All published articles of this journal are available on ScienceDirect.
Polymorphic Amplified Typing Sequences (PATS) Strain Typing System Accurately Discriminates a Set of Temporally and Spatially Disparate Escherichia coli O157 Isolates Associated with Human Infection
Abstract
Polymorphic Amplified Typing Sequences (PATS) is a PCR-based Escherichia coli O157 (O157) strain typing system. Here, we show that PATS compares excellently with Pulsed-Field Gel Electrophoresis (PFGE) in that both methods cluster geographically diverse O157 isolates similarly. Comparative analysis of the results obtained in this simulated “blind” study attests to the discriminating power and applicability of PATS to epidemiological/nosocomial situations.
INTRODUCTION
Escherichia coli O157:H7(O157) isolates are commonly typed by pulsed-field gel electrophoresis (PFGE), following digestion of genomic DNA with the restriction enzyme XbaI. The choice of this infrequently cutting restriction enzyme is dictated by the fact that it is an effective cutter that consistently yields a decipherable profile [1-3]. However, inherent limitations of the standard PFGE protocol have spurred several modifications, including: (i) digestion of genomic DNA with additional restriction enzymes to better relate strains; (ii) development of uniform size standards and protocols for universal use; and (iii) institution of protocols to decrease sample turn aroundtime and decrease technical errors. Despite these modifications and use of pattern-recognition computer software, resolution of co-migrating and undigested DNA bands poses a significant interpretive challenge [2-8].
Polymorphic Amplified Typing Sequences (PATS) was developed based on genetic events underlying the polymorphisms observed at the XbaI sites, and another infrequent restriction enzyme site, AvrII (BlnI), in the O157 genome [9, 10]. Primer pairs targeting 8 polymorphic XbaI sites associated with insertion-deletions (Indels) and 7 polymorphic AvrII sites associated with Indels or single nucleotide polymorphisms (SNPs) were incorporated, along with those amplifying four virulence genes (stx1, stx2, eae, hlyA), to develop the PATS O157 strain typing system [9-11]. PATS was shown to accurately discriminate between unrelated isolates and relate closely linked isolates when evaluated against 46 O157 isolates associated with human disease [9-11], as well as 25 bovine O157 isolates [12]. To further confirm the real world applicability of PATS, including epidemiological situations that demand strain typing of unknown isolates, we devised a “blind study” wherein the details of the O157 isolates being typed were withheld from the researcher until after analysis. Such a study, we reasoned, would minimize subjective biases. The results of PATS analyses were compared with those obtained using PFGE for the same isolates.
MATERIALS AND METHODS
Bacterial Strains
A total of 48 O157 isolates that were associated with human disease and isolated from diverse sources/geographic locations were obtained from collections maintained at the William A. Hinton State Laboratory Institute, Jamaica Plain, MA. Details about the strains/isolates were provided only at the end of the study, and can be observed on the two dendograms based on PATS and PFGE profiles (Figs 1 and 2). The isolates were labeled MLVA 001 to 048.
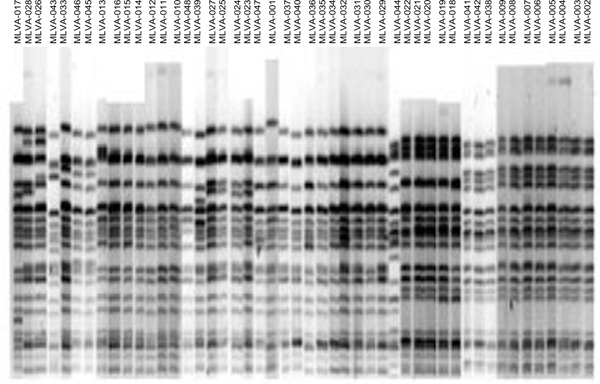
XbaI-based PFGE patterns of the 48 O157 isolates.

Analysis of the relatedness between 48 O157 isolates using XbaI-based PFGE. Dendogram was constructed using the UPGMA cluster analysis. PFGE gels were analyzed using Molecular Analyst Fingerprinting Plus software (Bio-Rad). Clades are labeled with roman numerals and groups with alphabets, A-D.
