RESEARCH ARTICLE
Rapid Identification of Mycobacterium Species with the Aid of Multiplex Polymerase Chain Reaction (PCR) From Clinical Isolates
Siddhartha Gupta1, Debasis Bandyopadhyay1, *, Suman Kalyan Paine1, Soma Gupta2, Surajita Banerjee1, Sujata Bhattacharya3, Ratan Gachhui4, Basudev Bhattacharya1
Article Information
Identifiers and Pagination:
Year: 2010Volume: 4
First Page: 93
Last Page: 97
Publisher ID: TOMICROJ-4-93
DOI: 10.2174/1874285801004010093
Article History:
Received Date: 12/7/2010Revision Received Date: 23/7/2010
Acceptance Date: 28/7/2010
Electronic publication date: 21/10/2010
Collection year: 2010

open-access license: This is an open access article licensed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted, non-commercial use, distribution and reproduction in any medium, provided the work is properly cited.
Abstract
Mycobacteria are aerobic, nonspore forming, non-motile,single-cell bacteria.Of more than 40 currently recognized species of mycobacteria, Mycobacterium tuberculosis, the causative agent of human TB is the commonest pathogen for pulmonary and extra pulmonary tuberculosis cases. The other members of the Mycobacterium tuberculosis complex (MTC) or the nontubercular mycobacterium (NTM) produces similar diseases which cannot be differentiated from tuberculosis by clinical symptoms and signs. But this differentiation is important as the chemotherapy varies widely according to the strain of mycobacterium. The burden of morbidity and mortality of tuberculosis is rapidly growing worldwide, particularly with the HIV/AIDS epidemic. The strain identification of Mycobacterium remains a cumbersome, labor intensive and expensive procedure, which requires 3 to 12 weeks of time. The conventional methods of strain identification lack proper standardization and precise diagnosis. The prime objective of this study is to overcome these problems.
A multiplex PCR using 3 amplicons of 165,365, and 541 base pair target sequences was done with a total number of 165 clinical isolates of suspected Koch’s patients. Strain identification was compared both by conventional methods and multiplex PCR. The results of the study show that this multiplex PCR is supposed to be less complicated, less time consuming, cost-effective and superior to the conventional methods. It is also applicable for culture negative samples where strain identification is not possible by conventional approach.
1.. INTRODUCTION
Based on surveillance and survey data, WHO estimates that 9.27 million new cases of TB occurred in 2007.Of these 9.27 million new cases, an estimated 44% were new smear positive cases, 14.8% were HIV positive cases and 4.9% were MDR-TB (which includes 3.1% new cases and 19% previously treated cases). An estimated 1.32 million HIV-negative people died from TB in 2007 and there were an additional 4,56,000 TB deaths among HIV positive people [ICD-10]. India is ranking top among 27 high MDR-TB burden countries. Incidence of MDR-TB in INDIA is reported to vary from 0.1%-5.4%.MDR-TB among all new TB cases are 2.8%, whereas 17% among the previously treated cases [1-5]. Moreover, the HIV/AIDS has produced a devastating effect on TB control worldwide [6]. Among the mycobacterial diseases (both pulmonary and extrapulmonary) M.tuberculosis complex has got the lion’s share. But the infection with Nontubercular Mycobacterium (NTM) or Mycobacterium Other Than Tuberculosis (MOTT) is also increasing rapidly.
This MOTT complex can cause pulmonary infection as well as extrapulmonary lesions (Lymph node, skin & soft tissue, joint or systemic disseminated infection) in immunocompromised hosts. The majority of such infections are due to M. avium & M. intracellure (MAI).Other responsible pathogens are M.scrofulaceum, M.kansasii, M.fortuitum, M.ulcerans etc. [7]. In 1959,Runyon classified NTM into four groups (I to IV) based on phenotypic characterization, particularly growth rate and pigmentation [8]. The treatment of diseases with M.tuberculosis complex and MOTT complex are different because many of the first and second line of antitubercular drugs are ineffective against many members of the MOTT organisms. The most effective drugs against these organisms are macrolide or aminoglycoside antibiotics. The treatment protocol heavily depends on the species of mycobacterium isolated, site and severity of infection [9]. The report of drug sensitivity testing is not always reliable and the duration of the treatment is protracted. On other side Pyrazinamide is ineffective in M.bovis infection [10]. So, strain identification from the clinical specimens is critical particularly in case of patients not responding to conventional therapy, relapse or in immunocompromised subjects. Moreover, an early diagnosis of species helps us to curtail huge burden of treatment with ineffective drugs and reduce patients’ morbidity and mortality [11].
