All published articles of this journal are available on ScienceDirect.
Microbial Efflux Systems and Inhibitors: Approaches to Drug Discovery and the Challenge of Clinical Implementation
Abstract
Conventional antimicrobials are increasingly ineffective due to the emergence of multidrug-resistance among pathogenic microorganisms. The need to overcome these deficiencies has triggered exploration for novel and unconventional approaches to controlling microbial infections. Multidrug efflux systems (MES) have been a profound obstacle in the successful deployment of antimicrobials. The discovery of small molecule efflux system blockers has been an active and rapidly expanding research discipline. A major theme in this platform involves efflux pump inhibitors (EPIs) from natural sources. The discovery methodologies and the available number of natural EPI-chemotypes are increasing. Advances in our understanding of microbial physiology have shed light on a series of pathways and phenotypes where the role of efflux systems is pivotal. Complementing existing antimicrobial discovery platforms such as photodynamic therapy (PDT) with efflux inhibition is a subject under investigation. This core information is a stepping stone in the challenge of highlighting an effective drug development path for EPIs since the puzzle of clinical implementation remains unsolved. This review summarizes advances in the path of EPI discovery, discusses potential avenues of EPI implementation and development, and underlines the need for highly informative and comprehensive translational approaches.
INTRODUCTION
The 20th century gave rise to many successful methods for preventing and controlling infectious diseases but fostered the mindset that the war against infectious microbes was over. In the 1980s consensus among pharmaceutical companies was that there were enough antibiotics already on the shelf and research efforts were redirected elsewhere [1]. Optimism was short-lived however when outbreaks and epidemics of new, re-emerging, and drug-resistant infections arose. Termed “superbugs”, emergent microorganisms possessing effective and dynamic pathogenic capabilities continue to be a dangerous threat. Each year over 13 million deaths worldwide are attributed to the emergence of new infectious diseases or to the re-emergence of diseases previously thought well-controlled.
One major component of resistance to many classes of antimicrobials as well as chemotherapeutic agents is multidrug efflux [2]. Efflux results from the activity of membrane transporter proteins known as multidrug efflux systems (MES) [3, 4]. MES perform essential roles in cellular metabolism and differ in membrane topology, energy coupling mechanisms, and, most importantly, substrate specificities [5]. Identifying natural substrates and inhibitors of efflux systems is an active and expanding research topic [6]. Based on their sequence similarity, efflux systems have been classified into the following six super-families (Fig. 1): ATP-binding cassettes (ABC), Multidrug and Toxic compound efflux (MATE), resistance-nodulation cell division (RND), small multidrug resistance family (SMR), multi-antimicrobial extrusion protein family, and multidrug endosomal transporters (MET). The first five families are found in microorganisms while the MET family appears restricted to higher eukaryotes. Representatives of all groups are expressed in mammalian cells [7]. ABC transporters are the largest super family, containing seven subfamilies designated A to G based on sequence and structural homology [8].
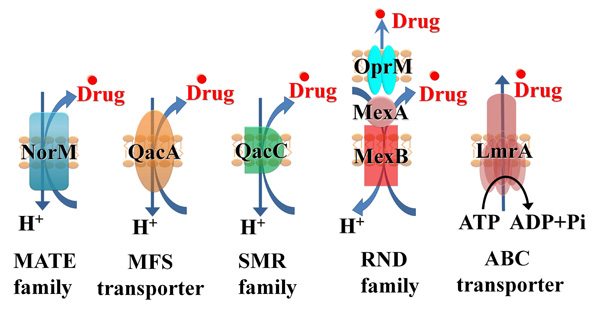
Schematic illustration for key members of the 5 super-families of the microbial efflux systems: NorM, multi-antimicrobial extrusion protein family (MATE), QacA major facilitators (MFS), QacC small multidrug resistance family (SMR), MexAB, resistance-nodulation cell division (RND), LmrA, ATP-binding cassettes (ABC).
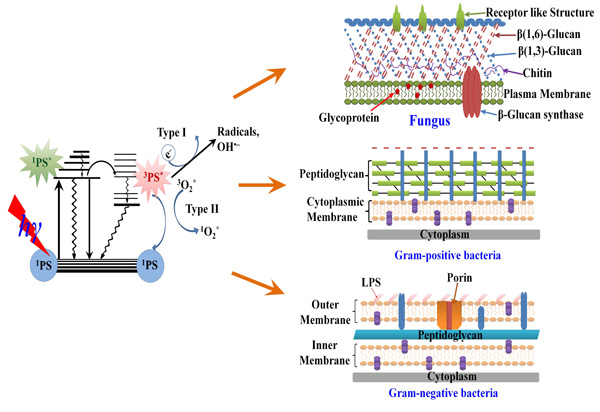
Schematic illustration for: the mechanism of photodynamic inactivation including the Jablonski diagram (left) and the microbial permeability barriers of Gram-positive, Gram-negative bacteria and fungi (right). The PS initially absorbs a photon that excites it to the first excited singlet state and this can relax to the more long lived triplet state. This triplet PS can interact with molecular oxygen in two pathways, type I and type II, leading to the formation of reactive oxygen species (ROS) and singlet oxygen respectively.

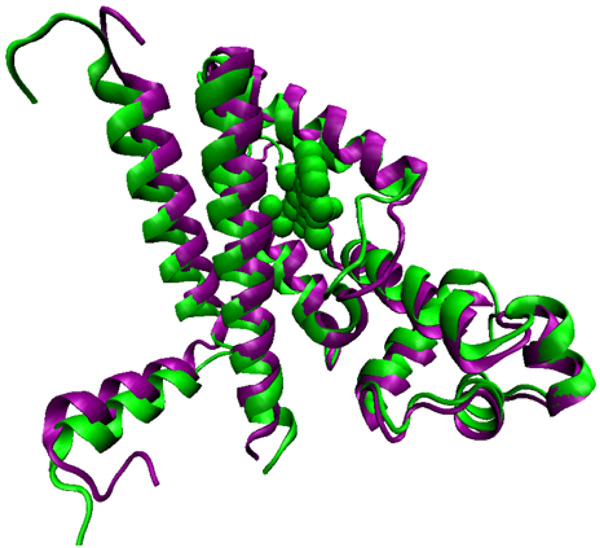
Left: Methylene blue PS bound to a hydrophobic pocket of an ABC transporter. The electrostatic potential varies from red (-) to blue (+). Right: The dye (blue tubes) shares the site with berberine (green tubes), a natural antimicrobial and also an ABC substrate.
Structure of the transcriptional repressor TetR in its apo form (violet) and bound to the tetracycline-Mg2+ complex (green); the latter is not competent for binding the TetA MFS promoter; this yielding expression of the MFS just upon antibiotic challenge.
