All published articles of this journal are available on ScienceDirect.
Characterisation of the MutS and MutL Proteins from the Pseudomonas avellanae Mismatch Repair (MMR) System
Abstract
The identification and analysis of the Pseudomonas avellanae mismatch repair system (MMR) were performed via sequencing and cloning the mutS and mutL genes and then analyzing the characteristics of the corresponding proteins studying their function and biological role in an E. coli heterologous system. In these studies, the P. avellanae MutS and MutL proteins were shown to localise at the nucleoid level, in a MutS-dependent manner as far as MutL is concerned, and were also able to complement the defect observed in both the mutS and mutL knockout strains of E. coli. In addition, their ability to form both homo and heterodimers between each other was shown by using the prokaryotic two-hybrid assay. Our results represent a first step to elucidate the MMR mechanism in plant pathogenic pseudomonads since the MMR genes were identified in P. syringae pathovars but there was no evidence on their action as effective repair products.
INTRODUCTION
Pseudomonas avellanae is the causative agent of bacterial canker and responsible for the decline of Corylus avellana L. This bacterium severely damages cultivated hazelnuts in northern Greece [1] and central Italy [2]. The main symptom observed in the fields is a sudden wilting of twigs, branches, and the tree. The disease is particularly prevalent during the late spring and summer.
It was reported [3] that P. avellanae, isolated directly from diseased hazelnut specimens in central Italy, showed, in some cases, a colony phase variation with the presence of transparent colony variants (i.e., water drop-like colonies) on nutrient agar medium supplemented with 5-7% sucrose (NSA). These variants appeared encapsulated, were possibly not pathogenic and did not induce a hypersensitivity reaction in tobacco leaves. Reversal towards typical P. avellanae colonies (i.e., creamy-white colour) was also observed following the re-streaking of transparent variants onto NSA plates [3].
Van den Broek et al., [4, 5] reported that MutS-dependent mismatch repair (MMR) is involved in Pseudomonas sp. PCL 1171 colony phase variation. The process occurs via the accumulation of spontaneous reversible mutations in the regulatory genes gacA and gacS, which belong to the two-component system that affects the production of secondary metabolites [6]. Indeed, mutations in the mutS gene increased the frequency of switching from phase I (opaque colonies) to phase II (translucent colonies) up to 1000-fold for the high frequency of mutations accumulated in gacA/S. These results indicate that the level of MutS-dependent repair is a determinant for the frequency of phase variation.
On the basis of these results and to investigate the characteristics of phase variation in P. avellanae, we preliminarily undertook an analysis of its mismatch repair system (MMR) by identifying and analysing its mutS and mutL genes.
Unlike the MutHLS pathway of Enterobacteriaceae [7], Oliver et al., [8] showed that the MMR system in P. aeruginosa lacks the mutH gene. Only mutS and mutL genes are present, and, as expected, their alteration is associated with a “mutator” phenotype. Although Pseudomonas aeruginosa does not have the corresponding E. coli MutH and Dam homologues, and consequently the MMR system seems to work differently, Jacquelín et al., [9] showed that the mutL gene from P. aeruginosa is able to complement a MutL-deficient strain of E. coli.
No prior studies were performed to examine the MMR system of plant pathogenic pseudomonads and all the available information (i.e., mutS and mutL gene sequences) stemmed from the complete genome sequencing of representative species, including several of the P. syringae pathovars. Additionally, as expected, no gene homologus of mutH has been identified.
MATERIALS AND METHODS
Bacterial Media and Chemicals
Nutrient broth (NB) and nutrient agar (Difco) supplemented with 5% sucrose (NSA) were used for bacterial culture and plating of P. avellanae. LB broth [10] was used for culturing and plating E. coli. SM (salt solution) was used for bacterial dilutions, as described by Miller [10]. The antibiotics used (Sigma) were ampicillin (50 µg ml-1), tetracycline (40 µg ml–1), kanamycin (30 µg ml–1), and nalidixic acid (40 µg ml–1). Synthetic oligonucleotides (Sigma) used for E. coli and P. avellanae gene amplifications and those used for P. avellanae mutS and mutL gene sequencing are listed in Table 1.
