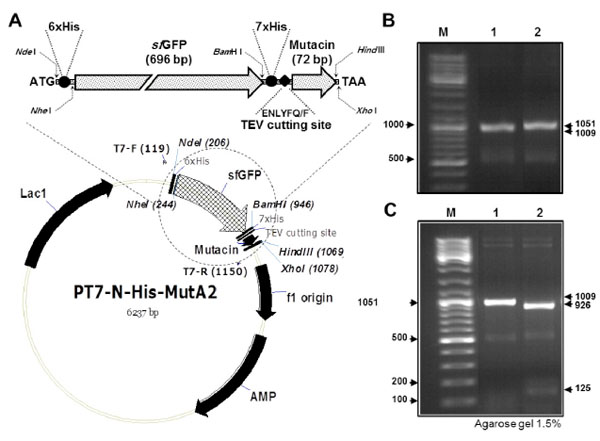
Construction of the plasmid pT7−His−sfGFP−mutA
(A) Schematic illustration of the plasmid pT7−His−sfGFP−mutA, with an enlarged part (up), which represents the coding region for the fusion protein sfGFP−mutacin. The most important elements of the plasmid are indicated. These include T7 promoter, His tags, TEV cutting site and the restriction sites used for cloning. Locations of T7 primers (F and R) used for PCR amplification and for sequencing are shown on the plasmid. (B) Agarose gel electrophoresis of PCR products using T7F/T7R primers and the plasmids pT7−His−mutA (lane 1) or pT7−His−sfGFP−mutA (lane 2) as DNA template. (C) DNA amplicons from the previous PCR were digested with NheI and separated by gel electrophoresis. DNA molecular weight marker was loaded in the first lane (M).
The new plasmid construct pT7(His(sfGFP(mutA was transformed into E. coli TOP10, then confirmed by colony PCR screening using T7F/T7R primers (Fig. 1B). PCR amplification of pT7(His(mutA containing colony (lane(1) resulted in a band of 1009(bp while in case of the new plasmid construct pT7(His(sfGFP(mutA (lane(2) a bigger band of 1051(bp was amplified from the colony. This was because of the presence of an additional N(terminal 6(His tag in the new plasmid construct. To further confirm the difference between the two plasmid constructs, PCR products, from previous reaction, were digested with NheI (Fig. 1C). Only amplified DNA fragment from the new construct contains a recognition site for NheI, resulting in the cleavage of this fragment into large and small bands of 926(bp and 125(bp, respectively (lane(2). To express the 6(His tagged sfGFP-mutacin, the new plasmid construct was transformed into the strain BL21 (DE3) and protein expression and purification was conducted by FPLC (Fig. 2). Obviously, the addition of N−terminal 6(His tag did not affect the solubility or the high level expression of the fusion protein, but resulted in its optimal purification (lane 5), as no further traces were remained in the flow(through sample (lane 4).
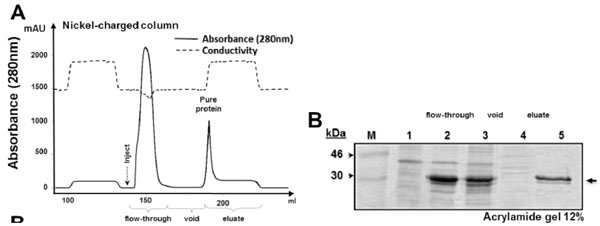
Expression and purification of His−sfGFP−mutacin fusion protein
(A) Diagram of purification procedure using nickel−charged column (5 ml). Continuous line represented the absorbance of the eluate, different purification steps are shown below and peaks of the flow−through sample and of purified sfGFP−mutacin are indicated. Dashed line represented conductivity of the eluate. (B) SDS−PAGE (acrylamide 12 %) of protein samples from the different steps of expression and purification; total cytoplasmic extract before IPTG induction (lane 1), after 16 h of induction (lane 2), after sonication (lane 3), flow−through sample from nickel−charged column (lane 4) and purified sfGFP−mutacin fusion protein (lane 5). Protein molecular weight marker (in kDa) was loaded in the first lane (M), and side arrow referred to the expected location of the recombinant protein in the gel.