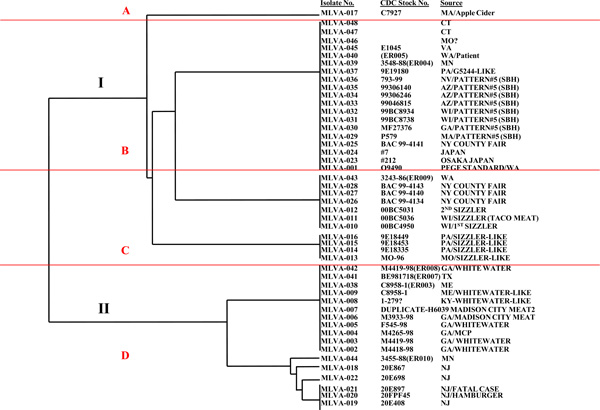
Analysis of relatedness between 48 O157 isolates using XbaI-based PATS. Dendogram was constructed using the UPGMA option in the phylogenetic analysis using parsimony (PAUP; Sinauer Associates, Inc., Publishers, Sunderland, Ma.). Clades are labeled with roman numerals and groups with alphabets, A-D, corresponding with those seen in the PFGE dendogram (Fig. 2).
PFGE
Standard PFGE method was used to analyze the 48 O157 isolates as previously described [4, 13]. Briefly, the genomic DNA of each isolate was embedded in separate agarose plugs and digested at 37°C for 2 h with 30 U of XbaI (Gibco BRL, Grand Island, N.Y.) per plug. The plugs were loaded onto a 1% agarose–tris buffer gel (SeaKem GoldAgarose; BioWhittaker Molecular Applications, Rockland, Maine), and PFGE was performed with a CHEF Mapper XA apparatus (Bio-Rad Laboratories, Hercules, Calif.). DNA was electrophoresed for 18 h at a constant voltage of 200V (6 V/cm), with a pulse time of 2.2 to 54.2 s, an electric field angle of 120o, anda temperature of 14°C, before being stained with ethidium bromide. PFGE gels were analyzed using Molecular Analyst Fingerprinting Plus software (Bio-Rad). The dendogram for the XbaI-based PFGE profiles was constructed using the UPGMA cluster analysis.
PATS
For PATS, a colony lysate of each O157 isolate was subjected to hot start, touchdown PCR with each individual primer pair as described previously [11]. PCR was carried out on the GeneAmp PCR system 2400 thermal cycler (PE Biosystems, Foster City, CA.), using 10 μl of colony lysate, 200 pmoles of each primer, 800 μM dNTPs, 1x diluted Ex TaqTM enzyme buffer and 2.5 units of TaKaRa Ex TaqTM DNA polymerase. An amplification segment of 20 cycles was set where the annealing temperature started at 73oC, to touchdown at 53oC at the end of those cycles. Then, another amplification segment of 10 cycles was set, using the last annealing temperature of 53oC. Each reaction was done in triplicate. PCR reactions were initially screened for the presence or absence of amplicons. Amplicons, when present, were purified using the QIAquick PCR purification kit (Qiagen, Valencia, Ca.), and digested with the AvrII restriction enzyme (New England Biolabs, Beverly, Ma.) to confirm the presence of an AvrII restriction site within the amplicon. Undigested and digested DNA fragments were resolved on a 4% agarose gel prepared with a combination of 3% Nusieve GTG agarose (FMC BioProducts, Rockland, Me.) and 1% agarose (Shelton Scientific Inc., Shelton, Ct.) and stained with ethidium bromide. The presence/absence of, and variations in the amplicons was tabulated as before for XbaI-based, AvrII-based,and combined-PATS; combined-PATS being XbaI-, AvrII- and virulence genes-based (Tables 1A and 2) [12]. Dendogram for the PATS data was constructed using the unweighted pair-group method with arithmetic means (UPGMA) option in phylogenetic analysis using parsimony software (PAUP; Sinauer Associates, Inc., Publishers, Sunderland, Ma.) as described previously [9-11].