The molecular mechanism of the transporter substrate recognition, the kinetics, the stoichiometry and the features of the antiport process, is an ongoing field of investigation. Structural studies indicate that the transporters may capture their substrates directly from the periplasm or from the outer leaflet of the cytoplasmic membrane [9,10]. In retrospective, the key steps of drug transport involve several and specific interplays between intramolecular and intermolecular conformational changes and intermolecular dynamics; It will be impossible to dissect without a coordinated effort that involves crystallography, molecular modeling, molecular biology, energy calculation and biological–functional investigations. The scenario is more complicated since some drug transporters show a cooperative interaction or can act sequentially [11,12].
A fundamental challenge associated with drug transport both for RNDs and ABCs efflux systems independently of the mechanistic differences between antiport and energy dependent efflux is associated is the broad specificity (or poly-specificity). It is very difficult to distinguish between a ‘single’ binding step and a correct recognition binding inside the high-affinity pocket with an active conformation, allowing efficient translocation of the drug to the outer membrane channel. The structural data including molecular docking calculations have provided some information [13], but they are rather far from model development without biological support [14,15]. The energy source is a key element of transport function and substrate recognition as for at least the proton antiporters efflux is a pH- dependent process [16]. This is also an active field of investigation in a variety of pathogens [17,18].
In order to address the obstacles of efflux to chemotherapy, an array of approaches have been investigated i) By-passing efflux systems by improving the molecular design of old antibiotics in order to reduce their efflux. ii) Decreasing the efficacy of the membrane barrier by a direct modification or a channel-blocker that induces a ‘traffic jam’ in the outer membrane channel restoring a high intracellular drug concentration. iii) Blocking the efflux capacity of microbial cells through competitive, non-competitive inhibition as well as direct or indirect energy depletion via an anti-porter site or via a collapse of energy driven mechanisms of the microbial cell envelope.
Natural products play a major role in drug discovery by providing bioactive scaffolds with activity against a variety of targets in infections and cancer. The strategy of efflux system inhibition employing natural product efflux pump inhibitors (EPIs) emerged in the last decade. Reports for novel and already known natural chemical entities which are active against virtually all of the major microbial efflux systems have populated the chemical space as well as the literature [6, 19]. The concept of restoring and enhancing the utility of antimicrobials by employing EPIs is appealing but is not yet at a therapeutic stage. Discovery efforts are multidimensional and continuously expanding but a number of conceptual and methodological gaps are barring identification of lead EPIs for clinical implementation. This review summarizes the tools for EPI discovery and validation with emphasis on those of plant origin. The path forward towards development and deployment in a clinical setting is also outlined.
MES in Gram-positive Bacteria
The major efflux systems implicated in Gram-positive bacterial related drug resistance are the chromosomally encoded MFS Nor-family (NorA, NorB, NorC), MdeA the MATE mepRAB (multidrug export protein) [20] and the SMR SepA [21]. There is also evidence for plasmid encoded systems such as QacA, QacB, [22,23] and Tet(K) that function as tetracycline-divalent metal complex/H+ antiporters [24]. A NorA homolog (EmeA) has also been validated in Enterococcus faecalis [25]. These systems have a broad and overlapping substrate specificity including quinolones, tetracyclines, monovalent and divalent antimicrobial cations (intercalating dyes, quaternary ammonium compounds, diamidines, biguanidines) and plant secondary metabolites [26, 27].
MES in Gram-negative Bacteria
Multidrug-resistant (MDR) and pandrug-resistant (PDR) Gram-negative bacteria, pose a grave threat of truly untreatable infections [28,29]. Efflux systems are key players in a variety of challenging clinical conditions. The resistance nodulation cell division (RND) family includes proton-driven systems with key members the AcrAB-TolC in E. coli with strong homology in Francisella tularensis,Yersinia pestis, and Brucella sp. AcrB functions as a multi-subunit complex in association with the outer membrane channel TolC and the membrane fusion protein AcrA. There are 12 RND-type efflux systems present in Pseudomonas aeruginosa of which three, MexAB-OprM, MexCD-OprJ, and MexXY-OprM have been shown to accommodate and provide resistance to β-lactams and fluoroquinolones (with the addition of MexEF-oprN) [30]. The partition of efflux in the resistance related to aminoglycosides, antimicrobial peptides and biofilm formation is established but is part of a more sophisticated cell response [30]. The characteristics of additional RND-type systems such as MexABC-OpmB [31] and TriABC-OpmH, a triclosan efflux pump, have been recently dissected [32]. The mex-locci in P. aeruginosa are phylogenetically related with a set of homologues systems in 3 key pathogens:
- The 11 Bpe-efflux systems in Bulkhoderia pseudomallei, and B. mallei [33]. BpeAB-OprB is a broad-spectrum MES which is only expressed at low levels and only marginally contributes to multidrug resistance (MDR). In contrast, strains expressing BpeEF-OprC are highly resistant to most of the clinically useful antibiotics for melioidosis treatment.
- The 16 RND efflux systems present in B. cenocepacia (Bcc complex, RND-1 to -16) [34] and bmeABC1-16 in the Bacteroides fragilis [35]. At least seven BmeB efflux pumps are functional in transporting antimicrobials and have overlapping substrate profiles, and at least four confer intrinsic resistance [35].
- At least 5 ade RND systems have been fully functionally characterized in Acinetobacter baumannii including AdeIJK [36] adeSR-adeABC, adeDE [37] and AdeFGH [38].
ABC efflux systems have been identified in several occasions including the heterodimeric SmdAB in Serratia marcescens [39] and the E. coli macrolide transporter MacAB [40]. The later has been also validated in Neisseria gonorrhoeae [41].
A number of MFS with efflux capabilities are present in Gram-negative bacteria. This list includes the Salmonella enterica sv. Typhimurium SmvA, most similar to QacA of S. aureus [42], the EmrAB-TolC in E. coli [43], and the QepA and OqxAB plasmids involved in quinolone resistance in E. coli and Klebsiella spp. [44] but with a wide distribution in Enterobacteriaceae. CraA in A. baumannii is homologous to the MdfA of E. coli, which extrudes only chloramphenicol and the acquired narrow-spectrum systems TetA, TetB, CmlA, and FloR [45]. AmvA, mediates antimicrobial and disinfectant resistance in A. baumannii [46]. Members of the SMR and MATE families have been detected in a variety of Gram-negative bacterial species. AbeM and AbeS have been characterized in A. baumannii, as have AdeXYZ, AdeDE and QacE for other Acinetobacter spp. [45]. SugE, contributes to antimicrobial resistance in Enterobacter cloacae [47].
