All published articles of this journal are available on ScienceDirect.
Circulating Genotypes of Hepatitis C Virus in Italian Patients before and after the Application of Wider Access Criteria to HCV Treatment
Abstract
Aims:
The aims of this study were to report a description of the HCV genotype distribution in adult Italians and non-Italians subjects tested in the Microbiology and Virology Unit of the Padova University Hospital from January 2016 (after about one year from the availability of DAAs) to December 2018 and to compare genotype frequencies in the 12-month period before and after the application of the wider access criteria to HCV treatment.
Background:
Hepatitis C virus (HCV) infection is a major health problem, but the availability of direct-acting antivirals (DAAs) has dramatically changed HCV disease natural history because these drugs have excellent tolerability and they can eliminate the virus in almost all treated patients.
Objective:
The objective was to describe the circulating HCV genotypes in high-income countries in order to help health authorities in the future organization of DAAs treatment strategies; this aspect is not limited to drug prescription, but it also includes the identification of infected individuals who are undiagnosed, which is the limiting step to achieve the HCV elimination goal.
Methods:
Adult patients who had HCV genotype performed from 01/01/16 to 31/12/18 in the Microbiology and Virology Unit of the Padova University Hospital were included in the study: the two 12-month periods were April 2016-March 2017 (before period, BEF) and April 2017-March 2018 (after period, AFT).
Results:
Italians were 2168 (91.2%) and non-Italians were 208 (8.8%). Italians median age was 55 years, and females were older. Italians had a lower genotype 1 (p=0.0012) and higher genotype 2 frequencies (p<0.0001) with respect to non-Italians. Most patients aged 38-67 years: Italians were more represented in class age 48-57 years (p=0.0138), 68-77 years (p=0.001) and ≥78 years (p<0.0001); subjects with genotype 3 were the youngest and those with genotype 2 the oldest. Italian patients typed in the AFT and BEF were comparable; only a lower frequency of genotype 1 males and younger age in genotype 3 were found in AFT.
Conclusion:
Italians were older with respect to non-Italians, which implies that a different age based screening program could be applied. Italian genotype 3 subjects represent a cohort to focus on for the risk of therapeutic failure. Patients tested after the extended criteria for HCV treatment were very similar to those tested before, suggesting that HCV burden in Italians is higher than expected.
1. INTRODUCTION
Hepatitis C virus (HCV) infection is a major health problem; HCV is characterized by a remarkable genetic variability, and eight genotypes have been identified, with different patterns of geographic distribution. Overall, genotype 1 is the most frequently identified (46.2% of the patients), followed by genotype 3, 2, 4, 6, and 5 (30.1%, 9.1%, 8.3%, 5.4%, and 0.8%, respectively). HCV genotype 7 and genotype 8 were the last ones identified and, now, they have been found to have a limited geographic diffusion; Democratic Republic of Congo for genotype 7 and Punjab (India) for genotype 8 [1-4].
The World Health Organization estimates that about 71 million people were chronically infected in 2015, and these subjects were at risk of developing complications as cirrhosis and liver cancer [5, 6]: the availability of direct-acting antivirals (DAAs) has dramatically changed HCV disease natural history because these drugs have excellent tolerability and they can eliminate the virus in almost all treated patients [7, 8]. Recently approved DAAs are pan-genotypic, but genotype identification has still a role in the clinical approach to HCV disease and in the epidemiological field. Both naïve and experienced patients treated with sofosbuvir/velpatasvir associated or not with voxilaprevir and with glecaprevir/pibrentasvir had extremely high sustained virological response rates, but the percentages were slightly different according to genotype [9-11]. Moreover, genotype-specific drug regimens with comparable efficacy are included in anti-HCV treatment recommendations [12].
