All published articles of this journal are available on ScienceDirect.
Current and Emerging Technologies for the Diagnosis of SARS-CoV-2
Abstract
Currently, there are numerous under development or developed assays with various sensitivities and specificities for diagnosis of the Coronavirus Disease 2019 (COVID-19) caused by the SARS-CoV-2 virus. The World Health Organization (WHO) has approved several detection protocols based on real-time reverse transcription PCR (RT-qPCR) and the reliability of tests to detect the N, S, or RdRp/Hel genes of the SARS-Cov-2 virus has also investigated. Among these targets, COVID-19-RdRp/Hel targets represented the highest sensitivity. Reverse transcription loop-mediated isothermal amplification (RT-LAMP) has also been developed to rapidly and efficiently amplify RNA under isothermal conditions. Other isothermal amplification approaches such as nucleic acid sequence-based amplification (NASBA), recombinase polymerase amplification (RPA), and rolling circle amplification (RCA) have also been reported for detecting coronaviruses but like LAMP assay. Different serological tests, including neutralization tests, immunofluorescent (IFA), enzyme-linked immunosorbent (ELISA), and western blotting assays, are available. Point-of-care tests (POCT) are emerging to detect the virus genome, IgG, or IgM antibodies against SARS-CoV-2. The advent of more sensitive, cheaper, and easier-to-perform diagnostic tests seems to be a fundamental prerequisite to improve the diagnosis of COVID-19 infection. Herein, we reviewed several commercially available diagnostic methods used in many clinical laboratories to detect COVID-19.
1. INTRODUCTION
Coronavirus Disease 2019 (COVID-19) was first described in Hubei, China and then quickly spread worldwide [1]. The patients present with a range of symptoms from mild to severe pneumonia. In the primary phases of disease, lymphopenia, leukopenia, elevated C-Reactive Protein (CRP) and erythrocyte sedimentation rate (ESR) are the most frequent laboratory findings [2-5]. Chest Computed Tomography (CT) is a major tool for diagnosis of COVID-19 and the most frequent finding in chest CT is Ground-Glass Opacification/Opacity (GGO) and consolidation. In addition, interlobular septal thickening in both lungs has been reported less frequent. Management of COVID-19 depends on prompt and precise detection of the virus using sensitive diagnostic assays. The symptoms of COVID-19 infection are not specific and they cannot be used for a precise diagnosis. Therefore, Nucleic acid–based method and CT scans have been widely used for detection of SARS-CoV-2 infection. Molecular assays are more appropriate than other diagnosis methods such as symptomatic diagnosis and CT scans [6]. The SARS-CoV-2 genome was sequenced soon after the initiation of the epidemic and facilitated development of precise reverse transcription PCR diagnostic tests.
Nasopharyngeal and oropharyngeal swabs as well as sputum, Bronchoalveolar Lavage (BAL) fluid and tracheal aspirates are the most common specimens used for detection of the virus [7, 8]. The ongoing technological improvements of molecular biology techniques have revolutionized virus-detecting methods [9]. In particular, Polymerase Chain Reaction (PCR), as the “gold standard”, provides a rapid and highly sensitive method for detecting viral genomes [10]. Moreover, non-PCR-based methods including the loop-mediated isothermal amplification (LAMP) have also been developed to detect the RNA of coronaviruses [9]. In the primary course of the epidemic, several nucleic acid detection kits, and a batch of fluorescent quantitative kits and sequencing systems were approved by the China Food and Drug Administration (CFDA) [11]. Current molecular diagnostic tests available for SARS-CoV-2 include real-time RT-PCR, reverse transcription loop-mediated isothermal amplification (RT-LAMP) and RT-PCR (Table 2). However, the last method is usually used for detection of RNA viruses but its application has been abandoned to diagnose COVID19 for the reasons explained in part 1.1 [12].
There are also other nucleic acid-based tests that can be used for detection of virus. SHERLOCK is an approach based on CRISPR-Cas13. Although CRISPR-Cas13 system has not been verified on clinical samples, it has been suggested as a rapid, cheap and easy-to-perform diagnostic test for SARS-CoV-2 detection [6].
Contrary to the nucleic acid tests, protein-based tests can be used for detection of virus after recovery. Antibodies against SARS-CoV-2 protein antigens are generated in response to the viral infection and can be used for the detection of infection. Alteration in the load of the virus during the infection may affect the results of testing [6]. For instance, high viral loads in saliva have been indicated during the first week after initiation of symptoms, which gradually decreased [13]. Therefore, diagnostic serology testing one week after onset of infection has been suggested. Detection of specific antibodies in serum have numerous advantages including easy procedure and availability of samples. Here, we have discussed the molecular and non-molecular assays that are currently available for detecting the novel SARS-CoV-2 virus.
