All published articles of this journal are available on ScienceDirect.
Pathogens in Urine from a Female Patient with Overactive Bladder Syndrome Detected by Culture-independent High Throughput Sequencing: A Case Report
Abstract
Introduction:
Overactive bladder syndrome (OAB) is described as urgency, with or without urgency incontinence. A range of medical conditions shares the symptoms of OAB, however the diagnosis is contingent on the exclusion of urinary tract infection (UTI). Knowing that urine dipstick and routine culture of bacteria can miss UTI diagnosis caused by low-count bacteriuria or “difficult-to-culture” pathogens, we examined a case of OAB with a culture-independent approach.
Case presentation:
A 61-year-old Norwegian female with a long history of urinary symptoms and a diagnosis of OAB was selected as a suitable subject for a culture-independent 16S rDNA analysis on the patient´s urine. The patient’s medical records showed no history of recurrent UTI, however, when the urine specimen was sent to routine culture at the time of study it showed a significant bacteriuria caused by a single bacterium, and the patient was prescribed antibiotics. The 16S rDNA analysis revealed not one, but many different bacteria, including a considerable amount of fastidious bacteria, indicating a polymicrobial state. One year later, the subject was still experiencing severe symptoms, and a follow-up analysis was performed. This time the urine-culture was negative, however, the 16S rDNA profile was quite similar to that of the first sample, again displaying a complex bacterial profile.
Conclusion:
The use of 16S rDNA pyrosequencing and sequence analysis to uncover “difficult-to-culture” bacteria should be considered when examining patients with chronic urinary symptoms. These methods may contribute to further elucidation of the etiology of overactive bladder syndrome and other urinary syndromes.
INTRODUCTION
Overactive bladder (OAB) syndrome is defined by the International Continence Society as urinary urgency with or without urge incontinence, along with increased frequency and nocturia in the absence of urinary tract infection (UTI) or any other obvious pathology [1]. This chronic condition often remains under-reported, under-treated and negatively impacts quality of life [2, 3]. The prevalence of OAB is estimated to be from 12 to 20% and is known to increase with age [4-6]. While gender does not appear to be a risk factor, the vast majority of patients receiving treatment are women [7]. The treatment mainly includes anticholinergic medications and bladder training, physiotherapy and electrical stimulation [7, 8].
The etiology of OAB is not completely understood and is likely multifactorial. The symptoms may be similar to those seen in other conditions like UTI, chronic urinary conditions, benign prostatic and bladder cancer [9]. There is no gold standard for the diagnosis. It is rather a diagnosis of exclusion, done based on patient’s history, physical examination to exclude other urogenital pathology and a voiding diary. However, culture-negative urine is a prerequisite for the diagnosis and UTI must be ruled out before making a diagnosis of OAB.
The standard way of excluding UTI in a patient with OAB symptoms in a clinical setting is urinalysis using a dipstick or mid-stream urine culture [10]. The dipstick is an indirect measure and identifies the bacterial conversion of nitrate to nitrite. For culture analysis, the traditional definition for a positive culture and a significant bacteriuria is >105 colony-forming unit per ml (CFU/ml). However, the reliability of these diagnostic aids in relation to OAB symptoms has recently come under scrutiny. Reports show that dipstick screening is not sufficient for UTI diagnosis in patients with symptoms of OAB [11, 12]. Further, the routine method for culturing urine applied to patients with OAB symptoms may also fail to spot genuine UTIs; due to the high microbial threshold required, fastidious bacteria can easily evade the culture conditions and intracellular microbes may also be present [13-15]. Lowering the threshold for a significant bacteriuria and use of enhanced culturing techniques has been advised, but the traditional methods remain common practice in most clinics.
The above-mentioned diagnostic limitations are evident in for example diagnosis of Ureaplasma spp.; these bacteria can easily escape detection by conventional microbiological investigations of urine samples due to slow or no growth on standard media. It is now recognized that these organisms play a more important role in human infections than previously anticipated and studies have shown a strong association between Ureaplasma spp. and urinary tract diseases including UTI, urethritis, and chronic urinary symptoms [16-18]. Of special interest with regard to OAB are findings by Lee et al. (2010) [19] documenting Ureaplasma urealyticum infection in 43% of the female patients with OAB symptoms included in their study.