MES in Mycobacteria
The Mycobacterial complex possesses a remarkable variety of efflux systems which are either members of the MFS or ABC super families. Mycobacterium tuberculosis presents one of the largest numbers of putative drug efflux pumps compared with its genome size. Bioinformatics as well as direct and indirect evidence have established relationships among drug efflux with intrinsic or acquired resistance in M. tuberculosis [48]. Tap is the first MFS system identified (Rv1258 in the annotated sequence of the M. tuberculosis genome) [49]. Genome analysis and homology search between the identified transporters and proteins characterized in other organisms have revealed 16 open reading frames encoding putative drug efflux pumps belonging to the MFS. The efflux function has been demonstrated in at least three of these including the sequence Rv0849. [50,51]. The list is expanding to include ABC Superfamily members such as DrrAB [52] Bcg0231 (M. bovis BCG) Rv0194 [53] Rv1218c[51] Rv1456c-Rv1457c-Rv1458c [54] and the SMR systems Rv3065 [51] LfrA in M. smegmatis [55] as well as the homologues of M. tuberculosis Rv1145, Rv1146, Rv1877, Rv2846c (efpA), [56] and Rv3065 (mmr and emrE) in both M. tuberculosis and M. smegmatis [57].
MES in Fungi
The best-studied families of fungal MES are from Saccharomyces cerevisiae, especially those responsible for pleiotropic drug resistance (PDR) [58]. Members of this family are highly conserved and are often responsible for drug-resistance among pathogenic fungal species [59]. Although fungal cells contain many genes for both types of systems, clinical azole resistance is most often associated with overexpression of ABC transporters [60]. Clinically important PDR transporters include C. albicans Cdr1p (CaCdr1p) and CaCdr2p, which are homologs of the S. cerevisiae Pdr5p (ScPdr5p) and mammalian G-type ABC transporters [60-62]. Fungal PDR efflux systems have relatively promiscuous substrate specificities that are presumably defined by their transmembrane domains. Analysis of the C. albicans genome has identified several efflux pumps (CDR1, CDR2, CDR3, CDR4, CDR5, SNQ2, and YOR1) belonging to the ABC superfamily.
Role of MES in Antimicrobial PDT
Given the broad spectrum of multidrug efflux pumps to raise resistance against various antibiotics and antimicrobials, an unconventional antimicrobial discovery effort is needed [1]. Photodynamic therapy (PDT), is a light-based technology platform [63], which uses harmless visible light in combination with non-toxic photosensitizers (PS) to control infections (Fig. 2). Historically PDT has had a prominent role in the cancer therapeutics and is also currently used to treat age-related macular degeneration [64]. Currently PDT is being investigated as an alternative treatment for localized infections [65]. PSs are usually organic aromatic molecules with a high degree of electron delocalization [66]. Porphyrins, chlorins, bacteriochlorins, phthalocyanines as well as a plethora of dyes with different molecular frameworks have been proposed as antimicrobial PSs [67,68]. These dyes include halogenated xanthenes (e.g. Rose Bengal (RB), [69] perylenequinones (e.g. hypericin), phenothiazinium dyes such as methylene Blue (MB) and toluodine blue (TBO) [70], cationic fullerenes (e.g. derivatives of C60), [71,72] and psoralens (e.g. furanocoumarins) [73].
PDT is quite different in comparison with conventional drug discovery platforms, since three elements (PS, visible light and oxygen) are essential for successful deployment. One of the main challenges for the rational design of efficient PSs arise precisely from the fact that the electronically excited states (singlet, triplet and reactive intermediates) of the photoactive agent causes cell toxicity [74,75]. This prompts the use of complex, computing-demanding high level Quantum Chemistry approaches for dealing with excited states [76-81]. Such in silico modeling, in combination with photophysical experimental studies are required for rationally tuning the desired properties of the PSs including dark toxicity, molar absorption coefficient, absorption wavelength, singlet-triplet intersystem crossing quantum yield and forthcoming singlet oxygen generation and probably also the study of open sheel species (radical cations in case of phototoxicity in the absence of oxygen) [82-84]. Examples from these interdisciplinary efforts yielded chemical modifications to enhance known dyes, as well as the re-consideration of dyes which absorb outside the desirable wavelength window (600-850 nm) by means of two-photon absorption [83-85] of yellow-orange dyes by means of potent red lasers, as well as the preparation of either highly positively charged derivatives [86] or conjugates of the PS and antibodies or nanoparticles for improving their specificity and/or cell accumulation [87-89].
Many of the elements of the bacterial phenotype may play an important role as PDT applications evolve. A set of technical challenges will have to be met using sophisticated tools and approaches that address complex biological questions regarding resistance mechanisms, biofilm inactivation, and persister cell formation. There are no validated examples of biofilm photoinactivation or PDT being used in reliable polymicrobial infection models. Minimal information exists for the design of host-pathogen studies exploring the ability of PDT to interfere with virulence determinants, although there is one notable exception; a report of a host-parasite model to assess intracellular targeting specificity of novel phthalocyanines against Leishmania parasites infecting macrophages and dendritic cells [90]. However the non-specific mechanism of action and successful clinical implementation prompt and direct a growing critical mass of antimicrobial PDT explorations.
The participation of efflux systems in PS mediated PDT has been observed with the ABC mammalian transporters ABCG2 (or Breast Cancer Resistance Protein BCRP), ABCC1 (or multidrug resistance-associated protein 1, MRP1) and in a lesser extent with ABCB1 (P-glycoprotein, P-gp). Primary evidence came from a variety of PDT investigations with porphyrins (phytoporphyrin, protoporphyrin IX) [91-93], pyropheophorbides and purpurinimides [94]. Both ABG2 and ABCC1 affected the outcome of hypericin-mediated PDT in HT-29 adenocarcinoma cells [95]. In these two systems it is clear that MES affect PDT for a variety of PS chemotypes. In contrast, for ABCB1 the evidence for PS substrates (chlorine e6 and psoralen) is sporadic and contradictory [96, 97]. In one only occasion, a hypericin-mitoxantrone (MTZ) cocktail plus illumination with blue light potentiated cytotoxicity in bladder and breast cancer cells that overexpress P-gp [98].
The two phenothiazinium dyes MB and TBO are amphipathic cations and physicochemically similar to the natural product antibacterial alkaloid berberine, a well-characterized substrate of MFS efflux systems in Gram-positive bacteria [27, 99]. This has raised the possibility that phenothiazinium PSs are substrates of microbial efflux systems. Experimental evidence indicated that phenothiaziniums were NorA (MFS) substrates in S. aureus and possibly MexAB (RND) substrates in P. aeruginosa [100]. Contrarily this evidence was not supported by a model study using sixty clinical isolates of P. aeruginosa overexpressing efflux systems where it was demonstrated that antibiotic-resistant P. aeruginosa was as susceptible to TBO-mediated PDI as susceptible strains [101]. The observation that ABC transporters and not MFS transporters affect MB-mediated PDT in the pathogenic yeast Candida albicans is perplexing [100, 102]. An in silico modeling venture, employing MB and the ABCB1 crystal structure adds in this perplexity. The study reveals a strong overlap for the PS binding site with the classic efflux substrates berberine and rhodamine 6G due to similarity in shape and electronic distribution (Fig. 3). It was concluded that MB appears to be at least as well substrate of ABCB1 as berberine and rhodamines [103]. Furthermore, the structurally related phenothiazines thioridazine and chromazine have been characterized as EPIs or functional transporter regulators as opposed to substrates of a variety of pathogen efflux systems [104-106].