2. EVIDENCE ACQUISITION
The most current evidences about the diagnosis methods for detection of SARS-CoV-2 infection have been provided in the current review. In the initial stage, a systematic search was done in some databases/search engines including ISI Web of Science, Scopus, PubMed, and Google Scholar. In the second stage, all selected documents were reviewed. The keywords used were: severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), COVID-19, diagnosis, PCR, LAMP, serology, coronavirus, and pneumonia.
3. NUCLEIC ACID BASED TECHNOLOGIES
3.1. Reverse Transcription Polymerase Chain Reaction
Reverse transcription polymerase chain reaction (RT-PCR) is one of the simplest virus detection methods and due to its simplicity; it is frequently used in clinical and research laboratories. In this molecular technique, ribonucleic acid (RNA) as genome template initially is converted into complementary deoxyribonucleic acid (cDNA) by reverse transcriptase enzyme and after that in a separate reaction, the cDNA will be amplified by DNA Taq polymerase enzyme leading to production of billions copies of amplicons [14]. Usually three primers are used in RT-PCR procedure including specific, random hexamer and oligo-dT primers. The random hexamer primer (sequence: NNNNNN) used for RT-PCR amplification is annealed to cell's mRNAs and pre-mRNAs and therefore, the synthesized cDNA usually is representative for cell total mRNAs [15]. The oligo-dT primer (T) 18-20n is bond to poly (A) sequence at the 3′ end of mRNAs, amplifies just cell's mRNAs and therefore it can only be used for recognition of cells' mRNAs. Viral RNAs are identified with random hexamer primer and the oligo-dT primer will not work due to lack of poly (A) sequence at the 3′ end of their RNA except some viruses [16, 17].
Although the RT-PCR method is a basic tool for several goals such as evaluation of gene expression, RNA virus detection and gene cloning, however, it has some disadvantages which cause its application to be limited in diagnosis of RNA viruses. These limitations include lack of ability to determine virus infectivity and inhibition by some compounds, leading to false negatives results [18]. According to published literatures, there are only limited reports for detection of COVID19 based on RT-PCR method.
3.2. Real Time RT-PCR
Currently, real time RT-PCR is the reference and standard method for the diagnosis of active COVID-19 infections [19]. This method employs the viral RNA extracted from patients’ samples (e.g., nasopharyngeal/oropharyngeal/nasal swabs, tracheal aspirates, sputum, bronchoalveolar lavage (BAL), pleural fluid, or lung biopsies) to synthesize complementary DNA (cDNA), a reaction catalyzed by the reverse transcription (RT) enzyme, which then serves as a template to amplify target viral genes [19]. The genes encoding nucleocapsid (N), glycoprotein spike (S), RNA polymerase-dependent RNA (RdRp), and Helicase (Hel) are common target genes detected by COVID-19 real time RT-PCR assays [20].
Chan et al. employed a very sensitive, specific and innovative real time RT-PCR method for detecting the RdRp/Hel in the clinical samples of COVID-19 patients. In the recent study, the researchers investigated the reliability of real-time RT-PCR to detect the N, S, or RdRp/Hel genes of the SARS-Cov-2 virus. Of the three mentioned targets, COVID-19-RdRp/Hel assay represented the highest sensitivity by detecting the lowest amount of the virus. Among 273 samples from 15 patients with confirmed COVID-19 infection in Hong Kong, 28.2% were positive for both RdRp-P2 and COVID-19-RdRp/Hel. The COVID-19-RdRp/Hel test rendered positive results in 42 (29 (24.2%) respiratory and 13 (8.5%) non-respiratory) samples that were negative for the RdRd-P2. While the presence of other respiratory pathogens and human coronaviruses in cell culture and clinical samples did not interfere with COVID-19-RdRp/Hel assay, a cross-reactivity phenomenon was observed between the SARS-CoV-2 and RdRp-P2 assays in cell culture samples. This study showed that COVID-19-RdRp/Hel method was very sensitive and specific for detecting COVID-19 infection [20].
Nevertheless, real-time RT-PCR has also been associated with false-negative results in patients with established COVID-19 infection [21]. Li et al. conducted a study on a confirmed case with COVID-19 pneumonia who had several real-time RT-PCR negative tests. In this patient, while the first five real-time RT-PCR tests were negative for COVID-19, the sixth test delivered a positive result. This observation suggested that in the presence of supportive epidemiological evidence and the typical clinical and imaging manifestations of COVID-19 pneumonia, real-time RT-PCR should be serially repeated, and suspected patients must be treated in isolation [21].
