All published articles of this journal are available on ScienceDirect.
Molecular Identification of Mycobacterium Species of Public Health Importance in Cattle in Zimbabwe by 16S rRNA Gene Sequencing
Abstract
Mycobacterium species are naturally found in the environment as well as in domestic animals such as cattle. So far, more than 150 species of Mycobacterium, some of which are pathogenic, have been identified. Laboratory isolation, detection and identification of Mycobacterium species are therefore critical if human and animal infections are to be controlled. The objective of this study was to identify Mycobacterium species isolated in cattle in Zimbabwe using 16S ribosomal RNA gene amplification and sequencing. A total of 134 cow dung samples were collected throughout Zimbabwe and mycobacteria were isolated by culture. Only 49 culture isolates that were found to be acid-fast bacilli positive by Ziehl-Neelsen staining. The 16S rRNA gene was successfully amplified by PCR in 41 (84%) of the samples. There was no amplification in 8 (16%) of the samples. Out of the 41 samples that showed amplification, 26 (63%) had strong PCR bands and were selected for DNA sequencing. Analysis of the DNA sequences showed that 7 (27%) belonged to Mycobacterium neoaurum, 6 (23%) belonged to Mycobacterium fortuitum, 3 (12%) to Mycobacterium goodii, 2 (1%) to Mycobacterium arupense, 2 (1%) to Mycobacterium peregrinum or M. septicum and 1 isolate (0.04%) to Mycobacterium elephantis. There were 5 (19%) isolates that were non-mycobacteria and identified as Gordonia terrae, a close relative of Mycobacterium. The study therefore provided a molecular basis for detection and identification of Mycobacterium species in animals and humans.
INTRODUCTION
The Genus Mycobacterium belongs to the Family Mycobacteriaceae that is found within the Order Actino-mycetales [1]. Members of this genus are very diverse and more than 150 species have so far been identified [2]. They are found in various environments such as natural and municipal water, soil, air and sewage [3,4]. Humans are exposed to environmental Mycobacterium in water through drinking, and other activities such as swimming and bathing [4]. Some Mycobacterium species are also found in the gut of several animals such as cattle and wild animals and these can be transmitted to humans [5,6]. Humans are exposed to Mycobacterium from domestic animals through contami-nated foodstuffs or by direct contact with the animals and their products [7,8]. Several species of environmental Myco-bacterium are now known to be of public health importance since they are pathogenic to humans especially those with underlying medical conditions such as being immunocom-promised [9-11]. Rapid detection and identifi-cation of environmental mycobacteria at species level is therefore critical if infections in humans and animals are to be controlled. Sequencing of 16S ribosomal RNA gene has become one of the standard tools for molecular identification of bacteria such as Mycobacterium species [12-14]. The objective of this study was to isolate, detect and identify Mycobacterium species of public health importance in cattle in Zimbabwe using 16S ribosomal RNA gene amplification and sequencing. The study was of great interest since humans live in close proximity with cattle in Zimbabwe and there is always a danger of zoonotic transmission of the bacteria especially to immunocompromised people.
METHODS
Collection and Laboratory Processing of Cow Dung Samples
The study was conducted from October 2013 to May 2014 in Zimbabwe, a country in Southern Africa. Fresh cow dung samples were collected around the country from cattle in districts of Bikita, Bindura, Chihota, Chitungwiza, Chivhu, Chiredzi, Chiweshe, Domboshava, Epworth, Goromonzi, Hatcliffe, Karoi, Kuwadzana, Lupane, Masvingo, Mavambara, Mazowe, Mhondoro, Mount Hampden, Mufakose, Musami, Murombedzi, Msengezi, Mvurwi, Norton, Seke, Snake Park, Zvimba and Zvishavane, The cow dung samples were stored at 4 oC before processing. All the samples were first decontaminated using 4% sodium hydroxide before culture to kill all other microorganisms save mycobacteria. One millitre of each of the decontaminated cow dung samples was inoculated on Peizer’s TB solid medium. The cultures were incubated at 37°C and examined daily for up to four weeks until visual colonies were seen. Smears of potential mycobacteria colonies were stained using the Ziehl-Neelsen. Colonies which were AFB positive were purified by subculturing on Peizer’s TB medium. Pure colonies from each culture were kept at -80 °C in 200 µl of distilled water until DNA extraction.