Synthetic Oligonucleotides used in this Study
| No. | Name | Sequence |
|---|---|---|
| 1 | p22 –31 SalI F | 5’-CATTGATTAGCATGGTTAGAGC-3’ |
| 2 | p22 +35 BamHI R | 5’-TTGTAAAACGACGGCCAGTGAATTCC-3’ |
| 3 | mutS P.avellanae 800 R | 5’-TCCAGGTTGCGACGGCTGGC-3’ |
| 4 | mutS P.avellanae 801 F | 5’-GCTGGACACCAACCTGTCCGG-3’ |
| 5 | mutS P.avellanae 1155 R | 5’-GCATCCAGGTCATTCA-3’ |
| 6 | mutS P.avellanae 1600 R | 5’-TCAGGTCTTCGAGCAACGTCT-3’ |
| 7 | mutS P.avellanae 1601 F | 5’-TCGACCACCTTGCGCCGCTGC-3’ |
| 8 | mutL P.avellanae 850 R | 5’-CGATACGCCTGACGCACCGCGT-3’ |
| 9 | mutL P.avellanae 851 F | 5’-CGATGTGCTGTTCAACGGCCGT-3’ |
| 10 | mutS P.syringae 1 F | 5’-ATGAATAAAGCAATTTCCGA-3’ |
| 11 | mutS P.syringae 2580 R | 5’-TCACAGGCGAGTTTGCAATG-3’ |
| 12 | mutL P.syringae 1 F | 5’-ATGACCGATCTTCTCCTCGA-3’ |
| 13 | mutL P.syringae 1938 R | 5’-TCATTGACCGCGCAGGAATA-3’ |
Bacterial Strains and Plasmids used in this Study
| E. coli Strains | Relevant Genotype | Source |
|---|---|---|
| R718 | SupE thy Δ (lac proAB) / F’ (proAB+ lacIq LacZ ΔM15) | Paolozzi collection |
| R721 | 71/18 glpT::OP434/p22lacZ | [11] |
| KM75 | (GM7698) F- As AB1157 but DmutS465::Tet | [19] |
| KM52 | (GM7532) F- As AB1157 but DmutL460::Cam | [19] |
| P. avellanae strain | ||
| CRA-ISFN1 | Pathogenic to Corylus avellana | CRA-FRU collection |
| Plasmids | Relevant genotype | Source |
| pGemT-easy vector | PCR cloning vector | Promega |
| pcIp22 434 | pcIp22 derivative harbouring the C-terminal domain of the 434 phage repressor | Paolozzi collection |
| pcI434434 | pcI434 derivative harbouring the C-terminal domain of the 434 phage repressor | Paolozzi collection |
| pcIp22 – mutSE.coli | pcIp22 derivative harbouring the E. coli mutS gene | This work |
| pcIP22 – mutLE.coli | pcIp22 derivative harbouring the E. coli mutL gene | This work |
| pcIP22 – mutSP.av | pcIp22 derivative harbouring the P. avellanae mutS gene | This work |
| pcIP22 – mutLP.av | pcIp22 derivative harbouring the P. avellanae mutL gene | This work |
| pcI434 – mutSE.coli | pcI434 derivative harbouring the E. coli mutS gene | This work |
| pcI434 – mutLE.coli | pcI434 derivative harbouring the E. coli mutL gene | This work |
| pcI434 – mutSP.av | pcI434 derivative harbouring the P. avellanae mutS gene | This work |
| pcI434 – mutLP.av | pcI434 derivative harbouring the P. avellanae mutL gene | This work |
| pTTQ18 gfp | pTTQ18 derivative harbouring the gfp gene | [12] |
| pTTQ18 gfp- mutSP.av | pTTQ18 gfp derivative harbouring the mutS gene of P. avellanae | This work |
| pTTQ18 gfp- mutLP.av | pTTQ18 gfp derivative harbouring the mutL gene of P. avellanae | This work |
| pTTQ18 gfp-mutSEc | pTTQ18 gfp derivative harbouring the mutS gene of E. coli | This work |
| pTTQ18 gfp- mutLEc | pTTQ18 gfp derivative harbouring the mutL gene of E. coli | This work |
Interaction of Pseudomonas avellanae MutS and MutL Proteins between Each Other and with the Orthologous Escherichia coli Proteins
| pcI434 pcIP22 |
MutSP.av | MutLP.av | MutSE.coli | MutLE.coli |
|---|---|---|---|---|
| MutSP.av | 38% ± 3.4 | |||
| MutLP.av | 17% ± 2.3 | 24% ± 4.3 | ||
| MutSE.coli | 36% ± 4.0 | 16% ± 1.9 | 34% ± 3.3 | |
| MutLE.coli | 28% ± 1.8 | 41% ± 2.5 | 11% ± 4.1 | 28% ± 2.0 |
Complementation of E. coli mut Deleted Strains
| Strain | Frequency of nalR Cells |
|---|---|
| R718 | 2.0x10-6 |
| KM75 ΔmutS | 1.0x10-3 |
| KM75/pcIP22-mutSEc | 1,2x10-6 |
| KM75/pcIP22-mutSP.av | 3,2x10-5 |
| KM52 ΔmutL | 1.0x10-3 |
| KM75/pcIP22-mutLEc | 1.4x10-6 |
| KM75/pcIP22-mutLP.av | 5.0x10-5 |
The results represent the mean of three independent experiments.