A detailed knowledge of the circulating HCV genotypes in high-income countries can help health authorities in the future organization of DAAs treatment strategies; this aspect is not limited to drug prescription, but it also includes the identification of infected individuals who are undiagnosed, which is the limiting step to achieve the HCV elimination goal [13]. The description of circulating genotypes should be updated and not only derived by global analysis of data obtained in multicenter studies; high DAAs treatment efficacy modifies rapidly the burden of HCV RNA positive patients so that peculiar epidemiological aspects may be lost, and both disease diffusion and genotype distribution are often non-homogeneous within a country [14, 15]. A survey conducted from November 2014 to December 2015 reported a 1.7% overall prevalence of patients with positive HCV-RNA in residents in 5 metropolitan Italian areas; frequency of active infection was comparable between age groups (75.% in<40 years old subjects, 79.3% in those 40–60 years of age, and 72.% in those>60 years old) [16]. This latter study focused on Italian patients, but HCV population resident in Italy also includes non-Italian subjects, who can be subjects living in Italy because of study or work reasons or migrants possibly coming from HCV-endemic countries [17]. Treatment with DAAs in clinical practice regimen is being practiced in Italy since 2014 for patients with advanced fibrosis and for other selected categories (i.e., subjects with extrahepatic disease and liver transplant recipients); revised criteria for access to antiviral therapy were outlined in March 2017, and included patients with any hepatic fibrosis degree [18].
The aims of this study were to report a description of the HCV genotype distribution in adult Italian and non-Italian subjects tested in the Microbiology and Virology Unit of the Padova University Hospital from January 2016 (after about one year from the availability of DAAs) to December 2018 and to compare genotype frequencies in the 12-month period before and after the application of the wider access criteria to HCV treatment.
2. MATERIALS AND METHODS
This retrospective study included all patients who underwent HCV genotype testing requested by the treating physician at the Microbiology and Virology Unit of the Padova University Hospital from 01/01/16 to 31/12/18; all genotype determinations were part of the routine patient management. Data of the patients were extracted from the hospital records, and this approach implied that no clinical or epidemiological data were collected; the correlations between these characteristics and HCV genotype were beyond the aim of this study. When a patient had more than one test available, only the first one was included in the statistical analysis; data were kept anonymous from the patients and discussed in aggregate form.
All patients were classified as of Italian or non-Italian origin on the basis of the place of birth reported in medical records. Subjects included in the comparative analysis were defined as ”before patients” (BEFpts) if tested from April 2016 to March 2017 and as “after patients” (AFTpts) if tested from April 2017 to March 2018. This study was conducted in accordance with the Declaration of Helsinki and local legislation and was approved by the Ethics Committee of Padova University Hospital (prot 61946).
2.1. HCV Genotype
The HCV genotypes were determined using the VERSANT® HCV genotype 2.0 assay (INNO-LiPA, Innogenetics, Ghent, Belgium) as per the manufacturer’s instructions from January 2016 to February 2018 and with Abbott RealTime HCV genotype II (Des Plaines, IL, USA) from February 2018. In-house sequencing of the core region (nt 429–741) was used when the genotype could not be determined [19, 20].
2.2. Statistical Analysis
Age was presented as median and interquartile range, and the Mann-Whitney U test was used to evaluate the differences between median values of the groups of patients. All genotypes but genotype 1 were classified by type; a separate analysis of subtype 1a and subtype 1b was performed. The Chi-squared test and Fisher’s exact test were used to compare categorical variables as appropriate (according to the frequencies).
The limit of significance for all analyses was established at p<0.05. All statistical analyses were performed with MedCalc® Statistical Software version 19.6 (MedCalc Software Ltd, Ostend, Belgium; https://www.medcalc.org; 2020).
3. RESULTS
A total of 3216 genotypes were requested in 2876 patients from January 1, 2016, to December 31, 2018. No valid result was obtained in 486 subjects (16.9%), almost in all cases because of undetectable HCV-RNA (434/486, 89.3%); a low-level HCV-RNA, under the sensibility cut-off reported in manufacturer instruction, was the reason for 39 samples (8%) and sample inadequacy for 13 patients (2.7%). Genotype 1 subtyping was available for 1305 out of 1334 subjects (97.8%).
Eleven of the 2390 subjects with HCV genotype identified were younger than 18 (median age 4 years, IQR 2-12 years), and a mixed infection was detected in 3 adult patients (1a-3 in 2 subjects and 1a-4 in one patient); these patients were excluded from the final analysis, which was conducted on 2376 subjects.