Advances in molecular-based methods like 16S rDNA sequencing have made overcoming the limitations of conventional microbial methods possible. Moreover, the progress in sequencing technology, such as high throughput sequencing, has allowed for extensive characterization of microbial populations in a massive parallel and cost-effective manner [20-26]. High throughput sequencing is giving a deeper insight than what was previously possible, and provides an increased knowledge about the urine microbiota in health and disease [20-23, 27]. These studies have revealed the polymicrobial nature of urine specimens and have clearly shown that anaerobic and “difficult-to-culture” bacteria are present even in culture-negative urine samples.
Here, we report the case of an OAB patient whose 16S rDNA analyses of two consecutive urine samples indicated a polymicrobial state. In both samples Ureaplasma was identified with relatively high abundance. The first urine sample was reported to be culture-positive with a single bacterium, while the second sample was reported culture-negative. Ureaplasma was neither reported nor investigated by the routine clinical lab, which followed the traditional guidelines for examining patients with chronic urinary OAB symptoms.
CASE PRESENTATION
A 61-year-old postmenopausal Norwegian female with a history of more than 10 years of urinary tract symptoms with urgency and urinary incontinence was studied. Having negative urine stick analyses, the diagnosis of OAB was made as an exclusion diagnosis based on absence of bladder pain, and most importantly, cystoscopy examination to rule out other urological disorders (e.g. interstitial cystistis, bladder cancer). Complaining of severe symptoms, the patient had been treated with various anticholinergic agents (Cetriprin, Detrusitol, and Vesicare) with only periodical relief. For her persistent symptoms, she had now been started on electric stimulation treatment. Finding the treatment satisfactory and wanting to continue, she was encouraged by the clinic to contact them as needed.
Knowing that patients with chronic urinary symptoms may actually suffer from an undetected UTI (e.g. low-count bacteriuria or “difficult-to-culture” pathogen), we investi-gated the urine sample from our case subject with a cultivation-independent approach. We characterized the urine microbiota using 16S rDNA amplicon analysis for two variable regions V1V2 and V6. Total DNA isolation, 16S rDNA amplification, followed by 454 pyrosequencing and sequence read analysis for taxonomic grouping of the reads was performed as described by Siddiqui et al. (2011) [20].
We performed both a urine dipstick analysis and a routine culture to ensure that the patient, as previously, did not have any traditionally detectable UTI. The urine stick analysis was negative, however she had a positive urine culture; she was reported to be infected with α-hemolytic green Streptococcus > 105 colony-forming unit (CFU/ml) (Streptococcus viridians group) (see Table 1). Antimicrobial susceptibility was determined and she was prescribed Trimethoprim 160 mg x twice a day for a week. In contrast to the clinical reports, the molecular analysis revealed a complex bacterial composition of the urine sample taken (Fig. 1A, 1B). Consistent with the culture report, Streptococcus was observed (sequences assigned to α-hemolytic Streptococci). The abundance of the bacterial sequences assigned to this genus was 23% and 30% for both V1V2 and V6 analysis. Several fastidious bacteria not identifiable by routine culture were also present. Atopobium was the major genus identified, with a total sequence abundance of 24% and 33,5% for V1V2 and V6, respectively. Interestingly, Ureaplasma was also present with a relatively high sequence abundance of 17% for V1V2 analysis, although V6 analysis only gave an abundance of 1.2%. Prevotella was also identified with an abundance of 16% for V1V2 and 8.6% for V6 analysis. The species Bacteroides urealyticus was also detected with abundances of 10.8% and 5.8% for V1V2 and V6, respectively.