The conventional method of identification of species involves culture on LJ slants and liquid medium followed by observing the rate of growth and pigment production (where applicable). Moreover, the growth of cultures was studied in different temperature gradient to identify the mycobacterial species. Biochemical tests like niacin production, nitrate reduction, semi quantitative and heat resistant catalase production etc are also required [12].
These are time consuming, complicated and laborious procedures whereas a multiplex PCR involving three amplicons (165bp,365bp and 541bp fragments) can identify and differentiate between different species of mycobacterium [13, 14] and this method is also highly sensitive and specific [15].
2.. MATERIALS & METHODS
2.1.. Specimen Collection
This cross sectional study was done with the Institutional Bioethical permission. Patients of all ages and both sexes recently diagnosed to be suffering from pulmonary or extrapulmonary tuberculosis were randomly chosen from in and out patients’ departments of three tertiary care hospital in Kolkata, Eastern part of India. The diagnosis of tuberculosis is based on the clinical examination, microscopy for AFB, manteaux test, Chest Skiagram, ESR, Cytological examination and ADA estimation of aspirated fluid . Sputum samples were collected from pulmonary tuberculosis patients. Body fluids (like CSF, pleural fluid, peritoneal fluid); blood, tissue or lymph node aspirates from patients with extrapulmonary tuberculosis were collected aseptically using all precautions. Total numbers of samples were 165.The total span of the study was conducted from July 2006 to February 2008.
2.2.. Specimen Processing
Sputum samples were liquefied and decontaminated using an equal volume of 4% NaOH, 2.9% Sodium citrate solution and N-Acetyl–L-Cysteine for 30 minutes incubation at room temperature. After decontamination, specimens were neutralized with sterile PBS (pH6.8). After centrifugation at 3000Xg for 30 minutes, supernatant was discarded and sediment was suspended in sterile water.The body fluids were concentrated in usual methods. Tissue or lymph node aspirates decontaminated and concentrated for AFB smear, culture inoculation. PBMC layer were collected from the blood samples of the patients by histopaque density gradient method. All the samples were ready for isolating DNA.
2.3.. Demonstration of Acid Fast Bacilli Smears
Smears were prepared using the dissolved sediment from all specimens, stained by Ziehl-Neelson (ZN) method, and examined for presence of AFB with a light microscope. Smears recorded as positive if at least 1 to 9 AFB per 100 high power fields were observed [16, 17].
2.4.. Isolation of DNA and Multiplex PCR for Species Identification
After decontamination and concentration, sample was moistened with adjustable amount of TE buffer (pH 7.6). After scrapping the cytological material, it was dissolved thoroughly in TE buffer. Mycobacterial DNA was extracted from clinical specimens using a modification of the lysis method described by Sritharan and Barker (1991). The sediments were resuspended in 100 μl of TE-Triton X-100 buffer (10 Mm Tris, 1 mM EDTA, and 1% Triton X-100, pH 8) with 200 µg/ml proteinase K, incubated at 56oC for 3 hours, and then incubated at 95oC for 30 min. After each incubation, the specimens were placed on ice for 5 min. The protease-treated and heated specimens were next extracted with Phenol: Chloroform: Isoamyl Alcohol (25:24:1). The aqueous phases of the organic extractions were precipitated with ethanol overnight. The DNA was dissolved in 200 μl of TE buffer [18]. Reference strains of M. tuberculosis (H37RV), M. bovis, M. smegmatis, M. intracellulare and M. avium were obtained from Tuberculosis Research Centre (TRC), Chennai, India. Extraction of genomic DNA was done from reference strains cultured on LJ media by standard method. Multiplex PCR as earlier developed in our laboratory was done to amplify the following target sequences [19, 20].
| Target Region | Primer Sequence |
| 65 KDa (hsp proteins) | 5´-CTA GGT CGG GAC GGT GAG GCC AGG- 3´ |
| 5´-CAT TGC GAA GTG ATT CCT CCG GAT- 3´ | |
| dna J gene | 5´-AAG AGG AAG GAG AGA GGC- 3´ |
| 5´-GTC GTT GAG GTT GAA CTC- 3´ | |
| IS 6110 insertion element: | 5´–GTG GGC ATG GTC GCA GAG AT–3´ |
| 5´–CTC GAT GCC CTC ACG GTT CA–3´ |
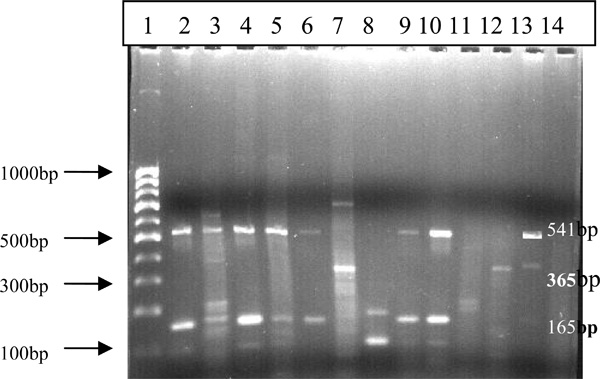
The PCR amplification was done in a Biometra Tpersonal DNA thermal cycler with an initial cycle of denaturation (5min at 85°C), followed by 40 cycles each of 1 min at 94°C, 1 min at 55°C, and 2 min at 72°C, with a final extension of 10 min at 72°C and a post incubation of 30 min at 4°C. Each DNA amplification experiment included a positive control (Reference strains like: M. tuberculosis H37Rv DNA) and a negative control (distilled water). The amplified DNA products were visualized under UV light and photographed after 2.5%agarose gel electrophoresis by ethidium bromide staining (Fig. 1).