Italian subjects were 2168 (91.2%) and non-Italians were 208 (8.8%), mostly European (69.7%). Seven hundred ninety-five patients (33.5%) were tested in 2016, 969 (40.8%) in 2017 and 612 (25.7%) in 2018. Most Italian patients were male in the three periods (59.3%, 55.7% and 59.5%, respectively); conversely, non-Italian males were more than 50% only in 2017 (53.5%), and they were significantly less represented with respect to Italian cohort in 2016 (42.9%, p=0.0078) and in 2018 (38.5%, p=0.0041).
Italians' median age was 55 years (IQR 48-67 years), and non-Italians were significantly younger (median age 47 years, IQR 38-57 years, p<0.0001). The difference was also confirmed in gender analysis: the male median age was 53 years (IQR 46-60 years) versus 41 years (IQR 33-55 years) (p<0.0001) and the female median age was 61 years (IQR 50-75 years) versus 50 years (IQR 42-58 years) (p<0.0001). Both in Italian and in non-Italian cohorts, females were older than men (p<0.0001 and p=0.0015, respectively).
3.1. HCV Genotype Distribution in all Italian and Non-Italian Patients
Most Italian and non-Italian patients had genotype 1 infection, but the relative prevalence was lower in Italians (p=0.0012): subtype b was more represented in both cohorts. This subtype had a gender-related distribution, being more frequent in Italian females (71.9% vs. 39.6%, p<0.0001) as well as in non-Italian females (94.9% vs. 65.3%, p<0.0001). Italians included a higher number of genotype 2 subjects with respect to non-Italian (p<0.0001). About a third of Italian females had genotype 2 infection versus a percentage of 13.8% in males (p<0.0001): despite this difference, the frequency was significantly higher than in non-Italians for both genders (p < 0.0001 for female and p=0.0357 for male). Conversely, patients with genotype 3 infection were more frequently male, and the prevalence in Italian females was significantly lower than in non-Italian females (p=0.0385). No differences were found between Italian and non-Italian men and between Italian and non-Italian females in patients with genotype 4 infection. A detailed description is reported in Table 1 and Supplementary Fig. (1a and Fig. 1b).
3.2. Age Distribution in the Overall Population
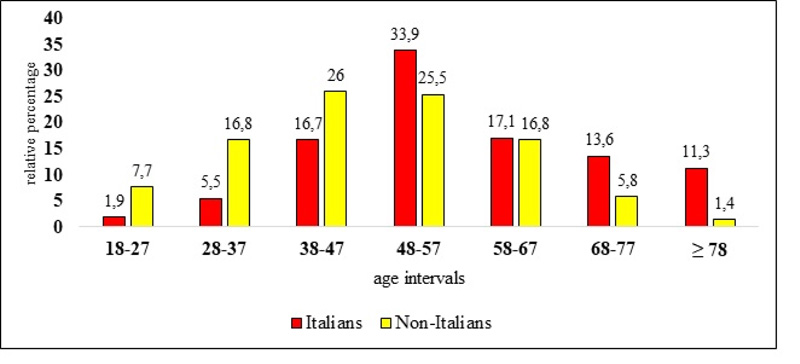
Overall, most Italians (67.7%) and non-Italians (68.3%) were aged 38-67 years; Italians were more represented in class age 48-57 years (p=0.0138), 68-77 years (p=0.001) and in the cohort of patients aged 78 year or more (p<0.0001), and less numerous in class ages 18-27 years (p<0.0001), 28-37 years (p<0.0001) and 38-47 years (p=0.0007) (Fig. 1).