3.2.. Purification of Mutacin After the Cleavage of sfGFP−mutacin Fusion Protein
Pure sfGFP-mutacin as well as sfGFP-TEV protease were prepared and their concentrations were determined by the Bradford method and adjusted to 1mg/ml. The mass ratio of the substrate His-sfGFP-mutacin; which contains the protease recognition site, to the sfGFP-TEV protease enzyme was 100:4, so that 25mg of the substrate were incubated with 1mg of the enzyme in the appropriate buffer for 16h at 30°C (Fig. 3). Protein samples from the cleavage reaction were prepared and then loaded onto 15% SDS(PAGE gel for electrophoresis, which was finally visualized by staining with silver nitrate (Fig. 3A). In this result, liberated mutacin (~2kDa) could be observed after the cleavage reaction (Lane 1), then nickel affinity purification was applied to eliminate his tagged moieties from the reaction mixture (sfGFP-TEV and His-(sfGFP) (Lane 2) and relatively pure mutacin was finally recovered in the flow(through fraction (Lane 3).
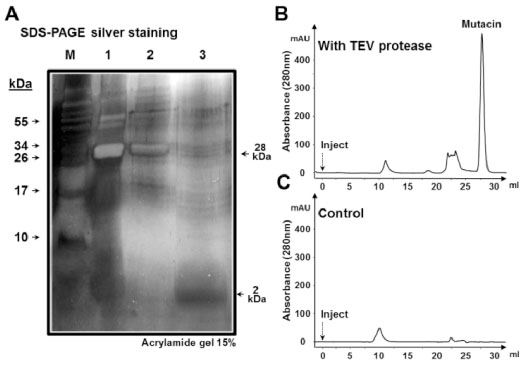
Cleavage, detection and purification of mutacin
(A) Silver staining of SDS−PAGE (15 %) of protein samples from the cleavage mixture before nickel affinity purification (lane 1), pure fraction containing His tagged proteins; His−sfGFP and sfGFP−TEV (lane 2) and the flow−through sample containing pure mutacin (lane 3). Gel filtration of injected nickel−purified flow−through sample from the cleavage reaction of sfGFP−mutacin in the presence of active (B) or of heat−inactivated form (C) of sfGFP−TEV. Gel filtration was achieved by injecting protein sample (Inject) into a Superdex200 column (10/30) at a flow rate of 1ml/min, and the fraction peak of pure mutacin is shown.
For more confirmation of the cleavage and to get highly pure mutacin, protein samples, from the flow−through fraction, before and after fusion protein cleavage and nickel affinity purification were concentrated and applied to a gel filtration column (Fig. 3B). The protein fractions eluted are detected by an in−line UV monitor and collected for subsequent specific analysis. When the flow−through sample from nickel affinity purification of a TEV digested sfGFP-mutacin were added, pure mutacin was eluted from the gel filtration column as a last single peak (Fig. 3B). This peak was clearly absent in the control condition where samples were incubated with heat−inactivated TEV protease (Fig. 3C).
3.3.. Production of Anti-mutacin Polyclonal Antibody
Polyclonal antibodies are produced in vivo in response to immunization of an animal, such as a rabbit, with different antigens. For small molecules, a carrier protein is required in order to be recognized by the immune system. In the fusion model of sfGFP-mutacin, the short peptide is already attached to a carrier immunogenic protein thus having the potential to elicit a good immune response [37]. For this aim, an adult female rabbit was immunized with four subcutaneous injections of purified sfGFP-mutacin, by two−weeks interval. Blood samples were collected at several time points from the start of immunization and tested in ELISA assay against sfGFP and mutacin to evaluate the rise of a specific immune response (Fig. 4A).
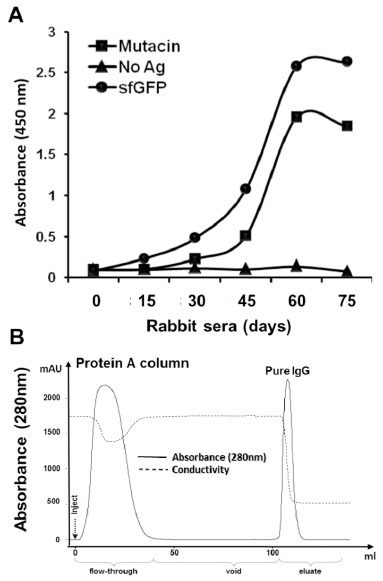
Evaluation of immune response and purification of anti−sfGFP−mutacin IgG.