In a study by Tao et al., the researchers examined the correlation between real-time RT-PCR results and chest CT scan findings in Chinese patients with COVID-19 infection. They showed that out of 1014 patients, while only 601 (59%) patients showed positive real-time RT-PCR results, 888 (88%) had characteristic chest CT scans. Considering real-time RT-PCR as a reference test, the sensitivity of CT scan for detecting COVID-19 infection was about 97%. In addition, 75% of the patients who had negative real-time RT-PCR results revealed positive chest CT scan findings. So, the results of this study indicated the high sensitivity of chest CT scan for detecting COVID-19 infection [22].
In another study conducted by Lan et al. to examine the applicability of real-time RT-PCR in Chinese patients recovered from COVID-19 infection, the results showed that four COVID-19 carriers who had the criteria of either quarantine discontinuation or hospital discharge (i.e., two consecutive negative real-time RT-PCR results and the lack of clinical symptoms) rendered positive real-time RT-PCR results within 5 to 13 days after the last negative test. The study's findings showed that real-time RT-PCR may also be associated with false negative results in coronavirus carriers [23]. Recently, Kasteren et al. compared several commercial COVID-19 detection kits and found that all of the studied kits are suitable for diagnostics of symptomatic COVID-19 patients [24]. They also recommended real-time RT-PCR kits from R-Biopharm AG, BGI, KH Medical manufacturers when the viral load in the specimen is low. The detection limits of studied kits are shown on Table 1.
Regarding high sensitivity, specificity, and reliability, as well as relatively rapid deliverance of results, it seems reasonable to use this technique to accurately detect the SARS-CoV-2 virus. In comparison with other diagnostic techniques, this method is considerably faster and has a lower potential to be affected by contaminations and errors. However, the test may rarely fail to discover a small ratio of infected people and carriers, so it is better to use other diagnostic methods along with this technique.
3.3. LAMP Method and Other Isothermal Assays
Nucleic acid amplification tests such as real-time reverse-transcription-polymerase chain reaction (RT-qPCR) have been rapidly developed to quantitatively detect the SARS-Cov-2 virus in different samples including nasopharyngeal swabs, sputum, alveolar lavage fluid, and blood. However, RT-qPCR requires fully-equipped laboratories and skilled staff.
Therefore, as RT-qPCR is a time-consuming and laborious process, an alternative specific nucleic acid amplification test, loop-mediated isothermal amplification (LAMP), has been developed to rapidly and efficiently amplify DNA under isothermal conditions [25, 26]. This approach uses a set of three to four specific primers targeting conserved sequences such as ORF 1ab. Furthermore, a DNA polymerase enzyme with chain displacement activity is employed to amplify the target DNA up to 109 copies which is accomplished at a constant temperature between 60 and 65°C in less than an hour. Out of the 4 designed primers in this method, two are “inner primers” (BIP and FIP) which synthesize new DNA strands [26]. The outer primers (B3 and F3) anneal to the template strand allowing the replication of the new DNA strand. Stem-loop DNAs with multiple inverted repeats are produced as the final products of the target DNA. The LAMP assay can be simply and rapidly (i.e. a one-step reaction) performed by minimally trained staff without requiring for well-equipped laboratories [27]. This method can be combined with reverse transcription reaction (i.e. RT-LAMP) to allow the direct detection of RNAs [28]. A multiplex LAMP assay was designed by Kim et al. for detecting SARS-CoV. Compared to real-time RT-PCR, it showed a detection limit of 10e5 RNA copies in each reaction [29]. This assay is a one-step nucleic acid amplification test that is used to diagnose infectious diseases caused by either bacteria or viruses. Similar to RT-qPCR, the RT-LAMP utilizes the reverse transcriptase enzyme to synthesize cDNA from RNA sequences and successfully detects pathogen-specific nucleic acids applying either pH-sensitive or bioluminescent indicators. Collectively, the LAMP technology seems to be a sensitive and stable method to rapidly detect viral pathogens. Accordingly, the method can be considered as a rapid and simple diagnostic test for infections caused by RNA-containing viruses such as the SARS-Cov-2. Nguyen et al. proposed using the LAMP method as a point-of-care test to detect zoonotic viruses [such as avian influenza virus (AIV)] and coronaviruses (such as the SARS-Cov2) [30].
In the recent decades, in addition to LAMP assay, several types of isothermal nucleic acid amplification techniques such as strand displacement amplification(SDA), signal mediated amplification of RNA technology(SMART), rolling circle amplification(RCA), isothermal multiple displacement amplifi- cation(IMDA), single primer isothermal amplification(SPIA), circular helicase-dependent amplification(HDA) and recom- binase polymerase amplification (RPA) have been introduced [31]. The mechanism of these procedures and their applications have been reviewed by Gill et al., comprehensively [32].