The oldest patients belonged to genotype 2 (both male and female) in the group of Italian patients and to genotype 2 and genotype 1 (female in both cases) in the group of non-Italian subjects; the youngest were all male (non-Italian genotype 1 and genotype 2, Italian genotype 3). Females were significantly older than men in Italian (p<0.0001) and non-Italian patient groups (p=0.0026) with genotype 1 and in Italians with genotype 2 infection (p=0.0036). A detailed description is reported in Supplementary Table 1 and Supplementary Fig. 2.
| Italian Patients | Non-Italian Patients | |||||||
|---|---|---|---|---|---|---|---|---|
| Total (2168 patients) |
Male (1254 patients) |
Female (914 patients) |
p | Total (208 patients) |
Male (96 patients) |
Female 112 patients) |
p | |
| Genotype 1, n (%) | 1195 (55.1) |
680 (54.2) |
515 (56.3) |
0.3273 | 139 (66.8) |
54 (56.3) |
85 (75.9) |
0.0028 |
| Genotype 2, n (%) | 445 (20.5) |
173 (13.8) |
272 (29.8) |
< 0.0001 | 12 (5.8) |
6 (6.2) |
6 (5.3) |
0.7836 |
| Genotype 3, n (%) | 381 (17.6) |
292 (23.3) |
89 (9.7) |
< 0.0001 | 45 (21.6) |
27 (28.1) |
18 (16.1) |
0.0358 |
| Genotype 4, n (%) | 147 (6.8) |
109 (8.7) |
38 (4.2) |
< 0.0001 | 12 (5.8) |
9 (9.4) |
3 (2.7) |
0.069536 |
3.3. Genotypes Distribution before and after the Application of Wider Access Criteria to HCV Treatment
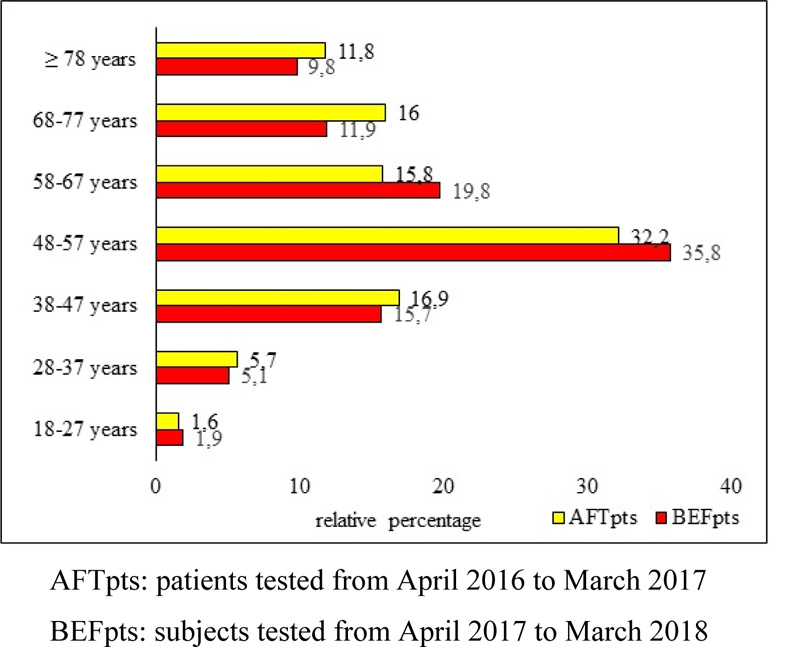
The number of Italian BEFpts and Italian AFTpts was comparable (784 subjects and 864 subjects, respectively); only Italian patients were studied because of the low number of non-Italian patients (74 BEFpts and 82 AFTpts). The percentage of male in BEFpts and AFTs was comparable (58.5% and 54.9%, respectively). Analysis by age showed that patients aged 58-67 years were less represented in AFTpts than in BEFpts (15.8% vs. 19.8%, p=0.0378) and, conversely, patients aged 68-77 years were more numerous in AFTs (16% vs. 11.9%, p=0.0164) (Fig. 2).