Bacterial Strains and Plasmids
Pseudomonas avellanae and E. coli K-12 strains as well as all plasmids used in this study are listed in Table 2.
DNA Sequencing and Sequence Analysis
The putative P. avellanaemutS and mutL genes were obtained by PCR amplification performed with synthetic oligonucleotides whose sequence derived on the analysis of the P. syringae pathovars phaseolicola, syringae and tomato sequence alignments. Because the 5’ and 3’ sequences of both genes from these pathovars are strongly conserved, we used the first and the last 20 nucleotides of the P. syringaemutS and mutL genes to design the oligonucleotides. The PCR products were then cloned in pGemT easy vector (Promega) to obtain pGemT-mutSP.av and pGemT-mutLP.av respectively.
The mutS and mutL gene sequences from Pseudomonas avellanae strain CRA-ISFN1 were obtained by sequencing the pGemT-mutSP.av and pGemT-mutLP.av vectors with primers 1-7 and 1, 2, 8 and 9 (Table 1), respectively. All sequences were determined at BMR Genomics.
Plasmid Construction and Bacterial Two Phages Two-hybrid Assay (TP-THA)
Recombinant plasmids (listed in Table 2) for the TP-THA were constructed by cloning the genes of interest in pcIP22 and pcI434 vectors. The DNA of the P. avellanae mut genes, obtained by PCR amplification using specific oligonucleotides carrying the SalI and BamHI restriction sites at the ends, listed in Table 1, was cloned in the same sites of the pcIP22 and pcI434 vectors, respectively [11]. All of the different combinations of recombinant plasmids coding for the chimerical repressors, obtained as described above, were transformed into the recipient E. coli strain R721, which carries the 434/P22 chimerical operator that governs the lacZ expression as reporter gene. β-galactosidase activity was assayed as described by Miller [10]. Bacterial cultures were grown at 37°C in LB medium supplemented with 0.1 mM isopropyl-β-D-thiogalactopyranoside (IPTG) to an OD600 of 0.3. β-galactosidase activity was evaluated for each strain and normalised to the value of E. coli R721 without plasmids. Residual β-galactosidase activity less than 50% was indicative of a protein-protein interaction, whereas activity greater than 50% indicated an uncertain or null interaction. The rationale for this cut-off is reported in previous studies [11-13]. E. coli R721 with the pcI434434 plasmid was used as a negative control and E. coli R721 with both the pcIP22434 and pcI434434 plasmids (Table 2) was used as a positive control.
Construction of GFP Derivatives for Microscopic Observation
The P. avellanae and E. coli MutS and MutL proteins fused with Aequorea victoria green fluorescent protein (GFP) were obtained by in-frame cloning of the PCR-produced DNA fragments of the corresponding genes. These were constructed in the PstI and HindIII sites, downstream of the GFP gene previously inserted in the EcoRI and PstI sites of plasmid pTTQ18, to give the plasmid pTTQ18-GFP [12].
Fluorescent Microscopy
Cultures of E. coli KM75 ΔmutS strain, harbouring the recombinant plasmids containing the genes of interest fused in-frame with GFP, were grown in LB supplemented with the appropriate antibiotics up to OD600 values in the range of 0.3 to 0.4. The addition of 0.1 mM IPTG induced the expression of the tagged genes, and the culture was further grown for 1 h at 30°C. Next, 1 µL of 4’, 6-diamino-2-phenylindole (DAPI, 5 mg mL-1) was added to a 100 µL aliquot of bacterial culture for 10 min in the dark. The sample was then centrifuged and resuspended in the same volume of saline buffer. Finally, 5 µL were loaded on poly-L-lysine-coated slides and examined using a Delta vision (AppliedPrecision) Olympus 1 X 70 microscope.