The reverse transcriptase-RPA (RT-RPA) method has been recently used to detect bovine coronavirus (BCoV) and the results of the study has shown that BCoV RT-RPA in comparison with real-time RT-PCR was more sensitive with detection limit of 19 copies RNA [33]. The authors of the study suggested that the RT-RPA method could be used in point-of-care diagnosis by applying a transferable fluorescence reading machine.
James et al. believe that RPA and also LAMP techniques have high potential performance to diagnose COVID-19 and it could also apply at community level [34]. It seems that applying the isothermal amplification methods to diagnose COVID-19 needs more researches to evaluate their sensitivities and specificities and furthermore, their reproducibility should be determined.
The RCA method also carries outs under isothermal condition without false-positive results and it needs minimal reagents for its performance. Recently, the method has been used for the detection of SARS-CoV in 7 clinical respiratory specimens and the results showed that the RCA technology was capable to amplify signals up to 109-folds in each amplification cycle [35].
NASBA is another isothermal amplification method for the detection of RNA targets. Three enzymes are involved in this method including T7 RNA polymerase, RNase H, and avian myeloblastosis virus reverse transcriptase. The method is a proven diagnostic tool and has more sensitivity than RT-PCR for detection of pathogens. Therefore, NASBA could be considered as a diagnostic tool for SARS-CoV-2. A study by Wat et al. used NASBA assay to detect respiratory viruses in nasal swabs which were taken from patients with cystic fibrosis. The results showed that out of 138 specimens, 46% samples were positive for respiratory viruses [36].
4. NON- NUCLEIC ACID- BASED TECHNOLOGIES
4.1. Serologic Assays of COVID-19 Infection
The patients infected by viruses develop antiviral antibodies during the acute phase of infections. Generally, IgM antibodies against several viruses become detectable within three days post-infection. The serological tests of COVID-19 infection measure the levels of serum antibodies resulted from the immune reaction against the SARS-COV2 virus. Compared to molecular methods, compatibility with moderate-quality samples and widely-available equipment (i.e., 96-well microplate ELISA automatic reader) is one of the main advantages of serological assays. Moreover, due to the high testing capacity of serological procedures, a larger ratio of febrile patients can be tested within a respectively short time.
The different types of serological tests include neutralization tests, immunofluorescent (IFA), enzyme-linked immunosorbent (ELISAs), and western blotting assays [37]. Using validated serological methods can improve the diagnostic process of COVID-19 infection, especially when there is a strong epidemiological evidence for the infection in the context of negative molecular tests and in both acute and recuperation phases [38].
Serological tests can be easily performed in hospital clinical laboratories, so they are much more widely used than molecular tests. A main target population for serological tests includes asymptomatic patients or those with mild COVID-19 infection who comprise a large portion of patients. This is particularly important as these individuals are not generally referred for molecular tests [38].
Although cross-reactions with other coronaviruses can be challenging, but new sensitive and specific commercial and non-commercial serological tests are being developed to resolve this problem [39, 40]. So far, multiple serological methods have been approved to diagnose COVID-19 infection. These include specific kits for detecting antibodies (IgM or IgG) against the whole virus by ELISA, chemiluminescence, or colloidal gold assays.
In addition to the nucleocapsid phosphoprotein (NP) which provides a sensitive antigen, other epitopes and specific molecules of the SARS-Cov2 virus can also be considered to develop serological assays.
Zhou et al. used the nucleocapsid (N) protein of the SARS-CoV Rp3 recovered from bats to detect IgG and IgM antibodies against the corresponding antigen in the SARS-Cov2 virus. This was due to the two antigens had a 92% similarity in their amino acid sequences. The applied antigen did not show cross-reactivity with antibodies against other human viruses than SARS. They assessed the levels of anti-viral antibodies in one patient at 7, 8, 9 and 18 days after the onset of the disease. There was a significant increasing trend in the titers of IgG and IgM antibodies over time, except that the titer of IgM showed a reduction in the last sample. They also evaluated anti-viral antibodies in samples from 5 out of 7 COVID-19 confirmed patients approximately 20 days after the onset of the disease. All the patients and healthy individuals’ samples rendered positive results for anti-virus IgG antibodies [41].
Shaoli et al. examined the epidemiological history and clinical features of seven patients with the novel coronavirus pneumonia in Lanzhou Pulmonary Hospital. After 7 days of the diagnosis, only one patient had negative results for anti-virus antibodies. This suggested that in patients who have a negative molecular test result but show supportive lung lesions, the screening of COVID-19 specific IgM antibodies can be of a great diagnostic value [42].