The well-documented promiscuity of efflux systems may explain the scattered reports for the role of porphyrins as potential substrates. Porphyrin uptake and efflux seem to be regulated by the TolC system in E coli [107]. In Streptococcus agalactiae, two co-regulated efflux transporters modulate intracellular heme and protoporphyrin IX availability [108]. In contrast the PDT pattern of amphiphilic protoporphyrin diarginate PPArg in a variety of efflux related S. aureus strains show no correlation for the PS with MES [109].
Tools for EPI Discovery
A number of assays to identify EPIs have been developed in the last 15 years. As the knowledge around efflux systems is advancing and due to their polyspecificity and overlapping roles in the microbial cell physiology, the majority of the EPI-discovery venture is an evolving “work in progress”. The activity of microbial efflux systems is not accurately detected by classical non-functional methods (Northern blotting, RNase protection, RNA in situ hybridization, RT-PCR or immunostaining). Efflux pump protein expression is often not correlated with mRNA levels, as transcripts are often present below the detection threshold, since relatively few active transporter molecules can cause major alterations in drug transport.
One key and generic distinction can be made between functional and phenotypic assays. Both groups of methodologies employ strains that lack or overexpress efflux systems and are usually robust with reproducible results, have been used extensively in low and middle throughput, and are amenable to miniaturization. Computational approaches have also been used but those efforts were not entirely independent as either an assay from the first two categories was coupled to provide proof of principal experimental information. Furthermore, the “on demand” regulation for many efflux systems in the presence of an antimicrobial agent complicates more these expeditions. One of the best understood examples at the molecular level involves the expression of TetA in response of tetracycline/Mg++ complex, mediated by the TetR transcriptional regulator [110,111] (Fig. 4).
A functional efflux assay is based on the ability of efflux systems to move compounds passively or actively against the concentration gradient, across the cell membrane and on the availability of a variety of fluorescent molecule-dyes to act as substrates of the pump under investigation. Upon loading of the cells with a lipophilic dye capable of diffusing across cell membranes, the fluorescence intensity of the cell will depend upon the activity of the efflux system. The cells, where highly active transporters are present, will display lower fluorescence intensity values because of the increased efflux of the dye/substrate. In the presence of an active EPI, these substrates will accumulate in the cell. The transporter function can be measured in cellular uptake, efflux, or steady-state distribution of fluorescent substrates over time. Two subcategories of functional transporter assays, using fluorescent dyes are commonly employed: i. the accumulation assay measures dye uptake in the presence or absence of known EPIs and ii) the retention assay where the cells are loaded with the substrate in the absence of any modulator-potential EPI, washed, and then further incubated without dye but in the presence of modulators to allow time for the substrate to be transported out of the cell by the efflux system under investigation. As substrates act differently under different experimental conditions, the distinction between accumulation and retention is essential when evaluating cells for efflux phenotypes. Both assay subcategories offer higher throughput, generic readouts (increase in fluorescence intensity), and are readily automated. As the critical mass and the variety of efflux-based expeditions is increasing, assay development and screening for EPIs have been transferred from conventional fluorimeters and plate readers to fluorescence microscopes and high-resolution multi-parametric flow cytometers.
Acridine orange, berberine, ethidium bromide (EtBr) and rhodamines (6G, and 123) have all been extensively used to report efflux activity in a wide variety of organisms including bacteria and fungi [6]. Recently, by using AcrAB-TolC efflux phenotypes Nile red was validated as a fluorescent dye substrate [112]. Variations of functional assays have been developed through automation to allow evaluation of EPIs against multiple bacterial strains. Classic examples are a) the agar-based method employing EtBr with modifications to evaluate as many as twelve bacterial strains and has been termed the EtBr-agar cartwheel method. Agar plates containing different concentrations of EtBr are swabbed with bacterial cultures [113] and b) a thermocycler based method for the real-time assessment of accumulation and efflux of EtBr under varying physiological conditions (temperature, pH, presence and absence of the energy source) as well as presence of EPIs. The method is sufficiently sensitive to characterize intrinsic efflux systems of multiple reference strains [114].
Unlike mammalian efflux systems, the capabilities of flow cytometry to contribute in microbial EPI discovery have not been extensively explored. In a recent example, an EtBr flow cytometric efflux assay has been developed to verify the EPI activity of celecoxib, (a cyclooxygenase-2 inhibitor) in S. aureus strains including MRSA and also M. smegmatis [115]. EPI activity for the RND efflux systems in E. coli and P. aeruginosa have been monitored using a microfluidic channel device under a fluorescence microscope. Fluorescein-di-β-D-galactopyranoside (FDG), a fluorogenic compound, is hydrolyzed by β-galactosidase in the cytoplasm of E. coli to produce a fluorescent dye, fluorescein. Both FDG and fluorescein were found to behave as substrates of E. coli efflux, and employed to validate the microfluidic channel device with two EPIs, the MexB-specific pyridopyrimidine (D13-9001) and non-specific Phe-Arg-β-naphthylamide (PAβN) [116].
In a few occasions uptake using radioactive labeled antibiotics have been used. Uptake of [14C]ciprofloxacin verified tariquidar and elacridar as EPIs in S. aureus and elacridar in S. maltophilia [117]. [14C]chloramphenicol was employed to identify the EPI activity of an alkylaminoquinazoline in Gram-negative efflux pump-overproducing strains [118].
The phenotypic assays are essentially growth inhibition assays with some modifications. The ultimate goal is to identify whether potential EPI compounds enhance the efficacy of antimicrobials for the interrogated efflux system(s). Growth inhibition of potential EPIs in the presence of substrates in sub-inhibitory concentration have been broadly used in low, middle and high throughput. Checkerboard or matrix assays are the most common and require both compounds (the antimicrobial and the EPI) to be diluted over a concentration range to assess through viability the nature of their interaction. The final readout set up and throughput may vary depending on the viability or imaging method used. The method is straightforward and calculates 1. The generic modulating factor (MF, fold potentiation of an antimicrobial present at sub-inhibitory concentration [119, 120] and 2. The robust Fractional Inhibitory Concentration indices (FICI, distinguishes whether two compounds together are demonstrating additive, synergism or antagonistic activity. FIC indices (FICI) are interpreted as synergic when values are ≥ 4. The results between synergy and antagonistic tendency are defined as additive or indifferent [120, 121].
An EPI targeting proton antiporters should not disrupt the proton motive force. This is clarified through protonophore counter-screen assays that employ for example the accumulation of radiolabeled lactose via the proton motive force (the membrane potential and the pH gradient) [120]. Independently of the targeting pump an EPI should be stable in potentiating the killing effect of antimicrobials over time. A microbiological time-kill assay is a final in vitro step for the functional characterization of a potential EPI [122]. In a classic example piperine effectively enhanced the bactericidal activity of rifampicin in time-kill studies and also significantly extended its post-antibiotic effect (PAE) [123].