(A) Reactivity of blood sera from sfGFP−mutacin immunized rabbit was ELISA tested against, sfGFP (0.5 μg/well) and mutacin (2 μg/well) or in the absence of antigens (No Ag). Serum samples (diluted to 1:5000) were taken from the immunized rabbit at several time points (days) from the first injection onwards. Antibody/ antigen interaction was detected by goat anti−rabbit HRP conjugated antibody (1:4000). (B) Protein−A affinity purification of IgG from rabbit serum. Five ml of diluted rabbit serum (at day 60) were injected (inject) onto HiTrap Protein−A column and washed with phosphate buffer (void) to remove the unbound proteins (flow−through), before eluting pure IgG (eluate).
A solid phase ELISA test of the rabbit sera (dilution 1:5000) prepared from these samples revealed an increase in reactivity toward sfGFP and mutacin after six weeks of immunization (Fig. 4A). From this serum sample (at day 60), rabbit IgGs were purified by affinity chromatography using a protein A column. An automated purification procedure was established using the AKTA prime system allowing a direct and confirmed purification of the total IgGs from 5 ml of serum (Fig. 4B).
3.4.. Preparation and Testing of Pure Anti-mutacin IgG
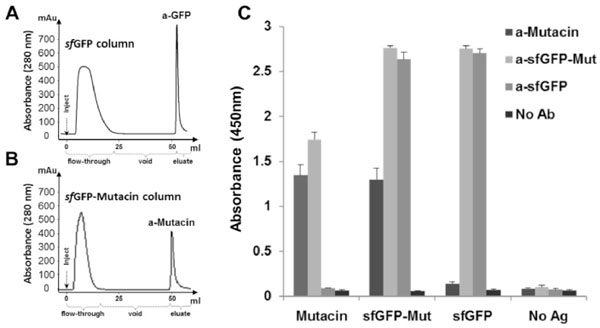
To extract anti−mutacin specific antibodies from total purified IgGs of the immunized rabbit, sfGFP and sfGFP-mutacin were covalently−immobilized on two NHS−activated columns. To establish these columns, pure recombinant sfGFP and sfGFP−mutacin were coupled by means of the NH2 group to the active NHS group of the resin. Firstly, total IgGs were purified by sfGFP(column. After column washing and calibration, total serum IgGs, from the previous step of protein A purification were loaded onto the column. Unbound IgGs, which contain the rest of antibodies including anti−mutacin, were recovered in the flow(through. After column washing, specific anti(sfGFP antibodies were eluted from the column by low pH buffer as a pure fraction (Fig. 5A). Secondly, unbound IgGs in the flow(through of the first step, were filtered using an sfGFP-mutacin(column to obtain specific anti−mutacin antibodies in the pure fraction (Fig. 5B).
Immunoaffinity chromatography of specific anti−mutacin antibody.
(A) Affinity 1 ml sfGFP−column was used in chromatography to purify specific antibodies from total IgGs which were purified from the immunized rabbit by protein−A column. Total antibodies sample (5 ml) was loaded in the column (inject) and unbound fraction (flow−through) was recovered. After washing, pure anti−sfGFP IgGs were recovered from the column by elution (100mM citric acid, pH3). (B) Affinity 1 ml sfGFP−mutacin−column was used in chromatography to repurify the previous unbound fraction of IgGs to get anti−mutacin antibodies in the pure fraction. (C) ELISA test in the absence of antibodies (No Ab) or in the presence of anti−mutacin, anti−sfGFP−mutacin and anti−sfGFP (purified IgGs). Antibodies (1 mg/ml stock) were tested at dilution (1:5000), whereas, anti−mutacin was tested at dilution (1:1000) in the presence of immobilized sfGFP and sfGFP−mutacin (0.5 μg/well), mutacin (2 μg/well), or in the absence of antigens (No Ag).