XbaI-based PATS Profiles of the 48 O157 Isolates
| PATS typea | PCR Amplification and Restriction Digestion Patterns of Ampliconsobtained Using 8 PATS Primer Pairsb | Isolatesc | |||||||
|---|---|---|---|---|---|---|---|---|---|
| IK8 | IK19 | IK25 | IK114 | IK118 | IK123 | IKB3 | IKB5 | ||
| 1 | 2 | 2 | 0 | 2 | 2 | 2 | 0 | 0 | MLVA 002, 003, 004, 005, 006, 007, 008, 009, 038, 041, 042 |
| 2 | 2 | 2 | 0 | 2 | 2 | 2 | 0 | 2 | MLVA 001, 023, 024, 025, 029, 030, 031, 032, 033, 034, 035, 036, 037, 039, 040, 045, 046, 047, 048 |
| 3 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 2 | MLVA 010, 011, 012, 026, 027, 028, 043 |
| 4 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 2 | MLVA 013, 014, 015, 016 |
| 5 | 2 | 2 | 0 | 2 | 2 | 0 | 2 | 0 | MLVA 018 |
| 6 | 0 | 2 | 0 | 2 | 2 | 0 | 0 | 2 | MLVA 017 |
| 7 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 0 | MLVA 019, 020, 021 |
| 8 | 2 | 0 | 0 | 2 | 2 | 2 | 2 | 0 | MLVA 022 |
| 9 | 2 | 2 | 0 | 2 | 0 | 2 | 0 | 0 | MLVA 044 |
AvrII-based PATS Profiles of the 48 O157 Isolates
| PATS type | PCR Amplification and Restriction Digestion Patterns of Ampliconsobtained Using 7 PATS Primer Pairsd | Isolates | ||||||
|---|---|---|---|---|---|---|---|---|
| IKNR3 | IKNR7 | IKNR10 | IKNR12 | IKNR16 | IKNR27 | IKNR33 | ||
| 1 | 2 | 1 | 1 | 2 | 1 | 2 | 2 | MLVA 002, 003, 004, 005, 006, 007, 008, 009, 038, 041, 042 |
| 2 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | MLVA 018, 019, 020, 021, 022 |
| 3 | 2 | 2 | 2 | 2 | 2 | 3 | 2 | MLVA 017 |
| 4 | 1 | 2 | 2 | 2 | 1 | 2 | 2 | MLVA 044 |
| 5 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | MLVA 001, 010, 011, 012, 013, 014, 015, 016, 023, 024, 025,026, 027, 028, 029, 030, 031, 032, 033, 034, 035, 036, 037, 039, 040, 043, 045, 046, 047, 048 |
a PATS types are designated arbitrarily with different numbers.
b Prefixes of each PATS primer pair A/B are indicated. 0, no amplicon; 2, amplicon with one XbaI site.
c E. coli O157:H7 isolates from different locations that fell within a given PATS type.
d Prefixes of each PATS primer pair A/B are indicated. 0, no amplicon; 1, amplicon without AvrII site; 2, amplicon with one AvrII site; 3, amplicon with an additional AvrII site due to a SNP.
PATS Profiles of 48 O157 Isolates based on Polymorphic XbaI and AvrII Restriction Sites, and Virulence Genes
| PATS Typea | PCR Amplification and Restriction Digestion Patterns of Ampliconsobtained Using 15 PATS - 4 Virulence Gene Primer Pairsb | Isolatesc | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Polymorphic XbaI Sites | Polymorphic AvrII sites | Virulence Genes | ||||||||||||||||||||
| IK8 | IK19 | IK25 | IK114 | IK118 | IK123 | IKB3 | IKB5 | - | IKNR3 | IKNR7 | IKNR10 | IKNR12 | IKNR16 | IKNR27 | IKNR33 | - | stx1 | stx2 | eae | hlyA | ||
| 1 (prev. 8)d | 2 | 2 | 0 | 2 | 2 | 2 | 0 | 0 | 2 | 1 | 1 | 2 | 1 | 2 | 2 | 0 | 1 | 1 | 1 | MLVA 002, 003, 004, 005, 006, 007, 008, 009, 038, 041, 042 | ||
| 2 (prev.18) | 2 | 2 | 0 | 2 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | MLVA 001, 023, 024, 025, 029, 030, 031, 032, 033, 034, 035, 036, 037, 039, 040, 045, 046, 047, 048 | ||
| 3 (prev.19) | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | MLVA 010, 011, 012, 026, 027, 028, 043 | ||
| 4 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 1 | 1 | 1 | 1 | MLVA 013, 014, 015, 016 | ||
| 5 | 2 | 2 | 0 | 2 | 2 | 0 | 2 | 0 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 1 | 1 | 1 | MLVA 018 | ||
| 6 (prev.11) | 0 | 2 | 0 | 2 | 2 | 0 | 0 | 2 | 2 | 2 | 2 | 2 | 2 | 3 | 2 | 1 | 1 | 1 | 1 | MLVA 017 | ||
| 7 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 1 | 1 | 1 | MLVA 019, 020, 021 | ||
| 8 | 2 | 0 | 0 | 2 | 2 | 2 | 2 | 0 | 2 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 1 | 1 | 1 | MLVA 022 | ||
| 9 (prev. 9) | 2 | 2 | 0 | 2 | 0 | 2 | 0 | 0 | 1 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 1 | 1 | 1 | MLVA 044 | ||
a PATS types are designated arbitrarily with different numbers.
b Prefixes of each PATS primer pair A/B and virulence gene primer pair F/R are indicated. 0, no amplicon; 1, amplicon without XbaI or AvrII site; 2, amplicon with one XbaI or AvrII site; 3, amplicon with an additional AvrII site due to a SNP
c Isolates of E. coli O157:H7 from various outbreaks that fell within a given PATS type.
d PATS number of the identical profile observed in a previous study [11].
