All published articles of this journal are available on ScienceDirect.
Evaluation of Immunoprotective Activity of Six Leptospiral Proteins in the Hamster Model of Leptospirosis
Abstract
Leptospirosis is a worldwide zoonosis caused by pathogenic Leptospira. The whole-genome sequence of L. interrogans serovar Copenhageni together with bioinformatics tools represent a great opportunity to search for novel antigen candidates that could be used as subunit vaccine against leptospirosis. We focused on six genes encoding for conserved hypothetical proteins predicted to be exported to the outer membrane. The genes were amplified by PCR from Leptospira interrogans genomic DNA and were cloned and expressed in Escherichia coli. The recombinant proteins tagged with N-terminal hexahistidine were purified by metal-charged chromatography. The immunization of hamsters followed by challenge with lethal dose of virulent strain of Leptospira showed that the recombinant proteins Lsa21, Lsa66 and rLIC11030 elicited partial protection to animals. These proteins could be used combined or in a mixture with novel adjuvants in order to improve their effectiveness.
INTRODUCTION
Leptospirosis is a highly spread zoonosis of human and veterinary concern. The disease is caused by pathogenic species of the genus Leptospira with a higher incidence in tropical and subtropical countries, where the conditions for transmission are particularly appropriate [1, 2]. Transmission to humans occurs by direct or indirect contact to water or soil contaminated with the urine of chronically infected mammals. In the urban surroundings, due to sanitation problems and large population of urban rodent reservoirs, the disease spreads and become prevalent [3]. Flu-like symptoms, such as fever, chills, headache, and severe myalgias take place in the early phase of the disease, making clinical diagnostic very difficult. Multi-organ system complications may occur in 5 - 15 % of the cases with 5 - 40 % mortality rates [1, 4].
Presently there are veterinarian vaccines derived from inactivated whole cell preparations of pathogenic leptospires. These vaccines afford protection most probably through the induction of antibodies against leptospiral lipopolysaccharide [2, 5, 6]. Therefore, they do not confer protection against leptospiral serovars not included in the preparation and as polysaccharide antigens, do not induce long-term protection. Human vaccines are available in Cuba [7] and China [8] with the same limitations of the veterinarian ones. The large number of pathogenic serovars (> 200) represents a major limitation to the production of a multi-serovar component vaccine. A cost-effective vaccine preparation against leptospirosis has long been pursued [3].
Due to their location, outer membrane proteins (OMPs) can participate of host-pathogen interactions and have become the focus of many research groups. OMPs have been shown to be surface-exposed and expressed during infection of the mammalian host [9, 10]. Several recombinant protein have been evaluated as vaccine candidates, such as, outer membrane protein OmpL1, lipoprotein LipL41 [11], hemolysis-associated protein 1 (Hap1) / lipoprotein LipL32 [12-14], immunoglobulin-like protein [15-18] and predicted outer membrane proteins [19]. Moreover, Yan et al. [20] have shown that leptospiral proteins with an OmpA-like domain conferred partial protection against lethal infection of leptospira. Recently, the evaluation of several recombinant proteins as potential antigens against leptospirosis were published but the results were not very encouraged [21]. Cell-mediated immunity has also been explored [22] and pathogenic leptospires have been shown to stimulate in vivo the production of type 1 cytokines involved in cellular immunity in hamster model [23].
In the present work, we evaluated the immune response promoted by six recombinant proteins in hamsters. Four of them were previously characterized, LIC10368 [24], LIC11030 [25], LIC10258 [26] and LIC12253 [27], but their immune protective activity has not been tested by our research group. LIC10821 and LIC10672 are hypothetical coding sequences identified by bioinformatics tools in the genome sequences of the L. interrogans serovar Copenhageni. LIC10672 has also been identified by proteomics studies with virulent leptospiral strain [28]. Our results showed that the recombinant proteins elicited partial protection against lethal challenge with L. interrogans serovar Kennewicki strain Pomona Fromm in hamsters.