|
BEFpts (784 pts) n (%) |
AFTpts (864 pts) n (%) |
p | |
|---|---|---|---|
| Genotype 1 (all pts) | 455 (58) | 462 (53.5) | 0.0626 |
| Genotype 1 M | 261 (33.3) | 242 (28) | 0.0201 |
| Genotype 1 F | 194 (24.7) | 220 (25.5) | 0.7372 |
| Genotype 2 (all pts) | 154 (19.6) | 186 (21.5) | 0.3451 |
| Genotype 2 M | 66 (8.4) | 65 (7.5) | 0.5024 |
| Genotype 2 F | 88 (11.2) | 121 (14) | 0.0904 |
| Genotype 3 (all pts) | 127 (16.2) | 158 (18.3) | 0.2631 |
| Genotype 3 M | 100 (12.8) | 124 (14.4) | 0.3450 |
| Genotype 3 F | 27 (3.5) | 34 (3.9) | 0.5979 |
| Genotype 4 (all pts) | 48 (6.1) | 58 (6.7) | 0.6256 |
| Genotype 4 M | 32 (4.1) | 43 (5) | 0.3840 |
| Genotype 4 F | 16 (2) | 15 (1.7) | 0.6494 |
pts: Patients
F: Female
M: Male
Overall analysis showed no significant difference between genotype frequencies in the 2 study periods, even if genotype 1 approached the significance (58% vs. 53.5%, p=0.0626). Male patients with genotype 1 were less represented in the AFTpts cohort than in BEFpts cohort (28% versus 33.3%, p=0.0201), but median age was the same (53 years). Conversely, male AFTpts with genotype 3 infection were younger (median age 49 years versus 51 years, p=0.0262), but the frequency was similar over time (14.4% in AFTpts and 12.8% in BEFpts). No significant difference was found in female subjects. A detailed description is reported in Table 2 and Supplementary Table 2.
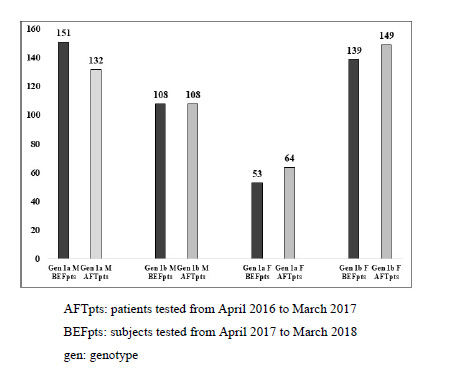
Genotype 1 subtyping was available for 904 out of 917 subjects (98.6%): overall, 1b subtype was the most frequently detected, and subtypes distribution pattern was similar in BEFpts and AFTpts. A complete description is reported in (Fig. 3). Females with subtype 1a infection were older in the AFTpts group (median age 51 years, IQR 44-56 years, versus 47 years, IQR 40-52 years, p=0.0105).
4. DISCUSSION
In this study, we have reported all HCV genotypes assessed in a regional hub center of Northern Italy from January 2016 to December 2018, that is, a description of HCV epidemiology of patients with known infection after one year of DAAs treatment availability in a low prevalence area.
In Italy, HCV infection prevalence is higher than in the other European Western countries, but it varies according to the geographical area; Southern regions have an anti-HCV patients frequency ranging from 2.4% to 7.5% [16, 21-23], while a value of about 1.5% has been found in the North of Italy [16, 24].
Many studies have evaluated HCV genotype changes in frequency over time before the availability of DAA.
Petruzziello et al. [25] compared the prevalence of HCV genotypes detected in 2012–2014 with those identified in 2006–2008 and 2009–2011 in Southern Italy regions; these authors reported an increase in genotype 2 and a decrease in genotype 3a, while genotype 1b subtype was the predominant in the three periods, but the first DAA was reimbursed by the Italian National Health Service in December of 2014 and other drugs of the DAA class from 2015 onwards [26]. A longer interval was chosen by Sagnelli et al. [27], who described the characteristics of two cohorts of patients referring to Italian liver units in 2001 and 2014, when only IFN-based treatments were available; HCV-genotype 4 frequency increased over time while the numerosity of patients with genotype 2 infection decreased.
The data available on HCV genotype distribution in Northern Italy is less. De Conto et al. [28] enrolled a lower number of subjects with respect to our work (1265 versus 2376) during a five years hospital-based surveillance in the Parma area; the relative genotype frequencies were comparable, even if De Conto’s work study years were 2009-2013, an interval in which only interferon-based treatments were available in routine clinical practice [29]. Furthermore, these authors performed an overall analysis including both Italians and non-Italians, who constituted 9.7% of the study population (8.8% in our work); we observed some differences in genotype prevalence distribution and in subjects' median age between Italian and non-Italian cohorts, so we believe that a separate analysis could provide a more reliable picture of known HCV viremic subjects to apply for a screening program.