In order to identify IgM and IgG against the SARS-CoV-2, Haveri et al. examined serum samples from COVID-19 patients by IFA after 4, 9, 10, and 20 days of the onset of symptoms. While the antibodies were undetectable at the 4th day, IgG titers rose to 80 and 1280, and IgM titers to 80 and 320 on the days of 9th and 20th, respectively [43].
In comparison with molecular methods, a study in China reported that the sensitivities of serum IgM and IgG antibodies for detecting COVID-19 infection were reported as 48.1% and 88.9%, respectively. Also, the specificities of these antibodies were 100% and 90.9%, respectively. The titer of IgM and its positivity rate showed no significant differences before and after recuperation [44]. The positivity rate of IgG was more than 90% showing no significant difference before and after recuperation.
To date, the FDA has approved only one serological test called the qSARS-CoV-2 IgG/IgM Rapid Test (Cellex Co.). This test which is planned to be used in clinical laboratories in the near future is based on the qualitative detection of IgM and IgG anti-SARS-CoV-2 antibodies in the plasma or serum samples of suspected COVID-19 patients [45].
4.2. Enzyme –linked Immunosorbent Assay (ELISA)
Recently, Zhang et al have developed an ELISA assay for detection of IgG and IgM against SARS-CoV-2. The SARS-CoV-2 Rp3 nucleocapsid protein has been used in this assay. This protein has 90% amino acid sequence similarity to the other SARS-related viruses. They studied the samples of sixteen SARS-CoV-2 positive cases whose infection were confirmed by RT-PCR and reported the increase levels of these antibodies during the first 5 days after initiation of the symptom. It has been reported that antibodies were detectible in blood, respiratory, as well as fecal samples [46]. Applying two ELISA kits based on recombinant nucleocapsid protein (rN) and spike protein (rS) for detecting antibodies in COVID-19 patient’s sera showed that the sensitivity of the rS-based ELISA was significantly higher than that of the rN-based ELISA [47]
4.3. Western Blotting
Western blotting is a precise technique used in molecular biology laboratories to identify proteins. In this technique, proteins within a mixture are initially separated through SDS-gel electrophoresis based on their molecular weights. In the next step, proteins resided on the gel are transferred on specific membranes (such as PVDF). Finally, the presence or absence of a target protein is confirmed through incubation of the membrane with a target-specific antibody [48-50].
In virology, western blot technique is widely used to identify viruses [51], and anti-viral antibodies [52, 53]. This technique can also serve as a confirmatory test to detect viral proteins or genes [54].
Generally, the applicability of western blot techniques depends on the availability of specific antibodies against target proteins. In fact, it is inapplicable to produce specific antibodies for all, especially unknown or unimportant, proteins. Unlike PCR-based methods, western blot techniques are relatively expensive limiting their wide application in clinical and research laboratories.
Despite these limitations, some advantages of western blot methods render them popular in many virology experimental fields. A western blot method has recently been developed to diagnose the coronavirus causing severe acute respiratory syndrome (SARS). In this study, a C-terminal fragment from the nucleocapsid (N) protein of the virus confirmed to have no cross-reactivity with the proteins derived from canine, pig, and chicken coronaviruses was used as a target to detect anti-SARS antibodies. The sensitivity and specificity of the method were obtained as 90.9% and 98.3%, respectively. The results of this western blot assay performed on 274 clinical samples showed strong correlations with IFA findings [55]. Nonetheless, a main disadvantage of western blot is that antibodies against pathogenic agents are not detectable in early or lag phases of infections. Generally, it takes at least 7 days for the body to develop antibodies against infectious agents or coronaviruses [56, 57].
The nucleocapsid (N) protein, as a highly conserved peptide among coronaviruses, seems to be an appropriate target for detecting these viruses in clinical samples. However, the extent of the humoral immune response against the various regions of this protein is variable. One study on human sera using rabbit polyclonal antibody described a great immunogenicity for the central region (amino acids of 174 to 300) of HCoV-OC43 nucleocapsid protein. On the other hand, variable immune responses were observed against the C or N-terminal domains of this protein in the course of HCoV-OC43 infection [58].
It seems that the nucleocapsid protein of coronaviruses is an acceptable target to be used in western blot techniques to diagnose infections by these viruses. For the COVID-19 infections; however, it is suggested to recruit a region with no or least cross-reactivity with proteins from other coronaviruses.
