All published articles of this journal are available on ScienceDirect.
Presence of a Human Diarrheagenic Escherichia coli Clone in Captivity Kept Psittacidaes
Abstract
Bacterial cultures of cloaca swabs from 86 captivity kept psittacidaes revealed 17 Escherichia coli bearing birds sharing strains which, on the basis of enterobacterial repetitive intergenic consensus (ERIC) PCR analysis, proved to be genetically similar. Further, triplex PCR specific for the genetic markers chuA, yjaA, and TSPE4.C2 was used to assign the strains to the E. coli reference collection (EcoR) B2 group. One strain of each, from the enteropathogenic (EPEC), enteroaggregative (EAEC) and Shiga toxin (STEC) E. coli pathovars were found among these isolates.
INTRODUCTION
Psittacidae comprise parrots and parakeets, which are often home cared as pets or found in many recreational zoos. In their natural habitats, they rarely have contact with the soil; their water supply comes from the fruits and some flowers rainfall deposits. As a consequence of this limited soil contact and eventual access to contaminated water, their alimentary tract´s microbiota shows a reduced presence of gram negative bacteria, such as E. coli [1-3]. In the case of home pets and institutional captivity kept birds, bacterial contamination is facilitated by contact with humans, aggregated animals and the environment [4]. In Brazil, illegally owned wild birds recovered by environmental police are usually kept in quarantine in Official Environmental Control Agencies or Non-Government Organizations facilities, which monitor eventual carriage of potential pathogens, before their re-introduction into natural habitats.
Pathogenic E. coli are classified in distinct categories according to their virulence properties, symptoms of diseases that they cause, species and age group where they are found. These criteria have been the basis for the division of human diarrheagenic E. coli (DEC) in at least five categories [5]: enteropathogenic (EPEC), enterotoxigenic (ETEC), enteroinvasive (EIEC), enterohemorrhagic (EHEC), and enteroaggregative (EAEC) E. coli. EPEC are an important agent of infantile diarrhea in developing countries; EIEC share several features with Shigella including the ability to cause dysentery in school-age children; EHEC potentially cause both hemorrhagic colitis and the hemolytic uremic syndrome; ETEC are a traveler’s cholera-like diarrhea agent; EAEC are an emerging agent of protracted diarrhea found in both developed and developing countries.
The possibility of captivity-kept psittacidaes to harbor pathogenic DEC was here investigated by analyzing cloacae swabs of 16 white-eyed conure (Aratinga leucophthalmus) and 70 blue-fronted parrots (Amazonia aestiva). The birds were in a good health condition, fed with fruits and sunflower seeds and potable water and separated in groups of eight per regularly cleaned wired suspended cages. Originally recovered from illegal owners, they were under care and observation quarantine by experts from the Floravida Institute, Botucatu SP Brazil, for at least one year, in preparation for re-introduction into their natural habitats. Following collection, swabbed samples (one per bird) were pre-cultured overnight at 40ºC in alkaline peptone water and in selenite-cysteine broth. The enrichment cultures growth was then simultaneously streaked in Hektoen and MacConkey agars and incubated under the above culture condition. Finally, a number from 3-to-5 distinct and well-isolated colonies were individually subjected to biochemical tests for bacterial identification. This enabled the detection of E. coli in 4 of the 16 (25%) white-eyed conure and in 13 of the 70 (18.6%) blue-fronted parrots. The isolates were then cultured at 37ºC overnight in trypticase soy broth (TSB) and the resulting growth added with 20% glycerol (Sigma) and stored at -80°C.
When multiple isolates from single birds showed identical biochemical profiles, only a single isolate per bird was taken for subsequent analyses, which consisted of screening for the following virulence-related features: Congo red adsorption (CRA), antibiotic resistance, hemolytic activity, HEp-2 cells adhesion, and presence of DEC-specific virulence genes.
CRA and hemolytic activity tests were performed by streaking overnight TSB cultures respectively in Congo red and blood agar (Columbia agar base mixed with 5% de-fibrinated sheep blood) plates, incubation for 18-24h and observation of corresponding colonies phenotypes. Antibiotic bacterial resistance was tested by the disc-diffusion method against the following drugs: ciprofloxacin, chloramphenicol, enrofloxacin, gentamicin, streptomycin, sulphamycin, tetracycline and trimethoprim. Unless otherwise indicated, all the culture media employed were from Difco (Heidelberg, Germany) or Oxoid (Hampshire, UK).