To evaluate the activity of purified antibodies against their antigens, indirect ELISA was performed on sfGFP, sfGFP-mutacin and mutacin pre−coated wells and bound rabbit IgGs were detected with a goat−anti−rabbit secondary antibody conjugated to HRP (Fig. 5C). As expected, pure total IgGs and anti(sfGFP were able to efficiently detect sfGFP and sfGFP-mutacin at high dilution (1:5000). Interestingly, anti-mutacin as well as total IgGs, but not anti−sfGFP, were able to detect mutacin in its fusion or singular forms (Fig. 5C).
3.5.. Mutacin Characterization
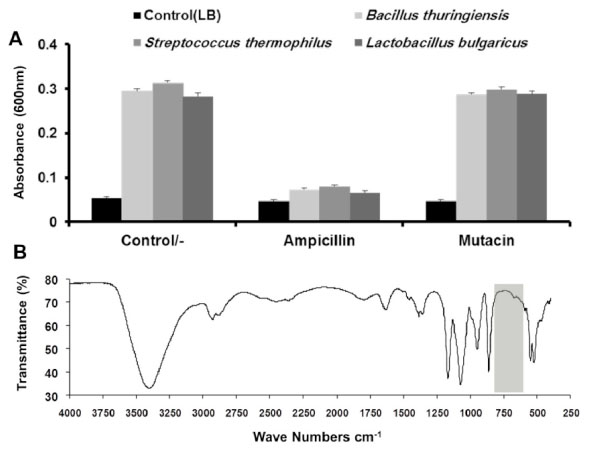
After mutacin was cleaved from the fusion protein and purified by gel filtration, its biological activity was tested by bacterial growth turbidimetry, using three different bacterial strains S. thermophilus, Bacillus thuringiensis, Lactobacillus bulgaricus. Turbidimetry by definition is the process of measuring the loss of intensity of transmitted light due to scattering effect particles suspended in the medium. Growth inhibitory effect of mutacin was tested in reference to ampicillin, and the LB medium was used as a negative control. All tested microorganisms appeared to be insensitive to mutacin, whereas the ampicillin was able to inhibit completely the growth of all used bacteria (Fig. 6A).
Characterization of purified mutacin
(A) Chart of turbidimetric assay for the determination of the growth inhibitory effect of mutacin (100 ìg/ml) and ampicillin (100 ìg/ml) in LB medium for S. thermophilus, B. thuringiensis, and L. bulgaricus. (B) FTIR spectrum using Thermo – Nicolet 6700 system for mutacin lyophilized sample dispersed on KBr disk. Grey region (600−800cm-1) referred to the position where the expected peaks of R−S−R bonds should appear.
We supposed that the ineffectiveness of mutacin was due to the absence of one or more of its important bonds that affect its biological activity. As we mentioned previously, mutacin belongs to the lantibiotic type IA, which contains 4 thioether bridges (R(S(R) that are formed in post(translation modification. These additional structures are often critical for the biological activity of lantibiotics [38]. To test this idea, FTIR analysis; which is based on the analysis of functional groups and identification of compounds, was applied on lyophilized pure mutacin. The hallmark finding obtained from this analysis was that no significant peaks were observed around 600 and 800 cm-1 which may indicate the absence of thioether bonds [39]. Without such formations, recombinant mutacin structure could be malformed and thus devoid of activity [40] (Fig. 6B).
4. DISCUSSION
Antimicrobial peptides are now being looked at as an interesting alterative to combat antibiotic resistant strains of bacteria. Because of their medical importance, production of these polypeptides on a large scale for commercial development is a significant challenge. Different peptides have been successfully produced using several methods, including expression as fusion proteins in bacterial systems [41]. Mutacin consists of 24 amino acid residues and exhibits a wide−spectrum antibacterial effect. A major obstacle for mutacin characterization and application is the difficulty of its purification from the original sources [7] and the high cost of its chemical manufacturing [42].
Fusion protein technology represents one of the best solutions to achieve rapid, efficient, and cost−effective protein expression and purification of recombinant proteins [43]. Several biological expression systems have been developed by fusing antimicrobial peptides to a partner protein to mask the intrinsic antimicrobial activity of the peptide inside variety of host cells such as bacteria [44], yeast [45] and insect cells [46, 47]. Many reports described the expression of different short peptides using variable strategies, including the application of GFP fusions in the expression of hard−to−fold proteins [48], toxins [49], proteases [50] and medical short peptides [36, 51] in both prokaryotic and eukaryotic cell types. Visibility of GFP is the most distinguishable element from other fusion tags [52]. Its unique green color (either visible blue light or UV light) allows us to monitor the existence of GFP fusion protein through purification steps even without any measurement apparatus like a UV spectrophotometer [53].