Although the X-ray crystal structures of some transporter families have been reported [124-132] only a limited number of computational methods have been used for microbial EPI identification and validation. Emphasis has been given in MFS and methods are available for the NorA and Rv1258c in M. tuberculosis). The virtual screening procedure employed Fingerprints for Ligands and Proteins (FLAP), a new methodology based on GRID force field descriptors [133] The 3D structure of Rv1258c, using in silico modeling, was analyzed to elucidate the binding of piperine to the active site [123]. Quantitative structure activity relationship [134] has been performed in order to obtain a highly accurate model enabling prediction of inhibition of S. aureus NorA of new chemical entities from natural and synthetic sources. Algorithm based on genetic function approximation method of variable selection in Cerius2 was used to generate the model. The model is not only able to predict the activity of new compounds but also explains the important regions in the molecules in quantitative manner [135].
Examples of Quantitative Structure-Activity Relationships (QSAR) models for prediction of pump inhibitory activity have been employed. The key example involves a model designed with genetic algorithm (GA) and partial least square (PLS) analyses with a good predictive power and yielded the compound 2-(2-Azidomethyl-5-phenoxy-phenyl)-5-nitro-1H-indole. The EPI lower the MIC of berberine to 0.091 mg/L in the S. aureus NorA over expression phenotype K2361 [136].
The National Institutes of Health Molecular Libraries Probe Production Centers Network (NIH MLPCN) is tasked with finding small molecule probe compounds for academic facilities and investigators lacking adequate HTS tools to exploit biological systems. The University of New Mexico Center for Molecular Discovery (UNMCMD) has pioneered the development of cell suspension HTS transporter inhibitor discovery assays utilizing a sensitive multiplex flow cytometry platform [137]. The approach incorporates profiling libraries such as the Prestwick Chemical Library (PCL) which consists of 1200 off-patent and known biologically active compounds along with the diverse Molecular Libraries Small Molecule Repository (MLSMR, http://mlsmr.glpg.com/ML-SMR_HomePage/) library of greater than 350K compounds. We set out to develop new small molecule scaffolds with distinct efflux inhibition selectivity profiles based on multiplex efflux system target screening. It paved the way for a series of innovations in chemical genetics including novel flow cytometry efflux assays in both mammalian (ABCB1, ABCB6, ABCG2, ABCC1) and yeast transporters including ABC (CDR1, CDR2 in Candida albicans, V-ATPase in Sacharomyces cerevisae) and MFS (MDR1) [138-141].
An HTS campaign was employed based on Nile red or R6G as the fluorescent probes and a phenotypic multiplex (triplex) of heterologous expressed C. albicans transporters in a S. cerevisiae system where endogenous efflux systems have been disabled. The PCL HTS campaign led to confirmation of enniatin B as a CDR1 inhibitor and revealed that the monoamine oxidase A inhibitor clorgyline is a broad-spectrum inhibitor of fungal ABC and MFS efflux pump activities which reverses the azole resistance of C. albicans and Candida glabrata. In a slightly different approach, a target-based functional flow cytometry assay that measures vacuolar pH-dependent fluorescence changes in response to V-ATPase function was developed. The experimental strategy uses the pH-sensitive fluorophore BCECF-AM specific for the yeast vacuole and a potent and highly specific V-ATPase inhibitor, concanamycin A. The assay was also established by using pH Luorin, a radiometric pH-sensitive GFP derivative that is used to monitor cytosolic pH.
This approach has been orchestrated as a center driven initiative with ultimate goal to set the foundation for the Transporter-Ligand Interactome. The foundation of the interactome is a UNMCMD predictive tool. This tool is comprised of a database and a visualization component that incorporates data from HTS flow cytometry campaigns with genomics, proteomics, structural informatics and knowledge mining using yeast and mammalian model systems for chemical probe discovery.
EPI Combined with Antibiotics
The NorA system in S. aureus has been the most prominent discovery target. An array of synthetic efforts has identified a number of chemotypes but there is wealth of structures and information for lead EPIs of natural origin (Table 1). There are numerous antibiotics categorized as first and second line countermeasures to treat TB. According to the WHO report more than 110,000 cases of deaths occur every year and, ~490,000 cases of MDR to first line TB-drugs, and 40,000 cases of extensively drug resistant (XDR) to first and second line TB-drugs emerge every year [142]. Although resistance to TB is multifactorial, the importance of efflux has been identified and documented in a variety of occasions [51, 143]. A number of synthetic EPI scaffolds have been validated in combination with antibiotics in vitro. Several NPs mostly flavonoids exhibit EPI activity against resistant Mycobacteria strains (Table 2) in a variety of efflux assays but there is no substantial evidence either for synergistic activity with antibiotics or specificity in respect with classes of efflux systems or inhibition patterns.
Representative NPs with EPI-activity Against Gram-Positive Bacteria
| S. aureus EPIs | Synergist | Isolated From | EPI [C] | References |
|---|---|---|---|---|
| Polyphenols | ||||
| 4,5 caffeoylquinic acid | fluoroquinolones | Artemisia absinthum | 10mg/L | [188] |
| 2-Arylbenzofuran | ||||
| spinosan A | Berberine | Dalea spinosa | 48 µM | [189] |
| pterocarpan | - | - | 56 µM | - |
| N-Caffeoylphenalkylamides | ||||
| N-trans-feruloyl 4’-O-methyldopamine | Norfloxacin | Mirabilis jalapa | 100 mg/L | [190] |
| Chalcones | ||||
| chalcone | Berberine | D. versicolor | 10 mg/L | [191] |
| Coumarins | ||||
| 4-{[(E)-5-(3,3-dimethyl-2-oxiranyl)-3-methyl-2-pentenyl]oxy}-7H-furo[3,2-g]chromen-7-one | Norfloxacin | grape fruit oil | 35.7 mg/L | [192] |
| 7-{[(E)-5-(3,3-dimethyl-2-oxiranyl)-3-methyl-2-pentenyl]oxy}-2H-2-chromenone | - | - | 30 mg/L | [192] |
| Galbanic acid | Ciprofloxacin | Ferula szowitsiana | 300 mg/L | [193] |
| - | Ethidium Bromide | - | 0.5 mg/L | - |
| Flavones | ||||
| baicalein 5, 6, 7-trihydroxyflavone | Tetracycline | Thymus vulgaris L | n/a | [194] |
| Flavonols | ||||
| chrysosplenol-D | Berberine | Artemisia annua | 25 mg/L | [195] |
| chrysoplenetin | - | - | 6.25 mg/L | - |
| Tiliroside | Ciprofloxacin | Herissantia tiubae | 32 mg/L | [196] |
| Flavonolignans | ||||
| 5’-methoxyhydnocarpin-D | Norfloxacin | Berberis aetnensis | 10 mg/L | [197] |
| Isoflavones | ||||
| genistein | Norfloxacin | Lupinus argenteus | 10 mg/L | [198] |
| orobol | - | - | - | - |
| biochanin A | - | - | - | - |
| Homoisoflavonoids | ||||
| bonducellin | Berberine | Caesalpinia digyna | unpublished data | |
| 8-methoxybonducellin | - | - | - | |