2.5.. Culture by Inoculation in LJ Medium and Liquid Medium
All the samples (N=165) were included in the study The samples were inoculated on LJ media and liquid medium Middlebrook 7H9 with OADC supplement in our laboratory. The observations like rate of growth both at 30°C & 37°C, Colony character and pigment production (if any) in light and dark (on LJ slants) for colonies were noted:
2.6.. Biochemical Tests
Then biochemical tests like Niacin Production, Nitrate Production, Catalase production (semi quantitative & heat resistant), TWEEN-80 Hydrolysis, Growth in presence of Paranitrobenzoic acid (PNB) were done to differentiate among different strains of Mycobacterium tuberculosis complex and NTM. To differentiate between Mycobacterium tuberculosis and Mycobacterium bovis, Growth on PZA containing media, Sensitivity to TCH, Niacin production, Nitrate Reduction were performed [12, 21].
3.. RESULTS AND DISCUSSION
In this study we found that out of 165 samples, 126 (76.36%) were PCR positive whereas 95 (57.57%) were culture positive. The overall AFB positive samples was 85 (15.5%) (Table 1). Out of total samples 136(82.42%) were sputum. Rest were pleural fluid(n=14),peritoneal fluid(n=02),CSF(n=06),Blood(n=04),Lymph node Aspirate(n=02) and tissues(n=01) (Table 2). Out of 126 PCR positive samples 102 (80.95%) showed all the three target amplifications and thus diagnosed as M.tuberculosis. Only 2(1.6%) cases were attributed to M.bovis which shows amplification only at 165bp region .18 (14.28%) cases were nontuberculous mycobacteria and the rest 4 (3.10%) were inconclusive showing nonspecific amplifications.
Comparisons of Tests Done in our Laboratory with PTB and EPTB Samples
| Experiments/Tests | Positive Result in Percentage | Distribution in Percentage |
|---|---|---|
| AFB | 85 (51.5%) | PTB:80(58.8) EPTB:5(12.6) |
| Culture | 95 (57.57) | PTB: 57 (64.77) EPTB: 7 (24.13) |
| Polymerase Chain Reaction | 126 (76.36) | PTB :110 (87.30) EPTB: 16 (55.17) |
Sample Distribution of PCR Positive Cases (N=165) of PTB & EPTB
| Type of Tuberculosis | Sample | Number of Cases |
|---|---|---|
| Pulmonary (82.4%) | Sputum | 136 |
| Extrapulmonary (17.6%) | Pleural Fluid | 14 |
| Peritoneal Fluid | 02 | |
| CSF | 06 | |
| Blood | 04 | |
| Lymph node Aspirates | 02 | |
| Tissue | 01 | |
| Total | 165 |
Types of Mycobacteria Identified By PCR (N=126) and Culture (N=95)
| Species | PCR Positive | Culture Positive |
|---|---|---|
| M. tuberculosis | 102(80.95%) | 72(75.78%) |
| M. bovis | 02(1.6%) | 03(3.16%) |
| MOTT | 18(14.28%) | 06(6.32) |
| Unidentified | 04(3.1%) | 14(14.74) |
| Total | 126 (100%) | 95 (100 %) |
The atypical mycobacteria (NTM) were identified either by amplification of both 165 and 365 bp target sequences or only 165bp M. intracellulare amplicons were 20–40 bp shorter than that of M.tuberculosis or M. bovis (target region 165 bp) due to small deletion. It can be used to differentiate this organism from the M .tuberculosis complex [19, 20].
Out of 95 culture positive samples 72(75.78%) were Mycobacterium tuberculosis whereas 3 (3.16%) were M.bovis and 6 (6.32%) were MOTT or NTM. 14 (14.74%) strains remained inconclusive (Table 3). This study was in accordance with the study made by Schultz et al. [22], who demonstrated the isolations were 71% of M.tuberculosis and 34% of atypical mycobacterial strains from 191 archival specimens of tissues. In the study of Ergin et al. the respective isolations were 79.1%and 20.8% in 120 specimens [23].