RESULTS AND DISCUSSION
Both PATS and PFGE distributed the 48 O157 isolates into two major clades, each including the same set of isolates. Although PFGE generated more clusters in one of the clades, examination of the identities of the isolates showed that PATS had clustered O157 isolates from the same geographical locations better than PFGE. Overall, PATS and PFGE yielded overlapping typing results, which confirmed the ability of PATS to make an unbiased assessment of bacterial isolates while maintaining the discriminatory ability of PFGE.
The PFGE patterns obtained for the 48 O157 isolates can be observed in Fig. (1). The dendogram based on these PFGE patterns distributed these O157 isolates into two clades containing three and seven clusters respectively (Fig. 2). PFGE clade I was the same as PATS clade II, and PFGE clade II as the same as PATS clade I, with the same O157 isolates in these corresponding clades (Figs, 2 and 3). The additional clusters observed with PFGE within the clade may have been due to its inherent over-discrimination of isolates, which may be attributed to misinterpretation of complex electrophoretic patterns. This often encountered drawback of analysis using PFGE has been largely addressed with specialized instrumentation and sophisticated software to interpret the patterns generated, development of standardized protocols with additional restriction enzymes and standards; yet DNA degradation, incomplete digestion, co-migration of similar sized DNA fragments, as well as non-homologous DNA migrating as same-size bands, often result in “untypeable” or incorrect profiles [3-8, 14-17]. DNA sequence-based typing methods such as multilocus sequence typing (MLST) and multilocus variable number of tandem repeats analysis (MLVA), are becoming potential alternatives to PFGE, with less subjectivity in the interpretation of results, comparable and transferable data and ease of automation [18, 19]. However, since these techniques still rely on specialized instrumentation and software to interpret data, the PATS technique might be useful for application in any laboratory, particularly those outside of standardized networks such as PulseNet [2, 8]. PATS grouped the 48 O157 isolates into 9 XbaI-PATS types (Table 1A), 5 AvrII-based PATS types (Table 1B), and 9 combined-PATS types (Table 2). Both the XbaI-based and combined-PATS placed the O157 isolates into same groups which suggests that much of the discrimination was provided primarily by the polymorphic XbaI sites with limited discrimination by the polymorphic AvrII sites. Of these, PATS type 1, 2, 3, 6, and 9 were the same as PATS type 8, 18, 19, 11, and 9 observed in a previous analysis of 46 O157 isolates associated with human disease (Table 2) [11]. Subsequent detailed analysis of the isolates used in this study showed that this repetition of PATS profiles could be associated with the same geographic locations for most of the isolates in these groups [11]. The dendogram based on the 9 PATS types distributed the 48 O157 isolates into two clades bearing three and four clusters, respectively (Fig. 3). The O157 isolates in each cluster could be linked by geographic location/sourceonce their identities were revealed.
As depicted in Figs. (2 and 3), the 48 O157 isolates could be divided into 2 clades and 4 broad groups (A-D) based on their placement in the dendogram, and these overlapped between PATS and PFGE. Similar to previous observations, isolates from different outbreaks that shared the same PATS clade/group also shared the same clade/group by PFGE [10,11]. Some of these isolates from different outbreaks were also from different years warranting a need to further investigate their epidemiologic links, however, as these were banked isolates with limited information available this evaluation could not be done. Nevertheless, shared PFGE profiles between O157 isolates from different sources as well as re-emergence of older profiles have been reported in the PulseNet database (www.cdc.gov/ecoli/2012/O157H7-11-12/index.html) making our observation less implausible.
CONCLUSION
As a user-friendly strain typing technique,which is both easy to perform and interpret, PATS favorably complements PFGE and provides comparable results without extensive instrumentation, less subjectivity in the interpretation of results and faster than the currently used 24-hour PFGE protocol. With profiles that are easy to track and transfer between laboratories, which is critical in both epidemiological and nosocomial outbreak situations, this simulated study reaffirmed the ability of PATS to rapidly and accurately type pathogens even in a “field” situation.
Acknowledgement
This study was supported in part by Grant Cooperative Agreement Number U60-CCU303019-15 from the Centers for Disease Control-Association of Public Health Laboratories to S. B. C. and I. T. K. This work was also supported inpart by the Center for Integration of Medicine and Innovative Technology (CIMIT), through U.S. Army Medical Research and Materiel Command Cooperative Agreement Number DAMD17-02-2-0006, award to S. B. C., I. T. K., and M. J. U.S. Utility Patent Pending for PATS- a bacterial strain typing technology (Filed Nov. 1, 2001).
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.