DNA Extraction from Mycobacterium Isolates
DNA was extracted from Mycobacteria culture isolates using Quick-gDNA™ MicroPrep (Zymo Research Corp, U.S.A) according to manufacturer’s recommendations. Briefly, bacterial isolates previously stored in 200 µl distilled water at -80°C were thawed at room temperature and 800 µl of Genomic Lysis Buffer added. After lysis, the supernatants were transferred to the Zymo- SpinTM columns and centrifuged at 10 000 g for 1 minute. The columns were then washed with 200 µl of DNA Pre-wash buffer, followed by 500 µl of gDNA Wash buffer. The DNA was eluted from the columns using 50 µl of DNA Elution buffer. The extracted Mycobacterium DNA was stored at -20°C until polymerase chain reaction of the 16S rRNA gene.
Amplification of 16S rRNA Gene by Polymerase Chain Reaction
The 5'-end of the 16S rRNA gene was amplified by polymerase chain reaction from the extracted Mycobacterium DNA. Each amplification reaction contained 32.8 µl distilled water, 0.2 µl Taq polymerase buffer (5U/µl stock), 3 µl MgCl2 (25 mM stock), 1 µl dNTPs (10 mM stock), 2 µl 16S rRNA forward primer (CCTGC-ACTTCGGGATAAGCCTG) (10 µM stock), 2 µl 16S rRNA reverse primer (CAACGCG ACAAACCACCTACGA) (10 µM stock), 5 µl 10X PCR Buffer and 5 µl of template DNA. The two primers were designed by Dr N Chin’ombe using Geneious Basic program (Biomatters, USA) to specifically amplify the 5'-end of the rRNA gene of Mycobacterium species. A thermocycler (Perkin Elmer GeneAmp PCR System 2400) was used for amplification with the following cycling program: initial denaturation of 5 minutes at 95°C followed by 35 amplification cycles of 30 seconds at 95°C, 30 seconds at 55°C, and 45 seconds at 72°C, and ending with a final extension step of 7 minutes at 72°C. The PCR products were analyzed by 2% agarose electrophoresis.
DNA Sequencing and Bioinformatics Analysis
PCR products with strong bands of the expected size (approximately 460 bp) were selected and sent to Inqaba Biotechnical Industries (South Africa) for sequencing. DNA sequence data (chromatographs and sequences) for all the samples were sent back by email for analysis. DNA sequences were analyzed using the Geneious Basic program (Biomatters, USA) and BLAST programs. Different Mycobacterium species were identified by comparison of the alignment with sequences in the Genbanks.
RESULTS
Mycobacterium Isolates
During the study, a total of 134 cow dung samples were collected from several districts of Zimbabwe (see Fig.(1) for one of the study sites). Out of all the samples (n = 134) that were cultured successfully, only 49 were positive by AFB smear. These were successfully sub-cultured to generate pure isolates. Different colony colours were observed on the Peizer’s TB medium (Fig. 2). The colony colors ranged from white, cream, yellow to peach (Fig. 2). The colonies had various types of morphology, with some being smooth, rough, and others shiny (Fig. 2).

One of the sites in Bikita district of Zimbabwe where cow dung was collected for Mycobacterium analysis.
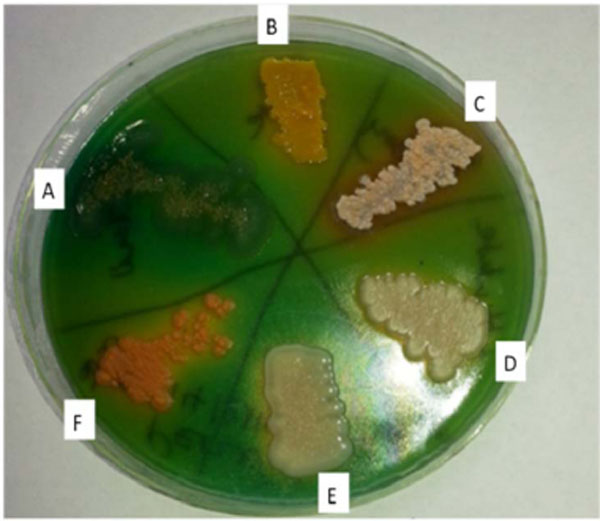
Typical examples of pure culture isolates growing on Peizer's TB medium. Only 6 representative cultures (A-F) are shown in this figure.