The first result of our study was that most Italian patients aged from 38 to 67 years, with a wide age range genotype-related; the youngest were patients with genotype 3 infection (median age 50 years in male and female), and the older were females bearing genotype 2 (median age 72 years). Patients’ ages have been analyzed with different approaches in previous Italian studies; 74.1% of individuals with more frequent HCV genotypes/subtypes were older than 41 years in the study by De Conto et al. [28], 76.6% of patients aged from 35 to 74 years in the work of Stasi et al. [30], who enrolled subjects referring to hepatology units in Tuscany from January 1st, 2015, to December 31st, 2015; Buonomo et al. [31] found that 54.8% of known HCV+ Naples’ patients who visited between January 2016 and March 2017 were >60 years old and the HCV Network Sicily [32] reported that the mean age of subjects registered on the web platform was 61 years and 34% of the subjects were >70 years. Taken together, the data seem to describe an HCV infected population aged similarly to ours: class age 58-67 years is less represented and class age 68-77 years more represented (16% vs. 11.9%) in Italian patients tested the year after the application of wider access criteria to HCV treatment with respect to those tested the year before (15.8% vs. 19.8%, respectively). We are not sure whether all genotyped patients underwent antiviral therapy; however, these last results suggest that the burden of expected HCV known viremic infected subjects is getting smaller under the pressure of an extensive access to antiviral treatment and that the increase in older patients was due to the inclusion of subjects with lower fibrosis degree, given the efficacy and safety of DAAs [33, 34]. The low and comparable frequency of genotyped patients aged less than 37 years both in AFTpts and in BEFpts possibly reflected the true epidemiological picture of HCV infection in Italy even if our work was not a population-based study; most patients with acute HCV infection aged 35-54 years but the number was extremely low [35], so probably these young subjects had a chronic disease recently diagnosed or recently linked to care. HCV genotype determination is a crucial pretreatment assessment, and the prescription to use an assay testing for two genomic regions at a time for an accurate genotyping/subtyping was already included in EASL Clinical Practice Guidelines published in 2011 [36], so in theory, previously identified subjects were not retested.
In our study, the youngest Italian patients had a genotype 3 infection; the relationship between young age and genotype 3 infection has been commonly assessed in previous studies on Italian patients [25, 28, 31, 37], and can be related to drug use as a mode of HCV acquisition. However, the overall relative frequency of genotype 3 we found (17.6%) was higher than those reported in the other studies involving Southern regions and ranging from 3.2% to 6.5% [25, 31, 37] but comparable to that describing epidemiological pattern in Parma (20.1%) [28]. This different epidemiological situation has clinical importance not only for the immediate therapeutic approach but also for the future health scenario because a higher percentage of genotype 3 subjects may predict a higher number of antiviral treatment failures [38, 39]. Differently from genotype 3, genotype 2 detection was associated with a more advanced age, and this was an expected finding [17]; the 75 percentile value was 79 years for female and 77 years for male, implying that there is a subgroup of very old patients. SVR rates were high in old patients [40, 41]; no age limit contraindicating antiviral treatment was established, and the only contraindication to therapy reported in European guidelines is the limited life expectancy because of non-liver-related comorbidities [12]. However, these patients can have comorbidities needing polypharmacy, and this implies the risk of drug-drug interactions, making advisable a collaborative network with general practitioners in the light of unchanged prevalence and the median age of genotype 2 subjects over time.