5. LABORATORY PARAMETERS
In the first stages of infection, peripheral blood lymphopenia is common. In addition, increased level of C-reactive protein (CRP) and erythrocyte sedimentation rate (ESR) is seen [59]. It has been also shown that the levels of IL-1β, IL-7, IL-8, IL-9, IL-10, IFNγ, and TNF-α are elevated, suggesting the possible role of immunopathology in COVID-19 infection [60]. A recent study has assessed the immune-inflammatory parameters of non-severe and severe patients [61]. They have reported the level of IL-6, CRP and hypertension as independent risk factors for evaluating the severity of SARS-CoV-2 infection.
6. COMPUTED TOMOGRAPHY
Chest computed tomography (CT) scan is a non-invasive diagnostic technique which uses X-ray and computer technology to produce images for detection of lung abnormalities. This technique can be used for clinical diagnosis of COVID-19. There are various characteristics of imaging for COVID-19 depending on the severity of the disease. However, the most common findings of chest CT include bilateral and peripheral ground-glass opacities (GGO) and lung consolidation [62, 63]. Numerous retrospective studies have suggested higher sensitivity in comparison with the RT-PCR. However, due to the similarity of imaging characteristics to that of other viral pneumonia its specificity is low. Since the shortage of PCR reagent kits and lack of availability of molecular tests, chest CT can be the alternative diagnostic method. However, this method has also some drawbacks including the necessity of expensive systems and expert staffs [64-66].
7. POINT OF CARE METHODS
Point-of-care tests (POCT) are medical diagnostic testing without the need to send samples to diagnostic laboratory. Antibody testing for SARS-CoV-2 recognized as lateral flow assay (LFA) is one of these techniques intending to detect the IgG or IgM antibodies against lateral flow antigen of SARS-CoV-2 [67]. The commercial LFA is based on a paper covered with capture antibodies and gold nanoparticle-antibody. Blood or urine sample is placed on the on the membrane and the analyte moves across the paper by capillary action until it passes the first line and interacts with the gold nanoparticle-antibody. This complex continues till reaches the second line. At this time, it is stopped by the capture antibodies and a colored line (blue/red) appears. Clinical specificity, sensitivity, and accuracy of LFA for IgM has been reported to be 100, 57%, and 69% and for IgG 100%, 81%, and 86%, respectively. The sensitivity of LFA which detects both IgM and IgG together is 82% [67].
8. NOVEL EMERGING TECHNOLOGIES FOR DETECTING SARS-COV-2
The microarray or biochip is a method used to determine many targets in clinical specimens. The technique used a microscopic slide that is usually made of glass, silicon chip, or nylon membrane. There are many microarrays types based on their targets including DNA microarrays, protein microarrays, carbohydrate arrays, cellular microarrays, tissue microarrays, chemical compound microarrays, antibody microarrays, phenotype microarrays and etc. Briefly the following steps are used to do a microarray from the coronavirus: 1. RNA extraction and measurement its quality and quantity, 2. converting RNA to labeled cDNA through reverse transcription method with specific dyes and primers, 3. hybridization of labeled cDNAs with fixed probes on the wells of microarray chip, 4. washing to remove unbound DNAs, and 5. detection of COVID-19 genome.
Although the microarray tools benefits from a novel methodology, there is not applied to detect COVID-19 yet. It seems that due to high prevalence of COVID-19, the application of the method will initiate early and its diagnostic kits will be commercialized. There are limited studies about application of microarray method for genotyping and diagnosis of viruses other than COVID-19. Recently, a DNA microarray method has been applied to detect some pathogenic viruses [68]. In this study, a set of 70 bp oligonucleotide probes have designed for more than 140 sequence viral genomes to simultaneously detect all respiratory tract and other viruses. However, there was a limitation for viruses without determined sequence in GenBank database. The microarray method has been also applied to identify single nucleotide polymorphism (SNP) among spike gene of SARS coronaviruses. The results of this research showed that all of 19 virus samples well genotyped using microarray with 100% accuracy [69]. These data conclude that the microarray is an accurate way for the detection and epidemiological surveillance of Coronavirus and it can be applied to detect and genotype COVID-19 in future investigations.
Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) is a well-recognized gene-editing technology which is currently used for the molecular diagnosis of various viral infections (2-4). As parts of the adaptive immunity in bacteria, CRISPR-Cas systems employ RNA-guided nucleases cleaving invading nucleic acids. The class 2 CRISPR-Cas systems are efficient tools in which an effector protein binds to the RNA component leading to the cleavage of a target sequence. In fact, these systems have unique features that can revolutionize medicinal diagnostic methods [70-72].