HEp-2 cell adhesion tests were performed according to the original method of Cravioto et al. [6]. Twenty microliters of overnight bacterial cultures in TSB were added to semi-confluent grown HEp-2 cell monolayers in chamber-slides (Nalge Nunc, Rochester, USA) containing 0.7 – 1.0 ml of Eagle essential minimum medium (Sigma, St. Louis, USA) supplied with 10% fetal calf serum (Cultilab, Campinas, Brazil) and added with 1% D-mannose (Sigma, St. Louis, USA). The preparations were incubated for 3h at 37°C in 5% CO2 atmosphere and then washed three times with phosphate buffered saline. Following methanol fixation, Giemsa staining, chamber removal and drying, the slides were observed at light microscope for cell monolayer attached bacteria. In surveys of at least 200 cells, adherent strains were those showing a minimum average number of 1 bacterium/HEp-2 cell. They were classified according to their adherent phenotypes in aggregative (AA), diffuse (DA) and localized (LA).
The E. coli isolates were screened by multiplex-PCR with primers specific for the following genetic markers [7]: ipaH, aggR, eae, stx and elt, respectively for EIEC, EAEC, EPEC, EHEC and ETEC. The reaction mixture, in a total volume of 50 μl, consisted of 1x reaction buffer, 0.15 mM MgCl2, 0.2 mM of dNTP mix (Promega, Madison, USA), 0.125 μM each of SK1, SK2, ipaIII, and ipaIV primers, 0.25 μM VTcom-u, VTcom-d, LTL, LTR, aggRks1 and aggRk2, 2.5 U Taq DNA polymerase (Promega, Madison, USA) and 5 μl of bacterial lysates. Reactions were run at 95°C/1min, 52°C/1min and 72°C/1min for 30 cycles and 72°C for 10 min in a Mastercycler pro S (Eppendorf, Hamburg, Germany). PCR products were submitted to electrophoresis in 1% agarose gels containing 104 X diluted GelRed (Biotium, California, USA) DNA stain and observed in an image capture device coupled UV transiluminator (Alpha Innotech, San Leandro, USA). In addition to the above mentioned genetic markers, PCR for the irp2 and shf – sequences commonly found in different pathogenic E. coli, particularly diffusely adhering strains and EAEC [8,9] – were performed. E. coli K12 HB101 strain was selected as a PCR-negative control and E2348-69, O42, 40T, EHY, and EIX, as EPEC, EAEC, ETEC, EHEC and EIEC positive controls, respectively. Genetic similarity among the strains were investigated by ERIC2-PCR [10], and their identity with E. coli reference (EcoR) groups [11] analyzed on the basis of combination of distinct amplicons generated by triplex PCR specific for chuA, yjaA, and TSPE4.C2, as previously described [12].
When submitted to ERIC2-PCR, DNA of the 17 strains from distinct birds generated identical band profiles (Fig. 1). Genetic similarity could also be observed from the EcoR PCR typing results: all the strains had chuA, yjaA, and TSPE4.C2, a gene combination corresponding to the EcoR B2 group [12].
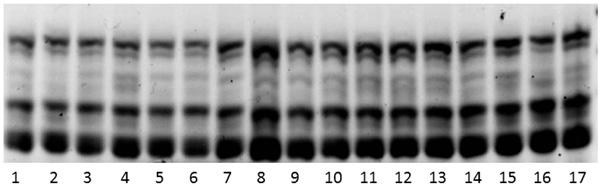
DNA fingerprints from ERIC2 PCR of 17 E. coli strains individually isolated from 17 distinct psittacidae birds.
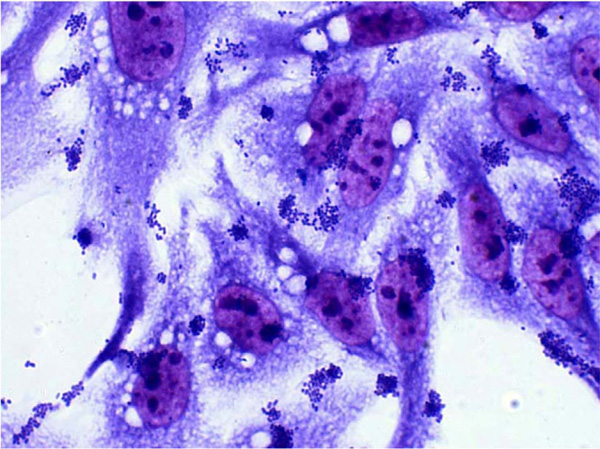
Localized adhesion to HEp-2 cells expressed by the irp2-eae-bfpA+ strain (N° 8 labeled in Table 1), after a 3h incubation period of contact bacteria-cells. May-Grunwald/Giemsa dye stained preparation. Magnification: 1000x.