MATERIALS AND METHODOLOGY
Leptospira Strains and Culture Conditions
The non-pathogenic L. biflexa (serovar Patoc strain Patoc 1) and the pathogenic L. interrogans (serovars Copenhageni strain M-20, Canicola strain Hond Utrechet IV, Hardjo strain Hardjoprajtino, Icterohaemorrhagiae strain RGA and Pomona strain Pomona), L. borgpetersenni serovar Whitcombi strain Whitcomb, L. kirshneri (serovars Cynopteri strain 3522C and Grippotyphosa strain Moskva V), L. santarosai serovar Shermani strain 1342K, L. noguchi serovar Panama strain CZ214 and the virulent strain of L. interrogans serovar Kennewicki strain Pomona Fromm (strain LPF) and serovar Copenhageni (strain Fiocruz L1-130) were cultured at 28 °C under aerobic conditions in liquid EMJH medium (Difco® - USA) with 10 % rabbit serum, enriched with L-asparagine (wt/vol: 0.015 %), sodium pyruvate (wt/vol: 0.001 %), calcium chloride (wt/vol: 0.001 %), magnesium chloride (wt/vol: 0.001 %), peptone (wt/vol:0.03 %) and meat extract (wt/vol: 0.02 %) [29]. Virulence of the L. interrogans serovar Kennewicki strain Pomona Fromm (strain LPF) and serovar Copenhageni (strain Fiocruz L1-130) was maintained by iterative passages in Golden Syrian hamsters[1]. The animals were infected with 104 leptospires and sacrificed after the appearance of symptoms, such as loss of weight and mobility (approximately 5 days post-infection). Kidneys were removed and macerated and the leptospires recovered through culture [1].
'In silico' Identification and Characterization of the Protein
Predicted coding sequences (CDSs) were selected from the L. interrogans serovar Copenhageni genome sequences [30] based on their cellular localization prediction by CELLO program, http://cello.life.nctu.edu.tw/ [31, 32]. SMART, http://smart.embl-heidelbergde/ [33, 34] and PFAM, http://www.sanger.ac.uk/Software/Pfam/ [35] web servers were used to search for predicted functional and structural domains within the amino acid of the selected CDS. The predicted lipobox sequence was evaluated by LipoP, http://www.cbs.dtu.dk/services/LipoP/ [36] program.
DNA Isolation and PCR Analysis
Leptospira cultures were harvested by centrifugation at 11,500 X g for 30 min and gently washed in sterile PBS twice. Genomic DNA was isolated from the pellets by guanidine-detergent lysing method using DNAzol® Reagent (Invitrogen), according to manufacturer's instructions. The DNA fragments were amplified using oligonucleotides designed according to L. interrogans serovar Copenhageni genome sequences (GenBank accession AE016823) (Table 1). PCR was performed in a reaction volume of 25 μl containing 100 ng of genomic DNA, 1 X PCR buffer (20 mM Tris-HCl, pH 8.4, 50 mM KCl), 2 mM MgCl2, 20 pmol of each specific primer, 200 μM of each dNTP, and 2.5 U Taq DNA Polymerase (Invitrogen). Cycling conditions were: 94 °C, 5 min, followed by 35 cycles at 94 °C, 50 sec, 62 °C, 50 sec, 72 °C, 1 min 30 sec, and a final extension cycle of 7 min at 72 °C. PCR amplified products were loaded on a 1 % agarose gel for electrophoresis and visualization with ethidium bromide.
Gene Locus, Protein name, NCBI Reference Sequence, Features, Gene Conservation, Sequence of the Primers Employed for DNA Amplification, and Molecular Mass of Expressed Recombinant Proteins
| Gene Locus1 | Recombinant Protein given Name | NCBI Reference Sequence Number2 | Description/Function | Conservation (Identity)3 | Sequence of primers for PCR amplification | Recombinant Protein Molecular Mass |
|---|---|---|---|---|---|---|
| LIC10368 | Lsa21b | YP_000355 | Putative lipoprotein | Lai (100 %) | F: 5´ CACCGATGAAAAAAAAGAAAATGAATTGAG 3’ R: 5’ AACGCGATTCATAGAGAGCG 3’ |
21 kDa |
| LIC10258 | Lsa66a | YP_000249 | Hypothetical protein with ompA domain | Lai (99 %) LBH (83 %) |
F:5´ GGATCCGAAGCCTTCTCACCCAATTG 3´
(BamH I) R:5´ CCATGGTTAAAGTGAAAGATAAAAATCGATTC 3´ (Nco I) |
65 kDa |
| LIC11030 | rLIC11030c | YP_001000 | Putative lipoprotein with a domain of unknown function (DUF1565) | Lai (99 %) | F: 5` CTCGAGCCAATTTCTTTCGATCCAAATC 3´ (XhoI) R: 5`AAGCTTTCAATCCTCTACTGCAGCCC 3´ (HindIII) |
37 kDa |
| LIC10821 | rLIC10821 | YP_000798 | Putative lipoprotein with a domain of unknown function (DUF1565) | Lai (100 %) | F: 5´ CTCGAGTGTGCCAACGAAAACG 3´