Urine culture and antibiotic resistance determination
| Microbe: | |
|---|---|
| Green α-hemolytic green Streptococcus >100, 000 CFU/ml | |
| Determination of resistance: | |
| Trimethoprim | S |
| Sulpha/Trimethoprim | S |
| Cephalotin | S |
| Ampicillin | S |
| Nitrofurantoin | S |
| Mecillinam | R |
S = sensitive, R = resistant
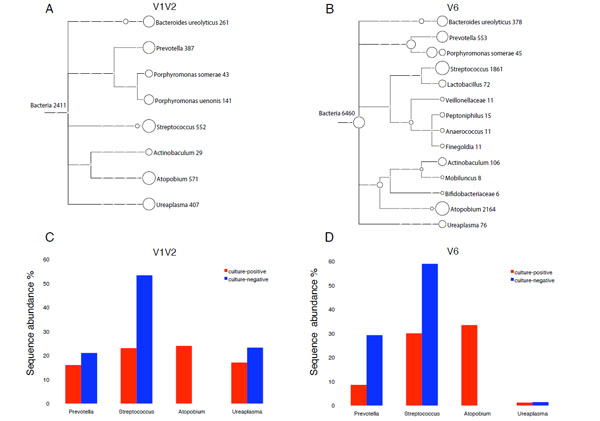
Taxonomical classification of 16S rDNA found in reported samples.
A and B: Tree view of 16S rDNA sequences from culture-positive urine (α-hemolytic Streptococcus >105 CFU/ml), assigned at different taxonomy levels (family, genus and species) as computed by MEGAN 3.4 for V1V2 (A) and V6 (B) amplicons. Each circle represents a taxon and is labeled by its name and the number of sequenced reads assigned. Taxonomical assignment was done by comparing the sequences to a curated version of the SSUrdp database, then assigning the sequences to the taxon of the best-matched reference sequence (lowest common ancestor). The size of the circles is scaled logarithmically to the number of reads assigned to the taxon.
C and D: Comparison of bacterial communities from culture-positive urine positive for α-hemolytic green Streptococcus (red bars) and culture-negative urine (blue bars) from a case subject diagnosed with overactive bladder syndrome. Sequence analysis is based on V1V2 (C) and V6 (D) amplicons. Only major genera with a sequence abundance ≥10 % for either V1V2 or V6 analysis are shown. The calculation of sequence abundance for each genus was relative to the total number of sequence assigned to Bacteria in each dataset.
About a month later after the antibiotic treatment the clinical reports for both urine dipstick and routine culture were negative, a sign of recovery from the incidence of significant bacteriuria and UTI.
However, the urinary symptoms lasted and were persistent. A year later, we again examined the patient´s urine with the 16S rDNA approach as a follow-up. Urine stick and culture was also performed and both were again negative. In accordance with the previous observations, 16S rDNA analysis of the urine specimen showed a composite bacterial profile. However, the previously dominant genus Atopobium was completely absent this time (Fig. 1C,1D). Interestingly, Streptococcus, found in the first sample and the reason the antibiotic treatment was given, was still detected in the urine with two times higher sequence abundance, 53.4% for V1V2 and 59% for V6. Ureaplasma was again identified within the same range of sequence abundance as before: 23.3% for V1V2 and 1.4% V6. While the V6 analysis showed Prevotella with an increased sequence abundance of 29.3% from 8.6%, V1V2 analysis identified this genus in the same range as previously (21% of the bacterial sequences). This time, the species Bacteroides urealyticus was observed with only trace amounts (< 0.3%) for both amplicons.
DISCUSSION
This report describes the urine microbiome of a female patient with OAB diagnosis using high throughput 16S rDNA 454 pyrosequencing on two successive urine samples. The first sample was culture-positive for a single bacterium, while the second sample was reported negative. However, a complex bacterial profile, with fastidious and anaerobic bacteria, was observed by the 16S rDNA analyses for both urine specimens, demonstrating that urine from this OAB patient displays a polymicrobial profile. These findings are in agreement with other recent reports on human urine microbiome in both health and disease [20-23, 27]. Another important observation is that, with respect to the major bacterial community membership, the 16S rDNA sequence profiles detected for the two urine samples were fairly consistent, suggesting only small variations in the microbial diversity of urine of this OAB patient over the one-year period between the two urine samples.