The unchanged genotypes prevalence of the study population before and after the wider availability of antiviral treatment is the second main result of our study; AFTpts could have a less severe disease but the same epidemiological root as BEFpts. This hypothesis is in accord with the comparable value of patient’s age for a specific genotype, with the only exception of males with genotype 3 infection (AFTpts were younger, 49 years vs. 51 years). This data may be possibly due to the persistent risk of HCV infection in drug users, as described by Spada et al. [42], who reported an overall HCV incidence as 5.83/100 person-years at risk in a prospective cohort study among drug users attending 17 Italian rehabilitation centers serving urban areas; on the other hand, prevalence and patients’ age suggest an increased access to treatment of this high-risk population. A notable finding is the confirmed frequency of genotype 2 infection at values of about 20%, lower than the pooled mean proportion of 27% reported in a systematic review of 43 Italian studies published from 2000 to 2017 and including 23,041 subjects [17]; the decrease in this calculated frequency could have been more evident in the light of SVR rates obtained with pegylated interferon and ribavirin [43, 44]. Italy has a peculiar diffusion of genotype 2 infection, with a higher prevalence with respect to European [45] and most non-European countries [1]. Published data suggest a transmission related to healthcare practices [46-48], and it has to be underlined that glass syringes use was dismissed in the late 1970s and that the NAT screening for HCV became mandatory for blood products donors in June 2002 [49]. Persistence of about 20% of patients with genotype 2, a historical “easy-to-threat” genotype, could suggest a great burden in older people or a persistent iatrogenic transmission or both; of note, Marascio et al. [50] reported a genotype 2 prevalence of 20.2% in patients presenting for testing in a teaching hospital of the Calabria Region in the study period 2008-2018, a value comparable to ours.
Finally, we had the opportunity to report HCV genotype distribution in non-Italian patients already linked to care; they had an overall higher frequency of genotype 1 infection and a lower one of subjects bearing a genotype 2 virus. Both findings are in accord with the previously published data [45], while it is interesting to note the higher diffusion of genotype 1 in females (75.9% vs. 56.3%), who were also older than males (52 years vs. 41 years) with respect to Italian patients. Non-Italian patients refer to patients included from countries having different rates of HCV prevalence and specific genotype frequencies, so it is hard to discuss the possible epidemiological clues. In our opinion, it is important to underline the differences in genotype distributions that can suggest a compartmentalized disease diffusion, as previously reported for HIV infection [51,52].
Our study involves three main limitations. First, it is a retrospective single-center study, and it is possible that HCV genotypic distribution is different in different parts of Italy; on the other hand, the number of patients tested was very high, and all of them were included, so no selection bias may have occurred. Second, non-Italian patients were analyzed as a single cohort and not by ethnicity because of the low number and, for the same reason, they were not included in the comparative analysis of genotype distribution before and after the application of wider access criteria to HCV treatment; moreover, our choice did not allow any comment on the possible influence of geographic origin on sensitivity to infection, as also reported for other diseases [53-55]. Third, no clinical data were recorded, and consequently, we could only speculate the possible risk factors for HCV infection in our patients. However, the data on age, gender and place of birth are easily available from electronic databases of laboratories, while clinical data may be included in medical charts; this characteristic made it more difficult to obtain data of a high number of subjects in a short time that may help organize a tailored screening. In our opinion, the knowledge of HCV genotype local epidemiology could be integrated with the known risk factors as a first step.
CONCLUSION
In conclusion, our study showed the genotype distribution to be different according to the geographical area in the same country and in the same area between Italians and non-Italians (who are younger) and, as the second report, the absence of modification in genotype prevalence in Italian patients after one year of wide antiviral treatment prescription. Taken together, these findings underline the importance of discussing HCV elimination approaches, starting from a picture of genotypes detected, which address the identification of risk groups, due to possibility of the main different transmission routes.
LIST OF ABBREVIATIONS
| AFT | = After Period |
| AFTpts | = After Patients |
| BEF | = Before Period |
| BEFpts | = Before Patients |
| DAAs | = Direct-Acting Antivirals |
| HCV | = hepatitis C Virus |
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study has been approved by the Ethics Committee of Padova University Hospital (prot 61946).
HUMAN AND ANIMAL RIGHTS
No animals were used in this research. All human research procedures followed were in accordance with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Informed consent was obtained from all the participants.
STANDARDS OF REPORTING
STROBE guideline was followed in this study.
AVAILABILITY OF DATA AND MATERIALS
Data supporting the findings of this research are available within the article.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest with respect to the publication of this article.
ACKNOWLEDGEMENTS
Declared none.
SUPPLEMENTARY MATERIAL
Supplementary material is available on the publisher’s website.