(XhoI) R: 5´ AAGCTTTCAAGGATTACAAGGTTTAG 3´ (HindIII) |
37 kDa |
| LIC12253 | Lsa25d | YP_002188 | Putative lipoprotein with a domain of unknown function (DUF1566) | Lai (100 %) LBH (77 %) LBP (39 %) |
F: 5´ CTCGAGGAGGAGAAACCGGACGATAC 3’ (XhoI) R: 5’ CCATGGTTAGGGAAGACTTCTAACACATC 3’ (NcoI) |
24 kDa |
| LIC10672 | rLIC10672 | YP_000656 | hypothetical protein | Lai (100 %) LBH (95 %) |
F: 5´CCGGATCCCAGAATAAAG 3´(BamHI) R: 5´GGGGTACCTCAGATGTCG 3´(KpnI) |
23 kDa |
Protective Efficacy of Recombinant Proteins in a Hamster Model
| Antigen | Experiment 1 | Experiment 2 | ||||
|---|---|---|---|---|---|---|
| % Protection (No. Surviving/Total) | P value* | Culture (Negative/Total) | % Protection (No. Surviving/Total) | P value* | Culture (Negative/Total) | |
| killed whole-leptospires | 100 % (10/10) | <0.0001 | 10/10 | 100 % (10/10) | 0.0002 | 10/10 |
| PBS | 0 % (0/10) | -------- | ND | 20 % (2/10) | -------- | 2/2 |
| Lsa21 | 30 % (3/10) | 0.0470* | 1/3 | 20 % (2/10) | 0.9224 | 2/2 |
| Lsa66 | 20 % (2/10) | 0.0092* | 1/2 | 30 % (3/10) | 0.6768 | 1/3 |
| rLIC11030 | 30 % (3/10) | 0.0144* | 2/3 | 30 % (3/10) | 0.5102 | 2/3 |
| rLIC10821 | 20 % (2/10) | 0.0640 | 2/2 | 20 % (2/10) | 0.8628 | 1/2 |
| Lsa25 | 20 % (2/10) | 0.0640 | 1/2 | 30 % (3/10) | 0.3784 | 3/3 |
| rLIC10672 | 0 % (0/10) | 0.4180 | ND | 40 % (4/10) | 0.3393 | 4/4 |
The statistical analysis was performed by Kaplan-Meier method [40] and Logrank test compared to PBS survival curves.
* P-values (< 0.05) considered to indicate statistical significance.
ND: not determined
Cloning, Expression and Purification of Recombinant Proteins
Amplification of the CDSs was performed by PCR from total L. interrogans serovar Copenhageni strain Fiocruz L1-130 genomic DNA using complementary primer pairs listed on Table 1. The gene sequences were amplified without the signal peptide tag, predicted by SignalP (http://www.cbs.dtu. dk/services/SignalP/) [37]. All cloned sequences were confirmed by DNA sequencing with an ABI 3100 automatic sequencer (PE Applied Biosystems, Foster city, CA). Cloning, expression and purification of the recombinant proteins LIC10368 (Lsa21), LIC10258 (Lsa66), LIC11030 and LIC12253 have been previously described [24-27]. The coding sequences for LIC10821 and LIC10672 were cloned into pGEM-T easy vector (Promega) and subcloned into the pAE expression vector [38] that allows the expression of recombinant proteins with a minimal 6 x His-tag at the N-terminus. Protein expression was achieved in E. coli BL21 (SI) strain by the action of T7 DNA polymerase under control of the osmotically induced promoter proU [39]. E. coli BL21 SI containing recombinant plasmids were grown at 30 °C in Luria-Bertani broth without NaCl and with 100 µg/ml ampicillin with continuous shaking until an optical density at 600 nm of 0.6 to 0.8 was reached. Recombinant protein synthesis was induced by the addition of 300 mM NaCl. After three hours, the cells were harvested by centrifugation, the bacterial pellets resuspended in lysis buffer (20 mM Tris-HCl, pH 8.0, 200 mM NaCl, 100 µg/ml of lysozyme, 2 mM phenylmethylsulfonyl fluoride (PMSF) and 1 % Triton X-100). The bacterial cell pellets were lysed on ice with the aid of an ultrasonic cell disruptor (Sonifier 450, Branson, USA). The bacterial lysate was centrifuged at 3,000 X g for 10 min at 4 °C. The pellets were resuspended in buffer containing 8 M urea; 20 mM Tris-HCl, pH8.0; 500 mM NaCl. The proteins were purified through Ni2+ -charged beads of chelating fast-flow chromatographic column (GE Healthcare). The rLIC10821 and rLIC10672 were refolded on-column by gradually removing urea (8-0M). The contaminants were washed away with low imidazole concentration and the recombinant proteins were eluted in 20 mM Tris-HCl, pH8.0; 500 mM NaCl and 500 mM imidazole. The efficiency of the purifications was evaluated by 12 % SDS-PAGE. The purified recombinant proteins were extensively dialyzed against PBS, pH 7.4, glycine solution (wt/vol: 0.1 %), at the proportion of 10 ml of protein per 1,000 ml of buffer, with at least five changes of buffer every 4 h for 48 h.
