All published articles of this journal are available on ScienceDirect.
Identification of Novel Mobile Genetic Elements Associated with Resistance to Macrolide and Lincosamide in Streptococcus dysgalactiae subsp. equisimilis
Abstract
Background:
Streptococcus dysgalactiae subsp. equisimilis (SDSE) is an important human pathogen. Recently, several studies have described the incidence of antibiotic resistance for SDSE worldwide, however, the data on the presence of corresponding genes and their possible association with mobile genetic elements are still limited.
Objective:
The objective of this research was to analyze the macrolide resistance in SDSE and to identify genetic determinants, mechanisms of resistance, and association with mobile genetic elements.
Methods:
A total of 9 SDSE strains from the collection of Joint Russian-Vietnamese Tropical Research and Technological Center (Hanoi, Vietnam) were used. These strains were previously isolated from throat swabs of children with pharyngotonsillitis in 6 provinces in Vietnam from 2012 to 2015. Antimicrobial resistance was tested by disk diffusion method. The presence of antibiotic resistance genes (ARG) was analyzed by PCR. The strains were characterized by emm typing and multilocus sequence typing (MLST). Illumina sequencing was employed for genome analysis of 4 representative SDSE isolates. Analysis of genetic elements with antibiotic resistance determinants was done using PubMed database and BLAST-searches. Artemis was used for comparative analysis of genetic elements.
Results:
In our study, we identified emm types that were similar to those reported in other studies. All SDSE isolates remained susceptible to penicillin, but presented alarming level of resistance to macrolides, tetracyclines, and fluoroquinolones. Most of the erythromycin-resistant strains were also characterized by clindamycin-resistance (MLSB phenotype). Both erm and different alleles of mef genes widely distributed among streptococcus pyogenes and Streptococcus pneumoniae were detected, except erm (TR) gene. The genetic elements carrying resistance determinants showed significant interspecies similarities, indicating conjugative transfer of antibiotic resistance genes between streptococcal species.
Conclusion:
Identification of the novel antibiotic resistance genes in SDSE indicates the necessity of monitoring of antibiotic resistance spreading and gene transfer in this bacterium.
1. INTRODUCTION
The human β-hemolytic streptococci (BHS) include streptococcus pyogenes (group A streptococci, GAS), streptococcus agalactiae (group B streptococci, GBS), and Streptococcus dysgalactiae subsp. equisimilis (group C and G streptococci, SDSE). SDSE colonizes the skin or mucosal surfaces, such as the respiratory tract, gastrointestinal tract, or vagina and causes a variety of diseases including invasive diseases. During the recent years the incidence of invasive SDSE diseases has increased, and in some geographic regions the rate of SDSE diseases was higher than those of GAS and GBS [1-3].
At present, penicillin is considered as drug of choice for treatment of β-hemolytic streptococcal (BHS) infections. Additionally, macrolides and clindamycin can be used for treatment of the patients intolerant to β-lactam antibiotics. Importantly, combined antibiotic therapy can reduce mortality in case of severe disease manifestations [4], probably through the abrogation of toxin synthesis [5]. Significant variations in resistance to macrolides, lincosamides (such as clindamycin) and streptogramin B (MLSB) in BHS were reported. MLSB resistance in β-hemolytic streptococci is mostly provided by the target modification enzymes encoded by erm genes, resulting in the resistance to all three classes (MLSB), or mef genes encoding efflux pumps specific for macrolides [6]. Lincosamide resistance genes lsa and lnu also occur in streptococci [7]. In GAS and GBS, all these resistance genes are located within mobile genetic elements. Their spreading among streptococci occurs by either horizontal genetic transfer or clonal expansion [6-9], however, their dissemination has not been extensively studied. In vitro studies showed the existence of conjugal transfer of integrative conjugative elements (ICEs) harboring resistance genes between streptococcal species [8].
As a consequence, the goal of the present study was to analyze the current status of antimicrobial resistance in SDSE. In addition, genome sequencing was performed to identify antibiotic resistance genes and their possible association with mobile genetic elements.