The P. avellanae and E. coli MutL localisations were examined in the E. coli KM75 strain in the absence and presence of P. avellanae MutS, coded by the pcI434mutSP.av plasmid.
Complementation Experiments
Cultures of E. coli KM75 ΔmutS strain, transformed with pcIP22mutSP.av and pcIP22mutSE.coli, respectively, were grown in LB supplemented with the appropriate antibiotics up to an OD600 equal to 0.9 and then plated on LB containing 40 μg/ml of nalidixic acid. The frequency of nal-resistant clones was compared to both the mut deleted strains without plasmids and to the R718 wild-type strain.
The same kind of experiments was performed with E. coli KM52 ΔmutL transformed with pcIP22mutLP.av and pcIP22mutLE.coli, respectively.
RESULTS AND DISCUSSION
To investigate whether the characteristics of phase variation in P. avellanae reported by Scortichini and Angelucci [3] could be related to an alteration of the MutS-dependent mismatch repair (MMR) system [4, 5], we preliminarily undertook a study of its mismatch repair system (MMR) by analysing the mutS and mutL genes.
Although MutH, MutL and MutS are essential components of the mismatch repair system in Escherichia coli only mutS and mutL genes are found in most organisms, whereas the mutH gene is limited to some proteobacteria [14] and, in particular, Pseudomonas species lack the mutH homologue [15]. In fact, no homologous sequence was found for the endonuclease MutH, necessary for cleaving the DNA strand with the mismatch in the E. coli MMR system, thus indicating that a different endonuclease is probably performing the cleavage in P. aeruginosa [8].
Identification of the Putative P. avellanaemutS and mutL Genes
Since the complete genome sequence is not available for P. avellanae, the synthetic oligonucleotide sequences used to amplify the putative P. avellanaemutS and mutL genes were obtained by the analysis of sequence alignments from P. syringae pathovars phaseolicola, syringae and tomato. Because the 5’ and 3’ sequences of both genes from these pathovars are strongly conserved, we used the first and the last 20 nucleotides of the P. syringaemutS and mutL genes to design the oligonucleotides. The DNA obtained by PCR amplification from a culture of P. avellanae strain CRA-ISFN1 with the two pairs of oligonucleotides, 10-11 and 12-13, for mutS and mutL, respectively, (Table 1) was cloned into the pGemT-easy vector and sequenced.
The P. avellanae strain CRA-ISFN1 mutS and mutL genes were entirely sequenced. The sequences were deposited at NCBI under the accession numbers AM982525 for mutS and AM982526 for mutL. The two gene sequences were roughly similar in size to the corresponding genes of P. syringae pathovar syringae (i.e., 2601 bp compared to 2568 for mutS and 1944 bp compared to 1938 for mutL, respectively). The phylogenetic analysis is enclosed as supplementary data file.
Screening of the putative MutS amino acid sequence revealed that the main differences between the P. avellanae and the P. syringae pathovars are in positions 367 to 373, where, in P. avellanae, an insertion of seven amino acids (EPGARPR) is present. The putative MutL protein shows a 96% identity to the P. syringae pathovars, with the main difference of a 2-residues (QY) insertion in positions 414-415.
A search for conserved P. avellanae putative MutS and MutL protein domains using RPS-BLAST with clusters of orthologous groups of proteins [16, 17] revealed typical Mut family domains in both cases (Fig. 1).