ELISA for Detection of Hamster IgG Antibodies
Hamster immunoglobulin G (IgG) antibodies against recombinant proteins were detected by ELISA. The plates were covered with 250 ng/well of each protein. Total IgG was evaluated by serial dilution of hamster’s sera followed by the addition of goat anti-hamster IgG peroxidase conjugated (1:5,000; KPL, Gaithersburg, MD, USA).
Hamster Immunization and Challenge Studies
Twelve male Golden Syrian hamsters (6 - 8 weeks old) were immunized subcutaneously with 50 μg of recombinant protein adsorbed in 10 % Alhydrogel (2% Al(OH)3), used as adjuvant. One booster injection was given after two-weeks with the same preparation of recombinant protein (50 μg). Negative-control group of hamsters were injected with PBS in 10% Alhydrogel. As a positive control, a group of hamsters was immunized with killed whole-leptospires (bacterin vaccine), prepared as described by Silva et al. [17]. In brief, washed pellets of L. interrogans serovar Kennewicki strain Pomona Fromm strain LPF cultures were heat-inactivated at 56 ºC for 20 min, resuspended in PBS and stored at -20 ºC until use. Hamsters were immunized with a dose of 109 inactivated leptospires in 10% Alhydrogel on days 0 and 14. Two hamsters of each group were bled, before the challenge, by cardiac puncture and the sera were analyzed by ELISA for the determination of antibody titers. The ELISA titer was considered to be the last dilution of serum that registered an optical density of 0.10. Two weeks after the second immunization, groups of 10 hamsters were challenged with an intraperitoneal inoculum of 3-5 X 107 leptospires assessed in a Petroff-Hausser counting chamber (approx. 400-10,000 LD50) from the same L. interrogans serovar Kennewicki strain Pomona Fromm, strain LPF, but in this case freshly isolated from liver of infected Golden Syrian hamster and no culture passaged. Hamsters were monitored daily for clinical signs of leptospirosis and euthanized when clinical signs of terminal disease appeared. For the challenges, LD50 was calculated as follows: hamsters were infected intraperitoneally with 10-fold serial dilutions. Inocula of 105–100 organisms were tested. Animals were monitored daily for clinical symptoms until 28 days post-infection. We then estimated how many virulent leptospires per injected hamster led to ca. 50% mortality, a dose therefore used for our LD50 experiments. Negative control animals were injected with the same volume of sterile EMJH media.
Ethics of Animal Experimentation
All animal studies were approved by the Ethic Committees of the Instituto Butantan and of the Faculdade de Medicina Veterinaria e Zootecnia, USP, São Paulo, Brazil.
Statistical Analysis
The statistical analysis was performed by using GraphPad Prism 3.00 (GraphPad software Inc.) which calculates survival fractions using the product limit or Kaplan-Meier method [40]. Logrank test (equivalent to Mantel-Haenszel test) was used to compare survival curves. All P-values were two-tailed and a P value of < 0.05 was considered to indicate statistical significance.