2. MATERIALS AND METHODS
2.1. Bacterial Isolates
A total of 9 SDSE strains from the collection of Joint Russian-Vietnamese Tropical Research and Technological Center (Hanoi, Vietnam) were used. These strains were previously isolated during 2012-2015 from 1359 children of 7-10 years old from different regions of Vietnam [10]. Bacteria were grown on Columbia base agar with 5% of sheep blood and in Todd-Hewitt broth with 5% of inactivated horse serum in 5% CO2 atmosphere at 37oC. Bacterial DNA was isolated by phenol/chloroform extraction. Species identity was identified with 16S rRNA gene sequencing [11].
2.2. emm Typing
emm typing of SDSE isolates was performed in accordance with recommendations available at CDC web site (www.cdc.gov/streplab/groupa-strep/emm-typing-protocol.html).
2.3. Antimicrobial Susceptibility Testing
The isolates were tested for susceptibility to penicillin G, cefotaxime, vancomycin, amikacin, norfloxacin, erythromycin, clindamycin, and tetracycline by disk diffusion method according to EUCAST guidelines. MLSB resistance phenotype was analyzed by double disc diffusion method [12]. Depending on resistance to erythromycin and/or clindamycin, the SDSE isolates were divided into several groups/phenotypes: constitutive MLSB-resistance (cMLSB), inducible MLSB-resistance (iMLSB), macrolide resistance (M-phenotype), and lincosamide resistance (L phenotype). Streptococcus pneumoniae strain ATCC 49619 was used as a control.
2.4. Antibiotic Resistance Gene Detection
PCR detection of antibiotic resistance genes erm(B), erm(TR), mef(A/E) (resistance to macrolides, and tet(M), tet(O), tet(T), tet(S) (resistance to tetracycline) was done using the primers previously published (Table S1).
2.5. Bioinformatic Analysis
The genome sequencing was performed for NT15, V123, B82 isolates with reduced susceptibility to erythromycin or clindamycin, and T201 isolate susceptible to erythromycin and clindamycin. Construction of the libraries and DNA sequencing on MiSeq platform was done as recommended by the manufacturer (Illumina, Essex, United Kingdom). Quality of the reads was tested using FastQC, trimmed with Trimmomatic [13], assembled by Spades [14], and subsequently annotated by the NCBI Prokaryotic Genome Annotation Pipeline (PGAP).
Multilocus sequence typing (MLST) was done using the Center for Genomic Epidemiology website [15]. Relevant resistance genes were identified using the ResFinder and CARD databases [16, 17]. Verification of potential integrative conjugative elements (ICEs) was performed using ICEberg 2.0 [18] together with PubMed database and BLAST-searches. Artemis was used for comparative analysis of genetic elements [19].
3. RESULTS
As previously published, a total of 152 β-hemolytic streptococci were isolated from 1359 children of 7-10 years old from different regions of Vietnam in the period 2012-2015 [10]. GAS were isolated from 49 of 1359 (3,6%) examined children, while group C and G streptococci were isolated from 8 (0,6%) and 75 (5,5%) children, respectively [10]. Using cpn60 gene based PCR approach [20] for differentiation of the species within groups C and G, a total of 9 SDSE strains (1 – group C, and 8 – group G) were identified. Other identified strains included group C S. anginosus (4 isolates), S. parasanguinis (1 isolate), S. сonstellatus (1 isolate), S. gardonii (1 isolate), and group G S. anginosus (54 isolates), S. сonstellatus (4 isolates), S. parasanguinis (3 isolates), S.sanguinis (3 isolates), S. mitis (2 isolates), S. australis (1 isolate).
3.1. SDSE Genetic Diversity
A high clonal diversity was discovered: a total of 6 emm types were identified among the 9 SDSE isolates, and stC5345 emm type was specific for 3 out of 9 isolates. MLST performed for 4 sequenced SDSE isolates revealed 3 different previously published STs (Table 1).