| Phenylpropanoids | ||||
| acetoxycavicolacetate | Ethidium Bromide | Alpinia galanga | 50 mg/L | unpublished data |
| Tannins | ||||
| epicatechin gallate | Norfloxacin | 20 mg/L | [199] | |
| epigallocatechin gallate | - | - | - | |
| epigallocatechin gallate | Tetracycline | Green tea | 0,0625-0.125mg /L | [200] |
| Diterpenes | ||||
| Carnosic acid | Erythromycin | Rosmarinus officinalis | 10mg/L | [201] |
| carnosol | Erythromycin | R. officinalis | 10mg/L | [201] |
| (+) Totarol | Ethidium bromide | Chamaecyparis nootkatensis | 5 mg/L | [202] |
| isopimarane diterpenes: methyl-1alpha-acetoxy-7alpha 14alpha-dihydroxy-8,15-isopimaradien-18-oate and methyl-1alpha,14alpha-diacetoxy-7alpha-hydroxy-8,15-isopimaradien-18-oate | Tetracycline | Lycopus europaeus | [203] | |
| ferruginol | Norfloxacin, Oxacillin | C. lawsoniana | 2 mg/L | [204] |
| Triterpenoids | ||||
| oleanolic acid | C. edulis | 10mg/L | [205] | |
| ulvaol | C. edulis | 10mg/L | - | |
| Oligosaccharides-Glycosides | ||||
| murucoidins XII-XVI | Norfloxacin | Ipomoea murucoides | 5-25 mg/L | [206] |
| orizabin XIX -IX-XV | Norfloxacin | Mexican Morning Glory Spp. | 1-25 mg/L | [207] |
| polyacetylated neohesperidosides | Berberine/fluroquinolones | Geranium caespitosum | 10 mg/L | [208] |
| Resin glycosides (murucoidins, pescaprein and stoloniferin) | fluoroquinolones | I. murucoides | [209] | |
| kaempferol-3-O-beta-d-(6''-E-p-coumaroyl) glucopyranoside (tiliroside) | Corfloxacin, Ciprofloxacin, Lomefloxacin, Ofloxacin | Herissantia tiubae | 10 mg/L | [196] |
| piperine | Ciprofloxacin | Piper nigrum | 50 mg/L | [210] |
| 2,6-Dimethyl-4-phenylpyridine-3,5-dicarboxylic acid diethyl ester | - | Jatropha elliptica | 2 mg/L | [211] |
| reserpine | Tetracycline, Norfloxacin | Rauwolfia vomitoria | [19] | |
| harmaline | Ethidium Bromide | Peganum harmala | [212] | |
| ergotamine | Norfloxacin | Claviceps purpurea | [19] | |
| pheophorbide a | Ciprofloxacin | Berberis aetnensis | 0.5 mg/L | [213] |
| julifloridine, juliflorine and juliprosine | Norfloxacin | Prosopis juliflora | [214] | |
| indoles, indirubicin | Ciprofloxacin | Wrightia tinctoria R. Br. | 12.5-25 mg/L | [215] |
| Pyridines | ||||
| 2,6-dimethyl-4-phenyl-pyridine-3,5-dicarboxylic acid diethyl ester | fluoroquinolones | Jatropha elliptica | [192, 211] | |
Representative NPs with Mycobacterial EPI-Activity
| Mycobacterial EPIs | Synergist | Microorganism(s) | EPI [C] | References |
|---|---|---|---|---|
| Plant Flavonoids | ||||
| (-)-epicatechin | Isoniazid | Mycobacteria spp. | 32 mg/L | [216] |
| kaempferol | - | - | - | - |
| isorhamnetin | - | - | - | - |
| taxifolin | - | - | - | - |
| rutin | - | - | 16 mg/L | - |
| myricetin | - | - | - | - |
| quercetin | - | - | - | - |
| resveratrol | Ethidium Bromide | M. smegmatis (MC2 155) | - | [217] |
| genistein | - | - | - | - |
| baicalein | - | - | 10 mg/L | - |
| biochanin A | - | - | 32 mg/L | - |
| Totarol | Isoniazid | M. tuberculosis (H37Rv) | [218] | |
| ferruginol | - | - | ||
| sandaracopimeric acid | - | - | ||
| 4-epiabetol | - | - | ||
| plumbagin | - | - | ||
| farnesol | Ethidium Bromide | M. smegmatis (MC2 155) | 32 mg/L | [219] |
| Curcumin | Isoniazid | - | 32 mg/L | |
| demethoxycurcumin | - | - | - | [220] |
Representative NPs with EPI-Activity against Gram-Negative Bacteria (EPI [C] is Ranging between 10-50mg/L)
| RND-EPIS | Synergist | Plant Source | References |
|---|---|---|---|
| Alkaloids | |||
| Cathinone | Ciprofloxacin | Catha edulis | [221] |
| Theobromine | Ciprofloxacin,Tetracyclin, Chloramphenicol,Ethidium bromide | Theobroma (cacao tree) | - |
| Epinephrine,catecholamine | Ciprofloxacin, Tetracyclin,Erythromycin, Chloramphenicol,Ethidium bromide | - | - |
| Norepinephrine, catecholamine | Ciprofloxacin, Tetracyclin,Erythromycin | - | - |
| Theophylline, methylxanthine | Ciprofloxacin | - | - |
| Caffeine | Ciprofloxacin | - | |
| EA-371alpha, EA-371delta | Levofloxacin | Streptomyces spp. | [157] |
| 5'-methoxyhydnocarpin-D (5'-MHC-D), pheophorbide a | CIprofloxacin | Berberis aetnensis | [213] |
| Extracts of Commiphora molmol, Centella asiatica, Daucus carota, Citrus aurantium and Glycyrrhiza glabra | Chloramphenicol,Nalidixic acid,Tetracyclines | Commiphora molmol, Centella asiatica, Daucus carota, Citrus aurantium and Glycyrrhiza glabra | [214] |
| Thanatin,peptide | Podisus maculiventris | [222] | |
| Ellagic tannic acids | Novobiocin, coumermycin, chlorobiocin, rifampicin, fusidic acid | n/a | [223] |
The pan resistance ability of Gram-negatives triggered a number of sophisticated EPI discovery strategies. The number of chemotypes is substantially smaller in comparison with Gram-positives but the number of available studies employing the core of EPIs is comprehensive. Phenyl-arginine-beta-naphthylamide (PaβN) was the first EPI identified in P. aeruginosa by assaying an array of synthetic compounds and natural products using strains over-expressing each of the three MES (MexAB-OprM, MexCD-OprJ, MexEF-OprN) in the presence of levofloxacin. It has been a valuable tool for drug discovery [144]. This approach has explored: i) the development of preclinical candidates including strategies for lead optimization; ii) activity in vivo through alternative scaffolds; iii) optimization of potency in the pyridopyrimidine series through the application of a pharmacophore model; and iv) extensive structural activity relationships in toxicity, stability and solubility. Mechanistically, it has been proposed that PAβN itself is an RND competitive substrate. It seems that PAβN may recognize and bind to the substrate pocket specific to the potentiated antibiotics. Alternatively, due to a proximal binding site, the EPI may also generate steric hindrance, impairing the antibiotic binding at its affinity site. PAβN has been validated against the AcrAB-TolC in a variety of Gram-negative pathogens (K. pneumoniae, E. coli, S. typhimuriumE. aerogenes) and multiple homologous systems (A. baumannii,Campylobacter sp.). Naphthylpiperazines, notably 1-naphthylmethyl-piperazine (NMP) are among the most potent unsubstituted arylpiperazines, with a dose-dependent ability to increase the intracellular concentration of several antibiotics [145]. NMP seems to be effective in A. baumannii and several Enterobacteriaceae [145,146]. The list also includes trimethoprim and epinephrine, indole derivatives and quinolone derivatives [147]. Recently, artesunate was found to enhance the antibacterial effect of beta-lactam antibiotics against E. coli by increasing antibiotic accumulation via inhibition of AcrAB-TolC [148]. A number of NPs have been implicated in Gram-negative efflux inhibition (Table 3). Again the list is chemotypes is limited and the mechanistic implications unexploited. As a general rule, it is challenging to identify NPs with Gram-negative EPI properties with the available functional or phenotypic efflux assays [149, 150]. In a recent study, total alkaloids of Sophora alopecuroides-induced down-regulation of AcrAB-TolC and reverse susceptibility to ciprofloxacin in MDR E. coli isolates [151].