RESULTS
Bioinformatics Analysis
The genes encoding LIC10368, LIC10258, LIC11030, LIC10821, LIC12253 and LIC10672 were identified by analysis of the genome sequences of the chromosome I of L. interrogans serovar Copenhageni and each one is presented as a single copy [30, 41]. The CDS LIC12253 and LIC10258 are predicted to be outer membrane proteins, while CDS LIC10368, LIC11030, LIC10821 and LIC10672 are predicted to be extracellular based on CELLO web server [31, 32]. PFAM and SMART programs found no putative conserved domain within the LIC10368 and LIC10672 CDS; the LIC10821 and LIC11030 presented a DUF1565 (domain of unknown function), the LIC12253 presented a DUF1566 domain, and the LIC10258 presented an OmpA-family protein domain. Blast analysis showed that the CDS LIC10368 has no homology with other known bacterial proteins; LIC10258 has partial identity (25%) with other spirochetal proteins; LIC12253, LIC10821 and LIC11030 present 30% similarity with other bacterial proteins due to the presence of DUF1566 or DUF1565 domain, respectively; CDS LIC10-672 has partial identity (20-30%) with other bacterial proteins (chaperone Skp of G. lovleyi, ferredoxin of T. oceani, transposase of C. botulinum) [42, 43]. The LipoP program predicted LIC10368, LIC11030 and LIC10821 CDS as putative lipoproteins, with a cleavage site for signal peptidase II at amino acids 18-19, 17-18, 17-18, respectively [36]. Similar putative coding sequences LIC10368, LIC10258, LIC11030, LIC10821, LIC12253 and LIC10672 were found in L. interrogans serovar Lai (99 - 100 % identity with LA0419, LA0301, LA3064, LA3340, LA1508 and LA3522, respectively) [44]. LIC10258, LIC12253 and LIC10672 CDS were also identified in the genome sequences of L. borgpetersenii serovar Hardjo (77 - 95% identity with LBL0511, LBL1929 and LBL2387, respectively) [45]. LIC10368, LIC11030 and LIC10821 CDS were not identified in L. borgpetersenii [45]. Only LIC12253 CDS were identified in the genome sequences of L. biflexa serovar Patoc (39 % identity with LEPBI_I0560) [46]. Table 1 summarizes the features of the selected proteins and gene conservation within the sequenced genomes.
Distribution and Expression of Genes Among Leptospira Strains
The presence of the genes LIC10368 (Lsa21), LIC10258 (Lsa66) and LIC11030 were reported previously [24-26] having DNA fragments amplified only in L. interrogans (serovars Canicola, Copenhageni, Hardjo, Icterohaemorrhagiae and Pomona). LIC12253 gene was amplified in all pathogenic strains and a less intense band was detected in the saprophytic strain [27]. The presence of the novel genes LIC10821 and LIC10672 was examined by PCR in ten pathogenic and in one saprophytic strain of Leptospira, using a pair of primers designed according to L. interrogans serovar Copenhageni genome sequences (Table 1). DNA fragments were amplified in all ten strains of pathogenic species of L. interrogans, except that LIC10821 was not amplified in the L. interrogans serovar Canicola and L. santarosai serovar Shermani. In the case of LIC10672 gene no significant amplification product was detected in the non pathogenic strain Patoc 1 (L. biflexa serovar Patoc) (Fig. 1). Template integrity was attested by 16S DNA amplification.
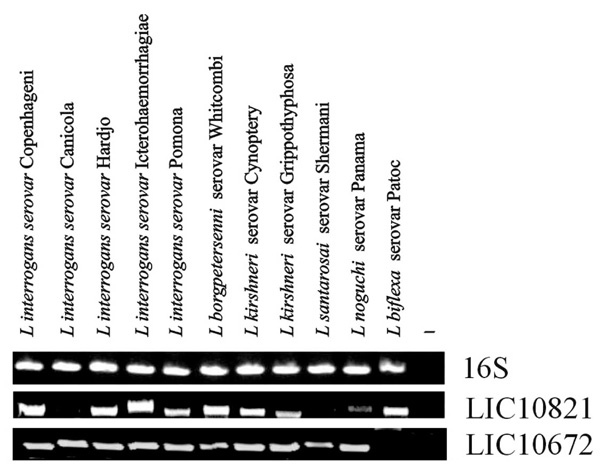
Distribution of genes among saprophytic and pathogenic leptospires. Genomic DNA from L. biflexa Patoc and from ten serovars belonging to the pathogenic species of Leptospira were subjected to PCR analysis with specific primers designed according to L. interrogans serovar Copenhageni genome sequences. (A) Amplification of 16S DNA shows template integrity; (B) LIC10821 and (C) LIC10672. No DNA was added to the negative control reaction (-).