| Isolate SDSE | emm | MLST | Phenotype | Genotype | ICE-family | Other Resistance Genes |
| B159 | stC5345 | nt | MLSB sensitive | - | - | tet(M) |
| HF196 | nt | MLSB sensitive | - | - | tet(M) | |
| T201 | 44 | MLSB sensitive | - | - | tet(M), pat(B) | |
| HF112 | stG480 | nt | cMLSB | erm(B) | Tn6002 | tet(M) |
| NT15 | 323 | cMLSB | mef(G) | ФNT15 | tet(S), lnu(B), lsa(E), pat(B) | |
| V123 | stG6 | 44 | M | mef(A) | Ф46.1-like | tet(O), lnu(C), pat(B) |
| V63 | emm44 | nt | cMLSB | erm(B) | Tn917 | |
| T122 | stG4831 | nt | M | mef(E) | mega | |
| B82 | stC36 | 499 | cMLSB | erm(B) | ICE-B82 | tet(T), pat(B) |
3.2. Antimicrobial Susceptibility
All SDSE isolates of this study were susceptible to cefotaxime, vancomycin, penicillin G and resistant to amikacin. The incidence of fluoroquinolones resistance (norfloxacin) was detected in 4 isolates that corresponded with the presence of pat(B) gene (Table 1). Resistance to tetracycline was observed in 6 isolates. The overall prevalence of resistance to MLSB antibiotics among SDSE was very high (6 isolates were erythromycin resistant, and 4 of them – additionally resistant to clindamycin (cMLS phenotype)).
3.3. Detection of Antibiotic Resistance Genes
In 6 SDSE isolates with M or cMLSB phenotypes (reduced susceptibility to MLSB) the corresponding resistance genes were detected (Table 1). Almost all isolates (3 out of 4) with cMLSB phenotype possessed the erm(B) gene, while mef(G) was discovered in one isolate which additionally possessed clindamycin resistance genes lsa(E) and lnu(B).
The M phenotype specific for 2 isolates was associated with mef(E) and mef(A) genes, and clindamycin resistance gene lnu(C) was detected in mef(A) positive isolate.
Among the 9 SDSE isolates, 4 isolates demonstrated co-resistance to tetracycline. tet(M) was the most common, while tet(O), tet(S) and tet(T) were also discovered (each by one) (Table 1).
Furthermore, we used whole genome sequencing of 4 SDSE isolates to identify mobile elements responsible for the spreading of antibiotic resistance genes. In these isolates, the gene pat(B) encoding ATP-binding cassette (ABC) for fluoroquinolone antibiotics efflux pump was also discovered.
3.4. Mobile Elements Involved in Macrolide and Lincosamide Resistance
3.4.1. Erythromycin-resistant SDSE Isolates Carrying the mef Genes
We identified 3 isolates harboring different alleles of mef gene (mefA/E/G). As expected, in tetracycline-susceptible isolate T122 PCR assay demonstrated that mef(E) gene was located within mega (macrolide efflux genetic assembly) element, which has been previously described for BHS and Streptococcus salivarius [21, 22].
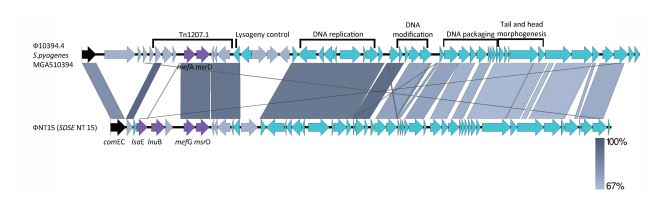
In SDSE strain NT15 (GenBank accession number JAFELF000000000) the mef(G) gene was located within the prophage-associated genetic element (54,700 bp). This genetic element consisted of 54 open reading frames (ORFs) and was chimeric in nature; it appeared due to the insertion of a Tn1207.1-related transposon into streptococcal prophage. BLASTN analysis revealed the high similarity of this prophage to other prophages, especially Tn1207.3 (52,491 bp) or Ф10394.4 (58,761 bp) discovered in tetracycline-susceptible GAS isolates [23, 24]. In both GAS isolates and SDSE NT15 strain these phages were integrated into the same chromosomal gene, comEC (Fig. 1). A Ф10394.4-like element in NT15, entitled ФNT15, harbored mef(G) instead of mef(A) and the additional 5 kb region containing unique lsa(E) and lnu(B) genes. The lnu(B) gene is involved in lincosamide modification/inactivation; and the lsa(E) encoding ABC transporter is responsible for active efflux of lincosamides, streptogramins A, and pleuromutilins. BLASTN analysis revealed that this region in SDSE strain NT15 is similar to the lnu(B)-containing sequences of S. agalactiae (JQ861959), Staphylococcus aureus (JX560992), Enterococcus faecalis (AF408195) and swine Enterococcus faecium isolate (KF421157.1) [25]. In addition, in NT15 isolate tetracycline susceptibility is associated with the presence of silent tet(S) gene.