Table 1 presents the virulence features of psittacidae E. coli. Five strains expressed DA, one AA and one LA (Fig. 2). One strain caused cytotoxic effect, leading to HEp-2 cell fragmentation and detachment (Table 1). A high prevalence (70.5%) of CRA+ strains was observed within this E. coli population. Over 50% of psittacidae E. coli were hemolytic and the majority of strains harbored some of the virulence genes searched for. The cytotoxic effect causing strain was PCR positive for stx and the LA+ strain had eae (Table 1). Since along with bfpA, LA and eae distinguish EPEC, an additional PCR for the bundle forming pilus gene permitted the typing of the LA+ strain as belonging to the typical EPEC pathotype. Based on the above results, three DEC strains were recognized among the 17 psittacidae E. coli: one Shiga toxin E. coli (STEC), one atypical EAEC (AA+, aggR-), and one typical EPEC.
Virulence Properties of 17 E. coli Strains Belonging to the EcoR B2 Group Isolated from Psittacidaes
| Strain a | bird b | Adhesion c | CRA d | Hemolysis | Gene (s) e | Antibiotic Resistance Profile f |
|---|---|---|---|---|---|---|
| 1 | White-eyed conure | NA | + | - | irp2 | Sensitive to all |
| 2 | White-eyed conure | AA | + | + | irp2 | CipR,CfcR,EnrR,SulR,TetR |
| 3 | Blue-frontet parrot | CE | + | + | stx | CfcR, EnrR,StrR,SulR,TriR |
| 4 | Blue-frontet parrot | NA | - | + | irp2 | CfcR,EnrR,SulR |
| 5 | Blue-frontet parrot | NA | + | - | irp2 | CfcR,SulR |
| 6 | Blue-frontet parrot | DA | + | - | none | SulR |
| 7 | Blue-frontet parrot | NA | + | - | none | CfcR,EnrR,SulR |
| 8 | Blue-frontet parrot | LA | - | + | irp2, eae, bfpA | CipR, CfcR, SulR |
| 9 | Blue-frontet parrot | DA | + | + | irp2 | CfcR,SulR |
| 10 | Blue-frontet parrot | DA | + | - | none | SulR |
| 11 | Blue-frontet parrot | DA | + | + | irp2 | CfcR |
| 12 | Blue-frontet parrot | DA | + | - | irp2 | CfcR,SulR |
| 13 | White-eyed conure | NA | + | - | irp2 | GenR |
| 14 | White-eyed conure | NA | + | + | irp2 | SulR |
| 15 | Blue-frontet parrot | NA | - | + | none | CfcR,EnrR,GenR,SulR |
| 16 | Blue-frontet parrot | NA | - | - | none | CfcR,EnrR,GenR,SulR |
| 17 | Blue-frontet parrot | NA | - | + | none | CfcR,SulR |
a Each strain was isolated from a distinct bird
b Bird species: White-eyed-conure: Aratinga leucophthalmus; Blue-fronted-parrot: Amazonia aestiva
c HEp-2 cell adhesion phenotypes; AA, DA, LA: aggregative, diffuse and localized adhesion, respectively; NA: non adherent; CE: cytotoxic effect
d CRA: Congo red adsorption.
e Virulence genes: bfpA: bundling, structural subunit of the EPEC bundle forming pilus; eae, EPEC intimin; stx, Shiga cytotoxins; irp2, yersiniabactin.
f Antibiotics: Cip, ciprofloxacin; Cfc, chloramphenicol; Enr, enrofloxacin; Gen, gentamycin; Str, streptomycin; Sul, sulphamycin; Tet, tetracycline; Tri, trimethoprim.
Antibiotic resistance was observed for at least one of the eight drugs tested and a higher prevalence of resistant strains was for sulphamycin and chloramphenicol (82.3 and 70.6%, respectively). Multidrug resistant strains include EAEC and STEC, which were the single TetR and StrR-TriR strains detected, respectively (Table 1).