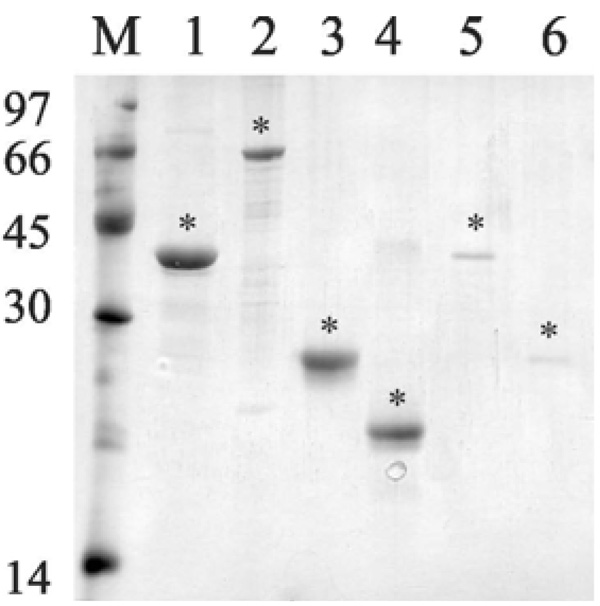
Analysis of purified recombinant proteins by metal chelating chromatography through SDS-12% PAGE. Purified protein eluted from Ni+2 - charged chelating sepharose column with 1 M imidazole. Lane M, molecular mass protein marker; lane 1, rLIC10821 (37 kDa); lane 2, Lsa66 (65 kDa); lane 3, Lsa25 (24 kDa); lane 4, Lsa21 (21 kDa); lane 5, rLIC11030 (37 kDa); lane 6, rLIC10672 (23 kDa). Protein bands were visualized by Coomassie blue staining. Positions of molecular mass standards are indicated on the left (in kDa). (*) denotes each protein band.
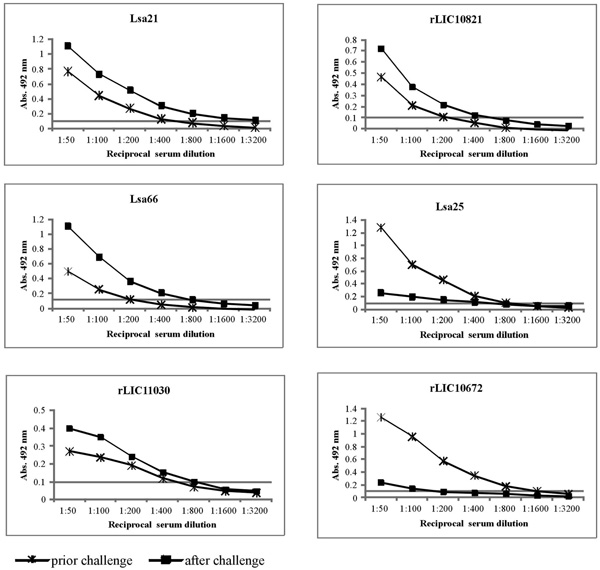
Induction of humoral immune response in hamsters. The animals were immunized subcutaneously with PBS (negative control), Lsa21, Lsa66, rLIC11030, rLIC10821, Lsa25 or rLIC10672, 50 µg each and boostered after two weeks. Animals were bled on day 30 (prior challenge) and 51 (after the challenge) to determine antigen specific total IgG responses in sera of immunized groups by ELISA method. Microdilution plates were coated with recombinant proteins and incubated with a serial dilution of serum from the immunized hamsters for IgG measurements. The data represent a pooled serum from two hamsters per group (days 30) or survival animals after challenge (day 51). All antibody titers refer to challenge assay 1.
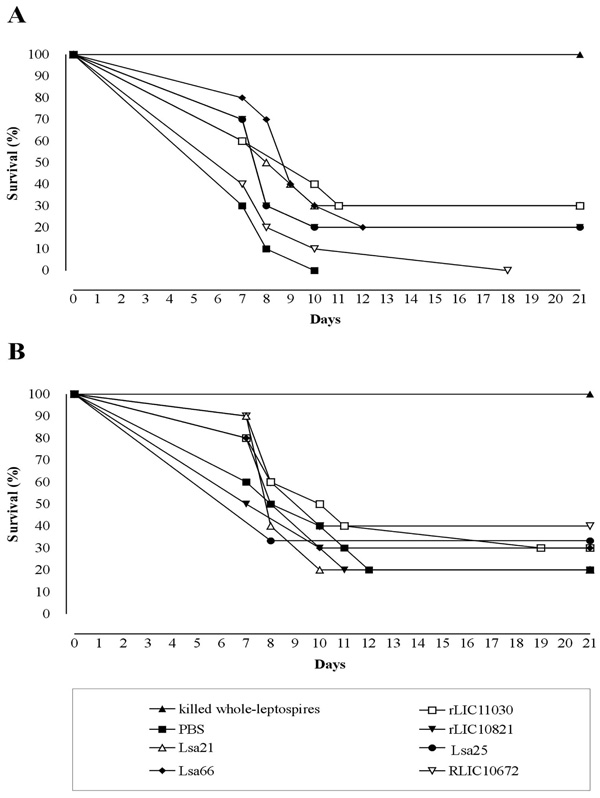
Survival curves of hamsters immunized with recombinant proteins and challenged with virulent leptospires. The animals were immunized subcutaneously with PBS (negative control), bacterin (killed whole-leptospires) (positive control), Lsa21, Lsa66, rLIC11030, rLIC10821, Lsa25 or rLIC10672. Two weeks after the second immunization (day 30) with the recombinant proteins, the animals were challenged intraperitoneally with 3 - 5 X 107 of L. interrogans serovar Kennewicki strain Pomona Fromm (LPF) in PBS. The animals were monitored for morbidity until day 21 post-challenge. A and B depicted the data of two independent experiments.