There is a core of natural products that can enhance the potency of antifungals: 1. Tacrolimus isolated from the fermentation broth of Streptomyces tsukubaensis and when added in sub-inhibitory concentrations (10-5 to 10-3 mM) can reduce 16-fold the MIC of itricanazole in the CDR1-expressing resistant strain C. albicans and 8–fold in a CaMDR-expressing resistant strain [152]. 2. The cyclodepsipeptides unnaramicin A and unnaramicin C isolated from the extracts of marine bacterium Photobacterium sp reduce the MIC of fluconazole by 64-fold (320 mg/L to 5 mg/L) against azole-resistant C. albicans at relatively low (1.25µM-5µM) concentration [153,154]. 3. Geodisterol-3-O-sulfite and 29-demethylgeodisterol-3-O-sulfite isolated from marine sponge Topsentia sp. enhanced the activity of fluconazole against overexpressed MDR1 in clinical isolate of C. albicans [155]. 4. Enniatin B reduced the IC50 of cyclohexamide 8-fold (0.13 to 0.016 µg/mL) against a Pdr5p-overexpressing strain of Saccharomyces cerevisiae at 6.0 µM [156]. 5. Milbemycins α9 found to enhance the activity of fluconazole and the triazole SCH-56592 against clinical isolates of C. albicans [157].
EPI Combined with PDT
The concept of synergistic action of antimicrobials with EPIs has been exploited in PDT to potentiate the phototoxic action of phenothiazinium PSs [158]. The PDT effect of MB or TBO was substantially enhanced by small molecule EPIs in S. aureus that affect NorA as assessed by both reduction of viable cells and fluorescent dye accumulation. The potentiation is less pronounced against P. aeruginosa with Mex AB [158].
Prates et al. tested this hypothesis by comparing MB- PDI for two pairs of isogenic C. albicans strains i) YEM 12 and YEM 13 mutant overexpressing MDR1 MFS; and ii) YEM 14 and YEM 15 mutant overexpressing CDR1/CDR2. Since efflux systems affect APDI of MB in C. albicans, a logical strategy was to explore whether EPIs could potentiate antimicrobial PDI (APDI) with MB. They used two well-documented EPIs, one for each corresponding efflux system, INF271 (targeting MFS) and verapamil (+) (targeting ABCs both in yeast and mammalian systems) and the reference C. albicans strain DAY185. Pre-exposure to INF271 followed by MB mediated PDI, resulted in a paradoxical protection rather than enhancement of phototoxicity. On the other hand, pre exposure of C. albicans cells to verapamil (+) and subsequent MB-mediated APDI led to an increase of phototoxicity when compared with the PDI efficacy of MB in the absence of EPIs. In addition, pre exposure of the CDR1/CDR2 overexpressing mutant YEM 15 in verapamil (+) and subsequent APDI revealed 2 logs10 of cell reduction in the presence of the EPI, in comparison with virtually no effect in the absence of the ABC pump blocker. The authors demonstrated that ABC pumps are directly implicated in MB efflux from the cell cytoplasm.
Animal Models of Infection, and the Challenge of Clinical EPI Implementation
A major obstacle to the path of microbial EPI implementation may arise from the fact that efflux systems manipulation could cause unexpected toxicities due to the multitude of physiological roles MES may play in human cells. In this context, efforts directed at specifically inhibiting efflux pumps operating only in prokaryotes may offer a greater chance of therapeutic success. Interestingly, it has been shown that target bacteria respond to clinical challenge with EPIs by developing resistance mutations that decrease the efficacy of the EPI [159,160]. In the same context, it was demonstrated that reserpine can select multidrug resistant Streptococcus pneumoniae strains [161].
The threat of cross-resistance to different antibiotics elevates the complexity of EPI discovery ventures. Addressing this requires that efflux substrates and inhibitors be clearly differentiated, this may be also of interest for the PS for use in antimicrobial PDT. The latter case will demand rational approaches that simultaneously address both the photochemical mechanistic aspects of PDT and efflux phenotypic variations. A reported strategy of “dual antimicrobial action” targeting NorA in Gram-positive bacteria provides a path forward towards addressing these issues. A hybrid compound (SS14) was created by fusing the plant antimicrobial berberine to the synthetic NorA EPI INF55. SS14 was established as an effective antimicrobial against S. aureus, (100-fold more effective than the berberine parent molecule) including mutant strains that over express NorA [162]. MIC’s for SS14 against S. aureus are interestingly 2-16 times lower than berberine in combination with the EPI INF55 when present together. The hybrid rapidly accumulated in bacterial cells and showed higher efficacy than vancomycin in a Caenorhabditis elegans model of enterococcal infection [162]. Analogs of SS14 exhibited similar antimicrobial activities [163,164] suggesting that significant structural changes can be made to these hybrids without adversely affecting their ability to block MES or their antibacterial activity. Further compelling the cytotoxicity associated with INF55 was found absent in the hybrid SS14. Such hybrids are predicted to have an advantage over separate compound administration in terms of synchronous or near synchronous delivery of both agents to the appropriate bacterial target sites. Interestingly –conjugation of the indole derivatives to ciprofloxacin rendered the hybrid orders of magnitude less effective than ciprofloxacin alone (Anthony Ball unpublished data). Perhaps the lesson is antibiotics are not ideal candidates for hybrid strategies; but rather antimicrobials; or agents with a non-specific mechanism of action are perhaps the better suited for conjugate approaches. It is noteworthy that plants evolved a complex milieu of toxins and synergists which are synthesized in defense against microbial attack –and that such a technique differs largely from that strategy employed by antibiotic producing microbes.