Although E. coli can be pathogenic for humans, the significance of their association with psittacidaes is controversial. Enteric microbiota in these birds is mostly comprised of gram-positive [1] and for some authors [2] the mere observation of E. coli in feces of a bird presented with enteritis could be significant enough to relate the bacteria with the symptoms. Yet, reports of E. coli isolation from psittacidaes are abundant, even in clinically healthy birds [1], sometimes in figures as high as 80% [4]. E. coli incidence varies according to the host psittacidae species [1], hygiene conditions and possibly the human contact [4]. The E. coli prevalence observed here – 25% for white-eyed conure and 18.6% for blue-fronted parrot – are not too different from previously reported results [1,3].
CRA, a frequently employed approach to identify chicken colisepticemic invasive E. coli, has previously been used to differentiate potentially virulent from innocuous psittacidae E. coli [13]. Accordingly, CRA+ isolates are found only in birds presenting clinical signs of illness, apparent behavioral change or physical debility. However data shown here do not support this conclusion, because, although 12 of 17 (70.5%) birds harbored CRA+ strains, all of them were in a good health and adapted to their condition at the time of sample collection.
Apart from the role played by these bacteria in their bird host, the virulence markers that they possess grant the recognition of them as human pathogens. Significant difference is observed in E. coli prevalence between birds kept under strict hygiene rules and limited human contact and those of public recreational parks and zoos, with very higher figures for the later [4], in an indication of birds’ bacterial contamination from humans. The identification of three DEC bearing birds here demonstrates that they can be carriers of these enteropathogens, configuring a public health problem. It should also be noted that one E. coli-free bird harbored a Salmonella enterica serovar Enteritidis isolate (data not shown). Since geographical distribution of psittacidae species is restricted, unmonitored commerce may turn them potential and generalized transmitters of these enteropathogens. In fact, EPEC a particular diarrhea agent from developing countries have been detected in a pscittacidae of unknown origin in Germany [3]. Besides the detection of EPEC, the finding of a stx+ strain in psittacidae was also of great significance in the present work, since strains with this marker has usually been found only in mammalian hosts. Although assays were not conducted on Vero cells, which is the appropriate model for Shiga cytotoxins detection, phenotypic evidence could be inferred from the bacterial action on HEp-2 cells in the adhesion test and the strain hemolytic activity. This strain was resistant to five of the eight antibiotics tested. The third detected DEC was EAEC, presently recognized as a universal enteropathogen [14] and apparently assuming a more significant role as an infantile diarrhea agent than EPEC in developing countries [15]. The ability to express the aggregative adhesion is the shared E. coli phenotype of this group of strains, which nevertheless are very heterogeneous with respect to other features. Some are usually isolated from feces of normal subjects and the clinical significance of isolation depends on a more complete strain characterization, besides adhesion assays. A number of genetic markers have been identified in EAEC, which based on the presence of aggR, a plasmid encoded transcriptional activator of virulence factors, have been classified in typical (aggR+) and atypical (aggR-) [14]. The role of atypical EAEC in human disease is as yet unclear. It has been found that shf+ atypical EAEC are associated with diarrheal cases [16], but none of the psittacidae strains was positive for this marker. However, the atypical EAEC of this study was multidrug resistant (Table 1), a feature adding some significance to its finding. The DA phenotype and the irp2 gene were very common among the psittacidae strains (Table 1). The later is part of the yersiniabactin operon, is relatively ubiquitous among EAEC and DAEC strains [8,] and seems to be involved in biofilm formation [17]. Like EAEC, DAEC is a very heterogeneous bacterial group including at least some pathogenic strains. No particular DAEC marker other than HEp-2 cell adhesion was investigated in these strains, thus limiting the interpretation of their high prevalence among the investigated strains.
Virulence of E. coli is a clonally distributed feature within its natural populations. Commensal strains are mostly assigned to EcoR A and B1 phylogenetic groups and those bearing pathogenicity markers to B2 and D [18]. Strains belonging to the later groups are suspected of being associated with inflammatory bowel diseases cases [19]. Thus, the present finding of a genetically homogeneous E. coli population of the EcoR B2 group, including traditional human pathotypes in psittacidaes suggests a role for these birds as potential reservoirs of these bacteria, and might represent an alert for the adoption of more strict control measures to prevent their carriage not only of E. coli but of other human bacterial pathogens.
ACKNOWLEDGEMENT
This work was supported by Fapesp Grant Nº 2008/10975-0 given to J.R.