Species identification by conventional cultural methods and biochemical tests are of limited value because of the comparatively low yield of the culture and long time to grow. Even the so-called rapid growers may require 1-3 weeks to grow. The differentiation of M.tuberculosis and M.bovis is not always conclusive. Niacin production and PZA sensitivity tests lengthen the reporting time span. Pigment production in light (photochromogens) and in both light and dark (scrotochromogens) are subjective observations that cannot be standardized To achieve good pigment production colonies should be young, actively metabolizing, isolated and well aerated. [24] Some members require prolonged exposure to light. Some others (like M. szulgi) need incubation at 35 ºC. In our study, we also observed the growth both at 30 ºC and at 37ºC.Moreover many important members of MOTT complex e.g. M.avium-intracellulare (MAI) and M.ulcerans etc. are nonchromogenic i.e. do not produce any pigment at all. Complicated biochemical tests like Niacin production, Nitrate reduction, Tween80 hydrolysis, Catalase etc. are required to differentiate them from M.tuberculosis complex as already mentioned. [12,25].
In the current study, even with this time consuming laborious effort the species identification by conventional methods is low. It was possible in 81 culture positive cases which are only 49.99% of total 165 cases. Schultz S et al. [22] also, in their study, found that only 41 out of 190 specimens (21.5%) were culture positive in which species identification could be made. On the other hand, species identification is much rapid and more accurate alternative where 122 cases out of 165(73.93%) could be categorized into different species. The overall isolation of M.tuberculosis Complex (M.tuberculosis and M. bovis) is 63% [M.tuberculosis 61.8%and M.bovis1.2%] in comparison to MOTT which is 10.9%. 2.4% cases gave ambiguous results by showing nonspecific bands on PCR followed by gel electrophoresis. This may be due to contamination. Ergin et al. [23] in their study, evaluated in share of M.tuberculosis as 79.1% and MOTT as 20.8% in 120 mycobacterial strains isolated from clinical specimens by PCR-RFLP. The result of our study is in accordance with the study made by Ergin et al. It is to be noted here that the failure in species identification by conventional methods is much higher (14 cases) which accounts for 8.48% of overall 165 cases.Over last decade, the rate of non-AIDS associated infection is also increasing and many of the newly identified NTM species have been identified using molecular biological tool (including DNA sequencing) rather than outdated phenotypic characterization. Thus PCR base studies along with some conventional tests are becoming indispensable for proper identification of NTM organisms [12].
The sensitivity and specificity of PCR depends on selection of the target sequence. Eisenarch et al. used primer designed from IS6110 sequence to diagnose pulmonary tuberculosis [25]. Norle et al. demonstrated that IS 6110 based PCR assay is 91% sensitive & 100% specific in detecting Mycobacterium tuberculosis in sputum [26]. But Kent et al. detected 24 false positive cases out of 31 non tuberculosis cases. It was thought that mycobacterial species other than Mycobacterium tuberculosis might be responsible for this [27]. In this study we have included hsp 65 genes & dnaj gene along with IS6110 and performed a multiplex PCR, so that it does not only diagnose Mycobacterium tuberculosis with greater accuracy but also help to identify the species of mycobacterium, as already mentioned. But one point should be kept in mind. Due to widespread and ubiquitous presence of MOTT organisms in nature, they can colonize in different human organs without causing infection. Hence, mere isolation of NTM organisms from clinical specimens is not tantamount to infection by itself. Therefore the American Thoracic Society recommends diagnostic criteria for NTM diseases to help physicians interpret laboratory results [12, 28].
4.. LIMITATIONS OF THE STUDY
DNA sequencing, DNA micro array and line probe assay to identify the causative organisms are definitely more conclusive. If these procedures could be included, the study would get enriched. But belonging to third world country and having a huge burden of tuberculosis patients, these sophisticated tests could not be done as a routine practice.
5.. CONCLUSION
To differentiate between M.tuberculosis complex and nontuberculous mycobacteria, multiplex PCR is a very good tool. If it is used in conjunction with culture and biochemical tests; it is likely to give best results. Moreover, our in house developed multiplex PCR will be much more cost effective than other molecular biological assay.
ACKNOWLEDGEMENT
Our team heartily acknowledges Department of Science and Technology (DST), Government of India for their financial assistance through the Project (SR/SO-HS-68/2006) to carry out the research work. Moreover we are grateful to RNTCP units of Kolkata Municipal Corporation for their regular support for detecting the tuberculosis patients.