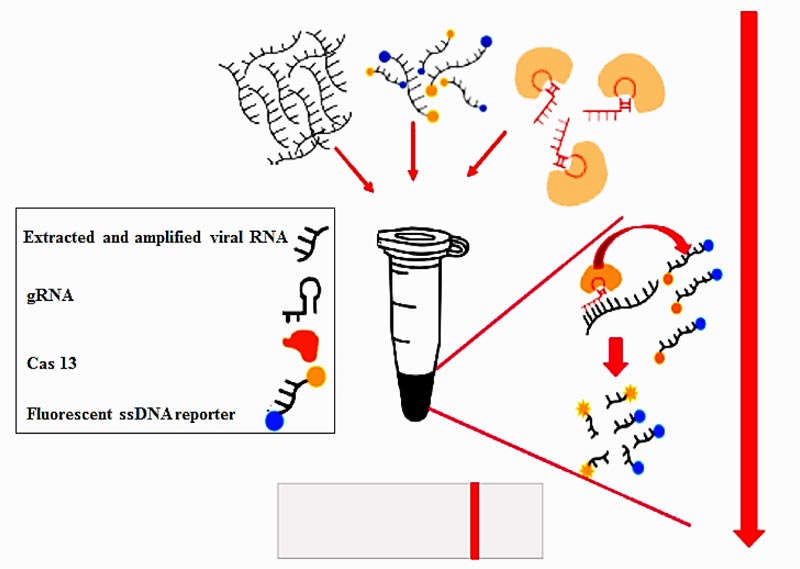
The CRISPR-Cas13 system has been developed as a rapid method for detecting RNA sequences (it schematically represented on Fig. 1). There are two CRISPR-based methods for detection of viral RNA. The first is according to the SHERLOCK [73] protocol, in which viral RNA is converted to dsDNA by RT-RPA and complementary RNA is produced from dsDNA template via T7 transcription. Subsequently, Cas13–tracrRNA complex binds to the target sequence, leading to activation of Cas13 to cleave the targeted sequence as well as the fluorescent RNA reporter. The second method is according to the DETECTR [74] assay in which viral RNA is converted to dsDNA by RT-RPA and Cas12a–tracrRNA complex binds to the target sequence leading to activation of nuclease enzyme activity of Cas12a to cleave the target sequence and the fluorescent RNA reporter.
| Kit | Country | Target gene(s) | Detection limit (copy/ml) |
|---|---|---|---|
| CerTest Biotec | Spain | ORF1ab, N | 18 |
| R-Biopharm | Germany | E | 4.3 |
| KH Medical | Korea | RdRp, S | 4.8 |
| Altona Diagnostics, | Germany | E, S | 3.8 |
| BGI | China | ORF1ab | 4.3 |
| PrimerDesign | England | RdRp | 23 |
| Abbreviations: ORF1ab, open reading frame; N, nucleocapsid protein; E, envelope protein; RdRp, RNA-dependent RNA polymerase; S, spike protein. | |||
| Platforms | Analyte | Testing Sample | Technology | Mechanism | Detectable Time | Expecting Sensitivity | Expecting Specificity | Reference |
|---|---|---|---|---|---|---|---|---|
| RT-PCR | -Nasopharyngeal swabs -BAL -Sputum -Blood -Feces |
RT-PCR | Detection of viral conserved region (D, S, and RdRP genes) | All stages of infection | >95 percent | >95 percent | [6] | |
| LAMP | Nucleic acid | -vector harboring virus sequence | LAMP | isothermal amplification of DNA | - | - | - | [29] |
| RPA | Nucleic acid | -Nasal Swabs -Fecal Swabs |
RPA | RPA | All stages of infection | >95 percent | >95 percent | [33] |
| NASBA | Nucleic acid | Nasal Swabs | real-time NASBA |
isothermal technique for amplification of RNA | All stages of infection | >95 percent | >95 percent | [36] |
| CRISPR | Nucleic acid | -Serum -Nasopharyngeal Swabs |
- RPA - RT-RPA |
- RPA, SHERLOCK protocol - Detection of fluorescence signal |
All stages of infection | >95 percent | >95 percent | [23, 39] |
| ELISA | rN protein rS protein |
Serum | ELISA | - Detection of IgG and IgM antibodies against SARS-CoV-2 | From 1-4 weeks after onset of symptoms | >95 percent | Not applicable | [47] |
| RPA: reverse transcription recombinase polymerase amplification; RT-RPA: Real-time reverse transcription recombinase polymerase amplification; LAMP: Loop-mediated isothermal amplification; NASBA: Nucleic Acid Sequence-Based Amplification; CRISPR: Clustered Regularly Interspaced Short Palindromic Repeats; ELISA: enzyme-linked immunosorbent assay | ||||||||
Zhang et al. recently released a method for identifying the SARS-Cov2 virus using the CRISPR system (https://broad.io/sherlockprotocol). The CRISPR-Cas12 is one of these RNA-guided DNases that non-specifically recognizes and cleaves single-stranded DNAs (ssDNA). The binding of the cas12a to a target DNA within a sample can be combined with the emission of a fluorescent signal by using a reporter molecule to produce a visual record on a paper strip [75]. Therefore, technologies based on the CRISPR-Cas12 system can provide in situ diagnostic tools for the infection caused by the novel coronavirus, SARS-CoV-2. In a recent study, the synthetic fragments of the SARS-CoV-2 genome corresponding to ORF1ab, ORF1b, and RdRp genes [blasted against the WH-human1 sequence (GenBank MN908947) as the reference sequence] were used as DNA templates. The CRISPR-detection complex was generated using commercially available LbCas12a endonuclease (NEB) and an in-house synthesized single guide RNA (sgRNA). Also, ssDNA reporters labeled with fluorescein amidites (FAM) were incorporated into the reaction mixture. In order to prepare samples, saliva specimens from healthy persons were incubated with the RNA fragments of the SARS-CoV-2 virus. The optimal limit of detection (LOD) results was related to the ORF1ab sequence retrieving up to10 copies/μl. This LOD was lower than the minimum level required detecting the virus in clinical samples. Overall, the CRISPR Cas12a system presents a fast, accurate, low-cost, and portable method for identifying the SARS-CoV-2 virus in clinical specimens. However, the validity of this system should be verified in studies on COVID-19 patients’ clinical samples [75].