Cloning, Expression and Purification of Recombinant Proteins
The genes were amplified, without the signal peptide sequence, and the DNA insert cloned and expressed as a full-length protein in E. coli. Recombinant proteins were expressed with 6 x His-tag at the N-terminus and purified by metal chelating chromatography. Purified recombinant proteins appeared as a single band by SDS-PAGE analysis with Coomassie staining (Fig. 2).
Antibody Response Induced by Immunization with Recombinant Proteins
To assess whether hamsters immunized with the recombinant antigens (Lsa21, Lsa66, rLIC11030, rLIC10821, Lsa25 and rLIC10672) and PBS (control) promoted IgG antibody response against the antigens, sera were collected from the animals on days 30th (prior) and 51th (after) the challenge. Two animals were bled by cardiac puncture and euthanatized. The antibodies were analyzed by ELISA using the recombinant proteins as antigen probes. The data obtained for each protein are depicted in Fig. (3) and refer to assay 1. Two weeks after the second immunization (day 30), hamsters produced an antibody response with titers equivalent to 1:400 of anti-Lsa21, 1:200 of anti-Lsa66, 1:400 of anti-rLIC11030, 1:200 of anti-rLIC10821, 1:800 of anti-Lsa25 and 1:1,600 of anti-rLIC10672. After challenge (day 51) the titers measured were 1:3,200 of anti-Lsa21, 1:800 of anti-Lsa66, 1:800 of anti-rLIC11030, 1:400 of anti-rLIC10821, 1:400 of anti-Lsa25 and 1:100 of anti-rLIC10672. No significant level of antibody was detected in control animals immunized with PBS-adjuvant at any time point. An increment of antibody titer was observed after the bacterial challenge with all the proteins, except for Lsa25 and rLIC10672 that an opposite effect was detected (Fig. 3). Although the reason for the antibody decay is unknown, we may speculate that a shift in IgG subclass response might have occurred or that because Golden Syrian hamsters are not isogenic animals, thus immune response may be different.
Protection Induced by Hamster Immunization with Recombinant Proteins
Evaluation of recombinant proteins protection in hamster model followed by challenge with virulent leptospires was performed in two independent assays. The surviving plots are presented in Fig. (4) and the data obtained with the statistical analysis are depicted in Table 2. Experiment 1 (Fig. 4A) shows that immunization with the Lsa21 and rLIC11030 proteins conferred partial protection of 30%, followed by 20 % protection for the proteins Lsa66, rLIC10821 and Lsa25, against lethal challenge with 3-5 X 107 leptospires when compared to the PBS-control (Fig. 4A and Table 2). Heat inactivated whole-cell leptospires (bacterin) afforded 100% protection while no survival animal was seen with PBS-control group. In the second experiment (Fig. 4B), 40% of the animals immunized with the proteins rLIC10672 survived, 30% survived with Lsa66, rLIC11030 and Lsa25, 20% survived with Lsa21, rLIC10821 and in PBS-control group. Bacterin afforded 100% protection (Fig. 4B, Table 2). Kaplan-Meyer evaluation [40] of the kinetics of the survival curves showed a statistically significant protective effect, for the animals immunized with Lsa21, Lsa66 and rLIC11030 proteins only with the data of experiment 1. The survival animals at day 21 post-infection were sacrificed and their kidneys collected for culture, isolation of leptospires and the results are summarized in Table 2. It is interesting to note that in the experiment 1, a shift showing a delay in animal death was observed with all proteins, except with rLIC10672. Moreover, 70% of survived animals had no leptospires in their kidneys (Table 2). The rLIC10672, however, produced inconsistent results between the experiments.