Among the potential roles, it has been demonstrated that efflux pumps are important for processes of detoxification of intracellular metabolites, bacterial virulence in both animal and plant hosts, cell homeostasis and intercellular signal trafficking [165]. Researchers have engaged in new antimicrobial development tractable whole-animal screens that utilize the well-studied nematode C. elegans, the great wax moth Galleria mellonella, the fruit fly Drosophilla melanogaster and the zebra fish. Those have been employed as model hosts to identify and develop new classes of antimicrobial agents with antivirulence or immunomodulatory efficacy and evaluate toxicity or efficacy. The amenability of these non-vertebrate hosts to large screens has made them invaluable in drug discovery against either bacterial or fungal pathogens [166-168]. The design of host-pathogen studies exploring the ability of efflux to interfere with virulence determinants sounds promising and highly informative but often requires more sophisticated tools and approaches as the results varies. For example, C. elegans was used to assess the fitness of in vitro selected P. aeruginosa MexABOprM (nalB) and MexCDOprJ (nfxB) multidrug resistant mutants [169] and to confirm that overproduction of MexEF-OprN does not impair P. aeruginosa fitness in competition tests, but produces specific changes in bacterial regulatory networks [170]. B. pseudomallei is able to cause ‘disease-like’ symptoms and kill the nematode but all the indication are that the killing mechanism is not related with efflux systems that pump out either aminoglycosides or macrolides [171]. In a similar fashion C. elegans and an array of antimicrobials were valuable tools in correlating membrane efflux and influx with both multidrug resistance and virulence of K. pneumoniae [172].
The nematode model was employed to validate the antimicrobial potential of the dual action antimicrobial hybrids in a variety of pathogens [162, 164] but also to demonstrate that the quorum sensing inhibitor C-30 (brominated furanone) can trigger rapidly efflux mutations in P. aeruginosa [173]. In a recent example employing C. elegans it was demonstrated RamA, a member of the AraC/XylS family, influences both virulence and efflux in S. enterica serovar Typhimurium [174]. In a unique example the nematode model was pivotal to dissect the mechanism that Pdr1p family members in S. cerevisiae and C. glabrata use and directly bind to structurally diverse drugs and xenobiotics, resulting in stimulated expression of drug efflux pumps [175].
The reports that correlate the rest of the model hosts with efflux are not extensive. G. melonella was employed to validate the efficacy of azithromycin against intracellular infections of Francisella [176]. AcrAB-TolC mutants were used only in vitro and were not employed to infect the caterpillars. D. melanogaster was a valuable tool to display the virulence characteristics and resistance patterns of the Liverpool epidemic strain (LES) of P. aeruginosa, a transmissible aggressive pathogen of cystic fibrosis (CF) patients [177]. Mycobacterium marinum-infected zebrafish larvae for in vivo characterization of antitubercular drug activity and tolerance. It was demonstrated that efflux systems required for intracellular growth mediate this macrophage-induced tolerance. This tolerant population also developed when M. tuberculosis infects cultured macrophages, suggesting that it contributed to the burden of drug tolerance in human tuberculosis. Verapamil was shown to reduce that tolerance [178].
The results for EPI efficacy in murine models are limited and inconclusive. The combination of piperine with mupirocin in a mice dermal infection model showed better in vivo efficacy when compared with the commercially available formulation of 2 % mupirocin [179]. Using a mouse model of lethal infection, it was demonstrated that concomitant administration of the lactoferricin-derived peptide P2-15 and erythromycin against P. aeruginosa afforded long-lasting protection to one-third of the animals [180]. A d-octapeptide EPI acts synergistically with azoles in a murine oral candidiasis infection model [181]. In a latest in vivo study, PDT employing EPIs was able to reduce bacterial load in P. aeruginosa burn wounds, to delay bacteremia and to keep the bacterial levels in blood 2-3 log10 lower compared to an untreated group [182].
A number of fitness and in vivo modeling studies have been conducted using different mice models. In MRSA MW2 (USA400 lineage) for example, NorB facilitated bacterial survival upon overexpression in a staphylococcal abscess and may contribute to the relative resistance of abscesses to antimicrobial therapy, thus linking bacterial fitness and resistance in vivo [183]. An intraperitoneal mouse model of systemic infection was used to evaluate the role of AcrAB-TolC on antimicrobial resistance in vivo employing clinical isolates of Enterobacter cloacae overexpressing or lacking parts of the RND efflux system. This approach, revealed reduced virulence in both E. cloacae clinical strains when either the acrA or tolC genes were inactivated [184]. Similar studies with emphasis in the AcrAB-TolC system have been conducted in a variety of F. tularensis strains including the live vaccine strain (LVS) [185, 186] and SCHU S4 [187].
CONCLUSION
The lengthy description of this array of endeavors leads in four main conclusions: a. Tools and technological platforms for EPI discovery are available and rapidly advancing with emphasis on flow cytometry and imaging. b. A core of NP chemotypes has been validated as EPIs in many pathogenic systems but the real challenge of discovery remains Gram-negatives. c. PDT and other alternative antimicrobial strategies may provide a niche for EPI implementation. This statement requires further elaboration: Efflux systems have been evolutionary co-ordinate to pump out a broad range of non-specific compounds including potential toxins which do not attack a single target. Indeed many efflux substrates are generally active against microbial membranes and DNA neither of which is essentially mutable. Antimicrobial PS, like amphipathic cations represent old and familiar enemies to the microbial efflux systems and APDI potentiation with EPIs NPs seems like a broader scheme with multi fold applications. NP EPIs have shown remarkable efficacy in potentiating largely ineffective antimicrobials such as berberine. PDT is able to exhibit substantially higher cidal efficacy than the traditional plant antimicrobials. This concept may apply by incorporating PS and PDT-based applications to the current hybrid models which employ traditional antibiotics (evolution has already refined to reasonably evade efflux systems with their neutral moieties) or plant based antimicrobials (they are only marginally effective and only within the presence of EPIs; or multiple EPIs within the plant’s natural environment). d. An apparent methodological gap between in vitro and preclinical studies remains and host-pathogen interactive expeditions incorporating EPIs are not providing conclusive results. Although the progress in highlighting the role that MES play in microbial ecology survival and virulence seems to be slow but satisfactory, the hurdles and challenges associated with antimicrobial drug EPI-based design remain overwhelming.
CONFLICTS OF INTEREST
The authors confirm that this article content has no conflicts of interest.
ACKNOWLEDGEMENTS
George P Tegos is supported by the NIH (grant 5U54MH084690-02). Research conducted in the Hamblin Laboratory was supported by NIH (R01 AI050875 to MRH) and US Air Force MFEL Program (FA9550-04-1-0079).