Amplification of Mycobacterium 16S rDNA Gene
Out of the 49 DNA samples amplified by PCR, 41 (84%) showed the expected band of approximately 460 bp (Fig. 3). There was no amplification in 8 (16%) of the samples. Out of the 41 samples that showed amplification, 26 (63%) had strong PCR bands and were selected for DNA sequencing of the 16S rRNA gene. The 15 samples (37%) with faint bands, though of the expected size, were not sequenced.
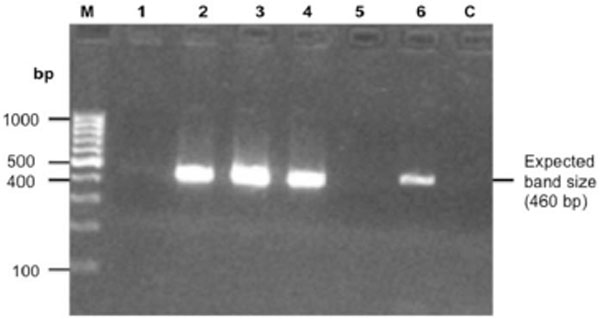
Representative gel of PCR amplicons of Mycobacterium 16S rRNA gene. M is the molecular weight marker, lane 1 to 6 contains the samples and lane C is the negative control. There was no amplification for samples 1 and 5 and there was amplification for samples 2, 3, 4 and 6 with a DNA band of approximately 460 bp.
Molecular Identification of Mycobacterium Species
DNA sequences of the 26 bacterial isolates were analyzed. A phylogenetic tree was constructed to check the closeness of the sequences (Fig. 4). The sequences were also compared against Mycobacterium tuberculosis (Accession number JX303298) and M. bovis (Accession number AM408590) sequences (Fig. 4). It was noted that some sequences had 100% homology while others had few base differences. None of the sequences from this study had 100% homology with sequences of M. tuberculosis or M. bovis (Fig. 4). The samples were grouped into clusters based on the phylogenetic analysis (Table 1).

Phylogenetic analysis of the 26 16S rRNA sequences of the bacterial isolates from cow dung collected in Zimbabwe. Mycobacterium bovis and Mycobacterium tuberculosis sequences were included for comparison.
Molecular identification of Mycobacterium species (n = 26). The bacterial isolates were grouped into 7 clusters according to phylogenetic data.
| Cluster | Frequency (%) | Mycobacterium Species Identified |
|---|---|---|
| I (S1, S4, S5, S8, S9, S16, S26) | 7 (27%) | Mycobacterium neoaurum |
| II (S3, S6, S10, S13, S21, S25) | 6 (23%) | Mycobacterium fortuitum |
| III (S12, S18, S23) | 3 (12%) | Mycobacterium goodii |
| IV (S2, S24) | 2 (1%) | Mycobacterium arupense |
| V (S11, S17) | 2 (1%) | Mycobacterium peregrinum or M.septicum |
| VI (S20) | 1 (0.04%) | Mycobacterium elephantis |
| VII (S7, S14, S15, S19, S22) | 5 (19%) | Non-Mycobacterium species (Gordonia terrae) |
The clusters containing related samples were identified to species level using the Geneious Basic program that was used to search all gene banks available for DNA sequence similarities (Table 1). Cluster I sequences were found to belong to Mycobacterium neoaurum. All the 7 sequences (S1, S4, S5, S8, S9, S16, S26) in this group had 100% similarity with each other. They aligned 100% with several strains of Mycobacterium neoaurum from the gene banks. Sequences from Cluster II (S3, S6, S10, S13, S21, S25) were 100% similar and they aligned 100% with strains of Mycobacterium fortuitum from the gene banks. The 3 sequences (S12, S18, S23) in cluster III were also 100% similar and aligned 100% with strains of Mycobacterium goodii. Cluster IV sequences aligned to Mycobacterium arupense. The two sequences in this cluster had 99.8% homology and differing by 1 base. Alignment of the 2 sequences with known Mycobacterium species from the gene banks showed that S24 was 100% similar with M arupense strains while S2 sequence was also closely related to M arupense (99.8% similarity). Cluster V sequences (S11 and S17) had 100% homology with both Mycobacterium peregrinum and M. septicum sequences from the gene banks. Cluster VI (S20) sequence matched 100% with Mycobacterium elephantis sequences. Cluster VII (S7, S14, S15, S19, S22) sequences did not match with any Mycobacterium species from the gene banks. They however matched with sequences from Gordonia terrae, which is a close phylogenetic relative of Mycobacterium species.