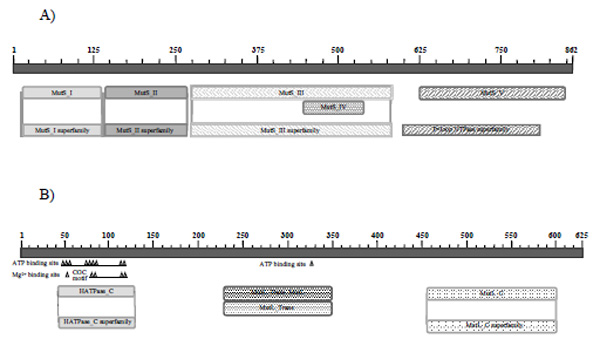
Structural domains of P. avellanae mismatch proteins. A) MutS: The aligned regions of the protein correspond: (i) with globular domain I (residues 12 to 124), which is involved in DNA binding. In this domain, there are two copies of the typical mismatch-binding motif, F-X-E, at positions 37 to 39 and 41 to 43 of the protein, as in P. syringae. In both cases, the sequence is FYE, as in E. coli, where only one copy is localised at position 36 to 38. (ii) with domain II (residues 132 to 257) defined as connector domain; (iii) with domain III or core domain (residues 263 to 572); (iv) in part with globular domain IV or clamp domain, which is also involved in DNA binding (residues 438 to 528); and (v) with domain V, which contains a Walker A motif and is structurally similar to ABC transporter ATPase domains (residues 588 to 801). B) MutL: For MutL, residues 43 to 129 constitute an ATP binding domain that contains two G-X-G motifs, present in the conserved domain of HATPase-C and located in loops defining the top and bottom of the ATP binding pocket. The transducer MutL_Trans_MutL domain extends from residue 228 to residue 349. Also, the C-terminal region of the protein (residues 456 to 601) defines the MutL C domain, which is involved in the homodimerisation of the protein.

Mut proteins localization at the nucleoid level. A) Fluorescence micrographs illustrating the localisation of P. avellanae and E. coli MutS in the E. coli ΔmutS strain. The experiments were performed as described in Materials and Methods. Cells were stained with DAPI to visualise DNA (blue). The GFP (green) highlights MutS and the merged images show the localisation of P. avellanae (A) and E. coli MutS (B) on the E. coli chromosome. B) Fluorescence micrographs illustrating the localisation of P. avellanae and E. coli MutL in the E. coli ΔmutS strain. The experiments were performed as described in Materials and Methods. Cells were stained with DAPI to visualise DNA (blue). The GFP (green) highlights MutL and the merged images show the localisation of MutL on the E. coli chromosome in the absence (A) or in the presence (B) of P. avellanae MutS.
- Localization of P. avellanae MutL.
- Localization of E. coli MutL.
Study of the P. avellanae MMR System
Since it was impossible to address directly to the question whether the two putative genes identified constitute the P. avellanae MMR system by generating and analyzing the behaviours of the mutants, due to the lack of suitable genetic tools in this organism, to assess their role in the MMR system, we studied their behaviour in an E. coli heterologous system.
The MMR mechanism predicts that, during initiation of DNA mismatch repair, a monomer of the mismatch recognition protein, MutS, after binding the mismatch on the DNA, homodimerises and then recruits MutL in the presence of ATP. The MutS (ATP)-MutL ternary complex forms and then further activates the repair process [reviewed in 18]. A consequence of this model is that MutL is unable to bind the DNA and therefore does not have the ability to localise at the nucleoid level in the absence of MutS.
To reconstruct the MMR system of P. avellanae, we studied the ability of MutS and MutL to interact individually and in combination, and to localise at the nucleoid level in the bacterial cell. The complementation of E. colimutS- and mutL-deleted strains and the study of the P. avellanae protein interactions with those of E. coli (where the MMR system is well characterised) were performed to assess their effective functionality as DNA repair proteins.
i). Interactions of the P. avellanae MutS and MutL Proteins between Each Other and with Those of E. coli
The ability of P. avellanae MutS and MutL to form homodimers and to interact between each other was studied using the prokaryotic two phages two-hybrid assay (TP-THA) [11]. In the TP-THA system, the interaction between two proteins is tested by their ability to form a chimeric lambdoid repressor. When the gene products interact they form a functional repressor that shuts down the expression of a chromosomal copy of the lacZ reporter gene, whose synthesis is governed by a hybrid 434-P22 promoter/operator region [11]. β-galactosidase synthesis, which is constitutive in the strain without plasmids, is repressed in the presence of the two plasmids only if the two proteins under investigation interact to form a functional chimeric repressor. The β-galactosidase activity of the E. coli strain R721 harbouring the pairs of plasmids was evaluated and normalised to that of the parental strain without plasmids. Residual activity less than 50% is indicative of an interaction between the two proteins [12, 13].
The two P.avellanae mut genes were cloned in frame with the N-terminal part of the lambdoid phage repressors 434 and P22 in the pcI434 and pcIP22 plasmids to form the recombinant plasmids, pcI434-mutS, pcIP22-mutS, pcI434-mutL, and pcIP22-mutL. In addition, to investigate the possible interactions between P. avellanae and their orthologous E. coli mismatch repair proteins, the E. coli mut genes were cloned in the same vectors described above. The assays were performed as described in Materials and Methods. The results represent the average of at least three independent determinations.