Proper and rapid detection of COVID-19 infection leads to early management of this disease and the reduction in its mortality rate. However, further studies are needed for a better understanding of this virus and finding effective strategies for early diagnosis [76-78]. Moreover the integration of new emerging sciences is an important issue to fight COVID-19 disease more efficiently in the future [79].
CONCLUSION
In conclusion, COVID-19 is a novel-emerged respiratory infection in human without specific treatments. There are several diagnostic methods to detect SARS-CoV-2 in clinical specimens with different sensitivity and specificity. Currently, the majority of the diagnostic procedures are based on qRT-PCR method which requires both longer sample preparation and analysis times. Isothermal amplification methods have though more variety, but the LAMP, RCA and NASBA have been only applied for detecting coronaviruses and more investigations are required for their application as commercial kit to detect SARS-CoV-2 virus. Serological assays have been suggested for the diagnosis and clinical management of COVID-19 infection and it seems that the serological assays are useful to widely screen suspected cases and evaluate the disease course. Finally, new CRISPR-based nucleic acid detection procedures provide highly sensitive, specific, rapid, affordable, and point-of-care diagnostic technologies to detect the SARS-CoV-2 virus. The development of portable tools including lyophilized reagents and microfluidic-based cartridges can escalate point-of-care testing rate at homes or in small local clinics which can help to effectively confine the rapid spread of the COVID-19 infection. Currently, there in a need for development of point-of-care tests including barcoding, isothermal amplification, and microfluidic tools to tackle the challenges of COVID-19 management.
LIST OF ABBREVIATIONS
| COVID-19 | = Coronavirus Disease 2019 |
| CT | = Computed tomography |
| CRP | = C-reactive protein |
| CRISPR | = Clustered Regularly Interspaced Short Palindromic Repeats |
| ELISA | = Enzyme-linked immunosorbent assay |
| GGO | = Ground glass opacification/opacity |
| HAD | = Helicase-Dependent Amplification |
| IFA | = Immunofluorescent assay |
| IMDA | = Isothermal multiple displacement amplification |
| LAMP | = Loop-mediated isothermal amplification |
| LFA | = Lateral Flow Assay |
| LOD | = Limit Of Detection |
| NASBA | = Nucleic acid sequence-based amplification |
| PCR | = Polymerase chain reaction |
| POCT | = Point of care tests |
| RCA | = Rolling circle amplification |
| RPA | = Recombinase polymerase amplification |
| RT-qPCR | = Real-time reverse transcription PCR |
| SARS | = Severe acute respiratory syndrome |
| SDA | = Strand Displacement Amplification |
| SMART | = Signal mediated amplification of RNA technology |
| SNP | = Single Nucleotide Polymorphism |
| SPIA | = Single primer isothermal amplification |
| WHO | = World Health Organization |
AUTHORS’ CONTRIBUTION
R. R. designed the study. D. A., S. O., S. H., F. S., AND M. A contributed to data gathering and paper writing. All authors revised and approved the final draft.
CONSENT FOR PUBLICATION
Not applicable
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest.
ACKNOWLEDGEMENTS
The authors would like to thank the Clinical Research Development Unit of Baqiyatallah Hospital, Tehran, Iran, for guidance and advice.