DISCUSSION
Cattle are hosts to a number of Mycobacterium species, some of which are of public health importance [6,8]. Environmental mycobacteria, including M. bovis from cattle can cause disease and other opportunistic infections in humans [5,6]. Cattle acquire environmental mycobacteria mainly through grazing and drinking contaminated water. Humans, particularly in Zimbabwe, live in close proximity with cattle (Fig. 1) and this poses a high risk of zoonotic transmission of Mycobacterium species. The use of cow dung as floor polish, plastering and cooking is widely practiced in rural Zimbabwe and this can be another source of infection in humans by Mycobacterium species. Humans can also get infected by consuming food products such as milk from infected livestock [8].
No previous studies have been done to investigate the Mycobacterium species that are prevalent in cattle in Zimbabwe. In this study, we set out to isolate, detect and identify Mycobacterium species present in fresh cow dung in Zimbabwe. Using a molecular approach, we were able to detect and identify various Mycobacterium species from the cow dung (Table 1). The species isolated were M. neoaurum,M. fortuitum, M. goodii, M. peregrinum or M. septicum, M. arupense and M. elephantis. The sequences, S11 and S17 needed further identification since they aligned 100% with both M. peregrinum and M. septicum species. In this study, some of the species we identified have already been detected in cattle in other studies. In Brazil, nine different species of environmental mycobacteria were isolated (M. nonchro-mogenicum, M. peregrinum, M. smegmatis, M. neoaurum, M. fortuitum, M. chelonae, M. flavescens, M. kansasii and M. scrofulaceum) in milk from cattle [15]. Most of the identified Mycobacterium species in this study were also of public health importance as they can potentially cause diseases in humans especially those with AIDS and are immunocompromised. M neoaurum has been shown to cause bacteraemia and other pulmonary infections [16,17]. M. fortuitum has been shown to be associated with several cases of wound and catheter infections [18, 19]. M. goodii was first isolated in 1999 and has been found to be associated with some human infections [20, 21]. M. peregrinum infects surgical sites and only few cases of infection have been documented [22]. M. septicum was previously recovered in blood cultures [23, 24]. M arupense has been isolated in a number of clinical specimens and may play a role in causing infection [25-27]. M. elephantis was first isolated from an elephant that died from chronic respiratory disease [28]. Since then, the species has been isolated from a number of human specimens in some countries, but its clinical relevance is still undetermined [29]. We also identified a phylogenetically close relative of Mycobacterium, Gordonia terrae that is also of public health importance as it can cause various health problems in humans [30-32]. The 8 ZN positive strains which did not show a PCR product could belong to the genera Rhodococcus, Nocardia or Tsukamurella. The genera Rhodococcus, Nocardia, Tsukamurella and Gordonia are also closely related to the Mycobacterium genus and they are also acid-fast rods. We have also noted that although the two primers used for PCR were specific for Mycobacterium, they could also weakly bind to the 16S rRNA gene of Rhodococcus, Nocardia, Tsukamurella and Gordonia species. This explains why some PCR bands were not very bright. There is therefore a need to further characterize these isolates that are found in animals as they can potentially cause various health problems in humans.
CONCLUSIONS
In conclusion, several Mycobacterium species were identified for the first time in cow dung in Zimbabwe and most of the Mycobacterium species identified are of public health importance. The Mycobacterium species detected are known to be potentially pathogenic to humans especially those with underlying medical conditions. Their rapid detection and identification using molecular approaches such as 16S DNA sequencing are important.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflict of interest.
ACKNOWLEDGEMENTS
The study was supported by the University of Zimbabwe, National University of Science and Technology and Government of Zimbabwe’s Central Veterinary Laboratory. Sequencing of 16S rDNA PCR products was done at Inqaba Biotech, South Africa.