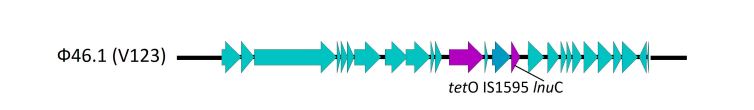
The complete sequence of SDSE V123 isolate (GenBank accession number JAFELD000000000) revealed that tetracycline resistance is provided by tet(O) gene, and tet(O) was linked with mef(A) within phage-like element. It is highly similar to well-known Ф46.1 of S. pyogenes, which resulted from the insertion of transposon Tn1207.1 into a prophage [26]. However, an addition of IS1595 element and gene lnu(C), which confer resistance to lincosamides, was found (Fig. 2). The Ф46.1-like phage was found to be integrated into a 23S rRNA uracil methyltransferase gene.
3.4.2. Erythromycin-resistant SDSE Isolates Carrying erm(B) Gene
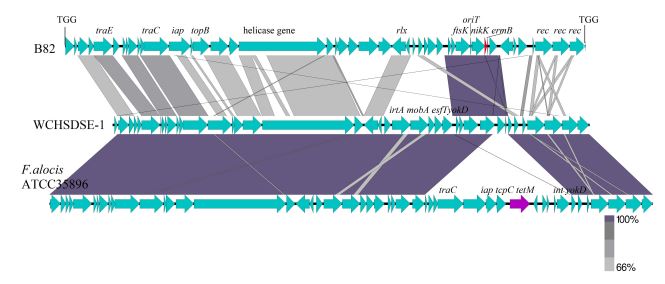
Among the 6 macrolide resistant isolates, 3 isolates had the erm(B) gene and exhibited constitutive resistant phenotype. According to the results of PCR analysis, the erm(B) was carried by the Тn917 in isolate V63 [27]. In another erm(B)-positive isolate (HF112), which was also resistant to tetracycline, erm(B) was associated with tet(M) on Tn6002 [28]. The third erm(B)-positive isolate, B82, was chosen for genome sequencing. The following analysis of B82 (GenBank accession number JAFELE000000000) revealed the presence of a 45.4-kb element. At present, this element was not found in other SDSE strains, but it has a partial similarity with SDSE strain WCHSDSE-1, which caused the streptococcal outbreak in China in 2013 [29]. Similar element, which contains the tet(M) gene instead of erm(B), is present in Filifactor alocis ATCC35896. This bacterium can cause periodontal diseases. In place of the 1.5-kb fragment containing erm(B) in SDSE, the strain F. alocis has a 14.1-kb fragment with tet(M) gene.
Thе 45.4-kb element of isolate B82 appears to be a conjugative transposon because it contains some specific genes, i.e., site-specific recombinase gene, relaxase-encoding gene (nicK), an origin of transfer (oriT), conjugal transfer coupling protein gene.
Transposition of this element is characterized by the presence of 3-bp direct repeat suggesting that the putative transposons were truly transposable elements (Fig. 3). This transposon was inserted between the genes encoding hypothetical proteins (SDSE167_0576 and SDSE167_0577 in strain 167). Additionally, Tn916 was found with gene tet(T) instead of tet(M) gene.
4. DISCUSSION
Based on the genome comparison, the SDSE which belong to group C and G streptococci, are closely related to GAS [2, 30, 31]. Almost all SDSE used in this study belonged to GGS. It corresponds to results of the previous studies, wherein Lancefield group G was found to be the most predominant among the human-recovered SDSE [32-34].
The emm genotyping was successfully performed for all 9 SDSE isolates, and 6 emm types were discovered. Four of them (stG6, stG480, stC5345, stC36) have already been found in SDSE isolated in North America, Europe and Australia [35], reflecting the successful dissemination of certain emm types in human.
Previously the certain correlation between stG480 and stG6 types and invasive infections was demonstrated [36]. However, in our study stG480 and stG6 strains were non-invasive.