Cleavage of the fusion complex usually is necessary because of the possible interference with the structural or functional properties of the recombinant target protein. TEV protease represents the best choice for our system because of its high specificity and activity towards its cutting site [50]. The P1' position of TEV protease cleavage site (ENLYFQ(G) is relatively relaxed and many different residues can be substituted for glycine with little or no impact on the efficiency of cleavage [31]. Because we did not want to make any modification to mutacin structure, and as TEV protease cleavage leaves an additional one amino acid (glycine; G) to the C−terminal moiety of its cutting site, the last amino acid residue (G) of TEV cutting site was replaced by (phenylalanine, F), the first amino acid of mutacin.
Many previous works have described a method to produce TEV as soluble recombinant 6×His−tagged protein in E. coli [23, 54, 55]. In our model, purified sfGFP−fused protein could be cleaved with TEV protease, and the desired native protein could be recovered after eliminating TEV protease by nickel affinity chromatography together with the 6(His−tagged sfGFP portion. Gel filtration chromatography was finally conducted for cleaved mutacin in order to eliminate impurities and to obtain a highly purified short peptide.
One important advantage for fusion protein technology is for enhancing antibody production [56]. Antibodies are powerful tools for protein detection and purification by affinity chromatography. Such techniques have not found an application in the bacteriocin field, mainly because it is difficult to develop antibodies against such antimicrobial peptides. The generation of antibodies against bacteriocins is a main research target because it permits their detection and quantification in samples, as well as their purification in a single step through similar strategies [57]. Small proteins in general are bad immunogens, which fail to solicit a significant immune response alone and should be conjugated to a large carrier molecule for efficient presentation to the immune system of a vaccinated animal. In fusion structures, sfGFP could play the role of a carrier for these small molecules in order to enhance specific antibody production. In our experiment, rabbit immune response against mutacin was induced by its fusion form with sfGFP, which has an important immunogenicity response in rabbits [37]. Rabbit immunization with sfGFP−mutacin resulted in a high titer immune response (~1:50000) towards sfGFP. To produce specific IgG FPLC fractions from immunized rabbit, recombinant sfGFP and sfGFP-mutacin were covalently immobilized on NHS(activated sepharose column by means of the N−terminus using free amine conjugation chemistry. The affinity chromatography column, which is created by this method, was successfully used to purify specific anti(sfGFP and anti−mutacin antibodies from total rabbit IgGs. FPLC purified polyclonal anti(sfGFP and anti−mutacin IgGs represent a valuable tool to detect GFP and mutacin in many biotechnological techniques.
An important production yield of pure mutacin was obtained using this system, estimated to be about 25mg/L of bacterial culture, and the manufacturing cost was considerably lower, since all necessary elements, such as TEV protease and the specific antibodies, were prepared "in house". However, pure mutacin was inactive and failed to inhibit bacterial growth of three different gram−positive strains of bacteria. It was suggested that losing activity could be related to the malformation of mutacin during protein expression, cleavage or even purification. Furthermore, protein synthesis machinery, used in E. coli BL21 (DE3) strain, was not suitable for the production of properly structured mutacin. In S. mutans and after mutacin is ribosomally synthesized, the resulting translated protein must be modified before becoming active [7]. Genes coding for the enzymes that facilitate these post(translational modifications are usually in close proximity to the structural gene of mutacin [58]. A major mutacin structural modification is the formation of lanthionine bonds, which are thioether−based (R(S(R) ring structures critical for the biological activity of lantibiotics [59]. As expected, FTIR spectrum of pure mutacin revealed an obvious absence of thioether peaks. It would be of great interest to conceive a solution to correct the structure of mutacin either by chemical reaction or by in vitro enzymatic modification using a recombinantly produced lanthipeptide synthetase [60].
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
LIST OF ABBREVIATIONS
ACKNOWLEDGEMENTS
The authors would like to thank the Director General of the Atomic Energy Commission of Syria and the head of the Molecular Biology and Biotechnology department for their continuous support throughout this work.