DISCUSSION
Vaccines are one of the most successful methods of preventing infectious diseases [47]. Predicted outer membrane proteins have been identified during genome annotation of pathogenic L. interrogans serovars Lai [44] and Copenhageni [30] and L. borgpetersenii [45]. Almost 50% of these proteins were assigned as conserved or hypothetical proteins of unknown function [30]. Several of these proteins are predicted to be located at surface of leptospires and therefore we believe that they may be involved in pathogenesis and protective immunity. Chang and colleagues [19] working with putative outer membrane proteins of Leptospira that had no similarity with other organisms deposited in the NCBI database, identified novel antigens that although afforded partial protection against lethal challenge could act as a multicomponent vaccine against leptospirosis. Putative leptospiral outer membrane proteins with an OmpA-like domain have also been shown to elicit protective activity against hamster leptospirosis [20]. Moreover, our group has shown partial protection conferred in hamsters by two recombinant proteins, rLIC12730 and rLIC10494 [48]. More recently, Felix and colleagues [21] have shown the data obtained in challenge assays with hamsters immunized individually with 27 recombinant proteins, but their end results were not very promising.
In this work, we report the immune response induced by two previously identified adhesins, LIC10368 - Lsa21 [24], LIC10258 - Lsa66 [26], and four hypothetical outer membrane proteins of unknown function, encoded by the genes, LIC11030, LIC10821, LIC12253 and LIC10672. The proteins Lsa21, Lsa66 and LIC11030 have been experimentally shown to be surface exposed in leptospires by live immunofluorescence microscopy [24-26]. DNA amplification was not observed in the non pathogenic L. biflexa strain for all studied genes, except for LIC10821 and LIC12253 [27].
The effectiveness of these proteins to elicit protective immunity in hamster model using alum as an adjuvant was evaluated. Our results demonstrated that Lsa21, Lsa66 and rLIC11030 proteins afforded partial protection, as revealed by enhanced survival and reduced leptospiral isolation in kidney. Kaplan-Meyer analysis [40] of the kinetics of the survival curves confirmed partial protective, statistically significant effect, for the animals immunized with these proteins in one of the experiments. The number of surviving in control animals in one of the assays (0/10 against 2/10) could be responsible for the inconsistencies obtained. Noteworthy are the displacement of the curves when these proteins were administered. Comparable data were shown when OmpA-like domain proteins [20] and recombinant proteins, rLIC12730 and rLIC10494, were administered in hamsters [48]. The possible synergistic immunoprotection effect of these proteins, in combination or not, with other antigens merits further evaluation. In fact, protective effect of combined proteins have been previously observed with LipL41-OmpL1 [11] and with rLp1454 - Lp1118 – MCEII [19]. Preparations using novel adjuvants can also be used to improve and maybe, modulate the immune response. Delivery systems can also be employed in order to increment the vaccine efficacy. Work performed with leptosome-entrapped leptospiral proteins, antigens entrapped with PC-liposomes and E. coli lipid liposomes [50] have shown to improve immune response. Moreover, leptospiral antigens delivered by Mycobacterium smegmatis lipids (smegmosomes) were shown an enhancement in both Th1 and Th2 immune responses [51].
Survival protection elicited in hamsters immunized with recombinant proteins could be correlated with the antibody responses for the proteins Lsa21, Lsa66, rLIC11030 and rLIC10821, suggesting the participation Th2 immune response, although Th1 contribution cannot be excluded. The recombinant proteins were able to promote sterilizing immunity in 70% of the survived animals, as bacteria could not be recovered from kidneys (see Table 2). Moreover, the fact that some of the animals immunized with these recombinant proteins were clinically normal may suggest that bacterial burden was probably lower when compared to negative PBS control group.
CONCLUSION
We introduced in this work six proteins, including two novel leptospiral proteins. Five of these proteins are potential antigens that induced partial immunoprotection activity against challenge with the virulent Kennewicki strain Pomona Fromm strain. The employment of new generation adjuvants and the combination of one or more proteins should improve the immune protective activity of these proteins and may lead to the development of a multicomponent vaccine that could generate cross-protection against a wide range of pathogenic Leptospira serovars.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflicts of interest.
ACKNOWLEDGEMENTS
This work was supported by FAPESP, CNPq and Fundacao Butantan, Brazil; MVA, MLV, RO, RFD, RSM and ATB have scholarship from FAPESP.