The β-galactosidase activity of the bacterial strain without plasmids was 2300 Miller units, and if it harbours only one of the two plasmids the residual activity was in the order of 100%.
The results (Table 3) suggest that, as expected from the behaviour of the Mut proteins, both the P. avellanae MutS and MutL proteins are able to homodimerise and also to interact with each other. In addition, although phylogenetically different, the P. avellanae MMR proteins were shown to be interchangeable with their homologous E. coli proteins during the interaction analyses.
ii). Complementation of E. coli mutS and mutL Deletion Mutants by P. avellanae Mut Proteins
Despite the phylogenetic distance between the E. coli and P. avellanae MMR proteins (see the supplementary data), the MutS proteins alignment shows about 57% identity that is mostly clustered on domains I and IV of the protein (Fig. 1). The identity decreases to 41% when the two MutL proteins are compared. In this case the best alignment is observed on the first part of the protein.
The sequence homology between the P. avellanae and E. coli Mut proteins and the results observed in the TP-THA suggest that the P. avellanae mismatch proteins could be functional in E. coli. For these reasons, we studied their ability to complement the defects of E. coli mutS and mutL deletion mutants. The E. coli strains deleted for the mutS and mutL genes show an increased mutation rate due to the loss of DNA repair. This phenotype can be evaluated by studying the mutation rate frequency of a particular marker, such as an antibiotic resistance derived by a mutation of the antibiotic target gene on the bacterial chromosome. In our experiments, we studied the frequency of nalidixic acid resistance in both the mutated and wild type strains, in the presence and absence of recombinant plasmids coding for the P. avellanae and E. coli mutS and mutL genes. The results of these studies are shown in Table 4. The frequency of nalidixic acid resistance increases about 1,000 fold in the ΔmutsS and ΔmutL strains when compared to the wild-type strain. The defects of the mut-deleted strains are fully rescued in the presence of recombinant plasmids carrying the E. coli mut genes. Additionally, the presence of the P. avellanae proteins can rescue, almost completely, the wild-type phenotype, confirming the results obtained using the two-hybrid assay.
These results indicate that the gene products identified as putative Mut proteins in P. avellanae are effectively involved in the MMR repair process and suggests that the P. avellanae Mut proteins are interchangeable with that of E. coli in performing the mismatch repair functions.
iii). Localisation of P. avellanae MutS and MutL in E. coli.
MutS localisation on the bacterial chromosome at the mismatch level is the first step in the repair process and is followed by the recruitment of MutL [18]. The onset of the P. avellanae MMR system was characterised studying the ability of its MutS protein to localise in the E. coli KM75 ΔmutS strain. This design was chosen to avoid any interference with the endogenous MutS protein.
As shown by Fig. (2A), P. avellanae MutS is able to localise in the E. coli at the same extent as the E. coli protein, whereas, according to the MMR model of action, in the absence of MutS either the P. avellanae or the E. coli, MutL does not appear to be condensed in foci, but diffused in the cytoplasm. On the other hand, the presence of P. avellanae MutS on a plasmid allows for both P. avellanae and E. coli MutL migration towards the nucleoid (Fig. 2B).
The ability of P. avellanae MutS to recruit the E. coli MutL protein on the chromosome is consistent with the result of the observed interactions between the E. coli and P. avellanae MMR proteins.
CONCLUSION
The characterisation of the P. avellanae Mut system is an early step in the study of its possible involvement in the colony phase variation of bacteria isolated from Corylus avellana plants infected by P. avellanae [3].
In their whole, our results confirmed that the gene sequences, caught by PCR amplification with the synthetic oligonucleotides designed on the P. syringae, correspond to MMR genes. Moreover, the coded proteins are interchangeable with the MutS-L proteins of E. coli. In addition, they represent a first step to elucidate the MMR mechanism in plant pathogenic pseudomonads since, until now, the MMR genes were identified in P. syringae pathovars but no evidence on their action as effective repair products was produced.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publishers Web site along with the published article.
CONFLICT OF INTEREST
Declared none.
ACKNOWLEDGEMENTS
This work was partially financed by NeCS society and “Progetti e piani per le Frontiere Tecnologiche nell’ambito dei progetti di Ricerca, Sviluppo e Innovazione delle PMI” supported by FILAS Lazio (Italy).