Urine culture is the gold standard for assessment of UTI. However, conventional culture techniques favour fast-growing bacteria. This is readily demonstrated in our observations where the molecular diagnostic tools such as 16S rDNA amplicon pyrosequencing and bioinformatics analyses independently identify fastidious and “difficult-to-culture” organisms not found by the standard cultivation procedures. The disagreement between the culture report and the 16S rDNA approach in this case opens up the possibility of the patient’s clinical chronic symptoms having a bacterial cause after all. Further, the more or less constant microbiome observed with the constant urinary symptoms regardless of the antibiotic treatment in between the two sampling time points again suggest a correlation between the patient’s chronic condition and the urine microbiome.
Since the resolution gained by using a single hypervariable region of the 16S rDNA gene may not be sufficient we therefore performed our analyses covering two regions (V1V2 and V6), which yielded complementary results. The molecular analysis revealed a considerable number of fastidious bacteria (Fig. 1A, 1B) most of them obligate anaerobes e.g. Atopobium, Prevotella, Actinobaculum and Bacteroides urealyticum, already associated with different urogenital diseases like UTI, chronic urinary symptoms and bacterial vaginosis [27-29].
The genus Streptococcus comprises a heterogeneous group of bacteria, some are major pathogens and others members of the normal human microbiota. OAB is associated with increased health risks and it has been suggested that these patients may be more prone to bacteriuria and UTI [30]. However, the clinical relevance of α-hemolytic green Streptococcus in OAB patients is unknown, in fact it is often regarded as non-uropathogen [31].
Like Streptococcus, Ureaplasma was present in both samples investigated. V1V2 analysis indicates its constant presence with an abundance of ~ 20%. Ureaplasma species are among the most common bacteria isolated from the human urogenital tract [32] and are often regarded as commensals. However, Ureaplasma has been shown to have a causal role in urogenital infections [33, 34] and their prevalence in chronic urinary symptoms in females has also been noted [17-19]. The slow growing and “difficult-to-culture” nature of Ureaplasma can cause chronic infections to be undetected and its possible intracellular localisation may protect from the host’s immune system [34-36]. It does not respond to conventional antibiotic treatment (e.g. Trimethoprim or Sulphonamides) for urinary tract pathogens [16, 33, 37]. It can therefore be hypothesized that Ureaplasma infections in females with chronic urinary symptoms are underreported. According to the patient’s hospital records, urine samples were not examined for Ureaplasma previously. However, our 16S rDNA high throughput sequencing easily identified this potential pathogen. Whether or not this “difficult-to-culture” bacteria is causing the chronic symptoms in this particular situation remains to be established.
CONCLUSION
The presence of “difficult-to-culture” organisms, which may be pathogenic, is clearly indicated in this case of OAB. Bacterial profiling of urine using high throughput 16S rDNA pyrosequencing in chronic urinary tract patients is therefore a promising method for diagnostics. An in-depth characterization of the urine microbiota of a particular case could direct the clinicians to a more precise diagnosis and treatment. This approach is now feasible in the routine diagnostic lab due to drastically reduced cost associated with sequencing.
The results presented in this study also suggest that 16S rDNA studies on a larger number of women with OAB diagnosis could be performed to explore which role Ureaplasma possibly play in development and persistence of this disease.
CONSENT
The Regional Committee for Medical Research Ethics East-Norway has approved the investigation (REK Øst Prosjekt 110-08141c 1.2008.367) and written informed consent was obtained from the patient for publication of this case report. A copy of the written consent is available for review.
CONFLICT OF INTEREST
The authors have no conflict or competing interests.
AUTHORS' CONTRIBUTIONS
HS, AJN, SLJ and KSJ were involved in study design; HS processed the samples and carried out the molecular techniques. KL and HS performed the bioinformatics and taxonomic analysis. HS interpreted the data and authored the manuscript. All authors edited and commented on the paper and all authors read and approved the final manuscript.
ACKNOWLEDGEMENTS
The authors would like to thank Hege Junita Gaup for technical assistance and the Norwegian Sequencing Centre (NSC, www.sequencing.uio.no), Department of Biosciences, University of Oslo, for sequencing services. We are very grateful to urotherapists Turid H Hoel and Bodil Svendsen at Aker University Hospital HF, Urological Clinic for specimen collection.
Financial support for this research was provided by grants from the Research Council of Norway to KSJ and from CEES to HS.