Results of this study demonstrated that beta-lactams are useful for treatment of SDSE infections that correlates with other studies [3, 4, 37]. However, during the recent years the MIC for penicillin was slightly increased for some GAS and SDSE (0.12 and of 0.25 μg/ml), respectively [38]. Tetracycline resistance was noted in 6 of 9 (67%) isolates. The macrolide and lincosamide resistance in SDSE were high, 67% and 45%, respectively. Inducible clindamycin-resistant phenotype (iMLSB) was not found among erythromycin-resistant SDSE. Macrolide resistance in SDSE occurs world-wide, e.g., in Hong Kong (24%), in the USA (19%), in Europe (16%) [12, 38]. Furthermore, the percentage of clindamycin-resistant SDSE is significantly higher than in GAS. It should be taken into account during the treatment of streptococcal toxic shock syndrome caused by SDSE. In case of tonsillopharyngitis, fluoroquinolones can be considered as a second choice because the prevalence of fluoroquinolone-resistance among β-hemolytic streptococci is still low (1%), and just a few publications reported fluoroquinolone-resistance in SDSE and GAS. Resistance rate to tetracycline is higher than 60%, and for this reason it can no longer be used for empiral treatment of SDSE infections [38].
The resistance genes of macrolides, lincosamides, tetracyclines, and fluoroquinolones were examined in this study. As result, the presence of erm(A) (0%), erm(B) (33%), mef(A/E/G) (33%) differed from those of human-recovered SDSE in China: erm(A) (0%), erm(B) (78.6%), mef(A/E) (5.4%), Korea: erm(A) (4.3%), erm(B) (20.3%), mef(A) (8.7%), and Japan: erm(A) (15.5%), erm(B) (11.3%), mef(A) (2.8%) [39, 40]. Among the tetracycline resistance genes, tet(M) was the most common in this study, but tet(O), tet(S) and tet(T) were discovered (each by one). Other publications confirmed that tet(M) was more predominant than tet(O), e.g., in China and Korea (tet(M): 73.2% and 29.0%, and tet(O): 5.4% and 1.4%, respectively) in human SDSE [12, 39, 40]. Mutations in the quinolone resistance determining regions (QRDR) of gyr(A) or par(C) are considered as a major mechanism of fluoroquinolone resistance [12]. In this study, resistance to fluoroquinolone antibiotics in 4 isolates was associated with the presence gene pat(B) mediating antibiotic efflux pump.
The association of macrolide resistant determinants with mobile genetic elements was previously demonstrated for major streptococcal pathogens that explains spreading of antibiotic resistance. The resistance phenotype has been reported to be transferable by conjugation [8, 41] and transduction [42, 43]. In our study most of erythromycin-resistant strains were additionally resistant to clindamycin (phenotype MLSB). In SDSE strains we detected a number of novel genetic elements, including the new Ф10394.4-like element characterized by the insertion of lnu(B)/lsa(E)-containing sequence; and the new phi-m46.1-like prophage with insertion of lnu(C)-containing sequence, which seems to be acquired from other bacterial species [44]. The novel putative transposon carrying erm(B) gene was identified in SDSE, which is likely to be acquired from the bacteria of oral microflora. It represents one more example of genetic exchange of antibiotic resistance between gram positive cocci.
CONCLUSION
Identification of the novel antibiotic resistance genes in SDSE indicates the necessity of monitoring of antibiotic resistance spreading and gene transfer in this bacterium.
LIST OF ABBREVIATIONS
| SDSE | = Streptococcus dysgalactiae subsp. equisimilis |
| MLST | = multilocus sequence typing |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No animals and humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Not applicable.
AVAILABILITY OF DATA AND MATERIALS
The data that support the findings of this study are available from the corresponding author, [D.A.V.] on special request.
FUNDING
This study was done in the frames of projects supported by the Ministry of science and higher education (075-01135-22-00).
CONFLICT OF INTEREST
Dr. Alexander Dmitriev is on the Editorial Advisory Board of The Open Microbiology Journal.
ACKNOWLEDGEMENTS
Declared none.
SUPPLEMENTARY MATERIALS
Supplementary material is available on the Publisher’s website.


