All published articles of this journal are available on ScienceDirect.
Production of Poly-γ-Glutamic Acid (γ-PGA) by Clinical Isolates of Staphylococcus Epidermidis
Abstract
Background and Objective:
Poly-γ-glutamic acid (γ-PGA) is a constituent of the Bacillus anthracis capsule and a potential virulence factor of S. epidermidis. In this study, a methodology for the isolation, purification and quantification of γ-PGA in the isolates was adapted. In addition, the fate of the produced γ-PGA and its antiphagocytic activity were investigated.
Methods:
The capB gene was investigated by the PCR method in 50 isolates of S. epidermidis. A modified methodology was used for the extraction, purification, and quantification of γ-PGA using Cetyltrimethylammonium Bromide (CTAB) solution. The fate of γ-PGA was determined in Tryptic Soy Broth (TSB) medium, as well as the effect of ethanol, NaCl and KCl on the induction of the polymer production. The ability of neutrophils to phagocyte both FITC-labeled latex particles in the presence of free γ-PGA and S. epidermidis with and without anchored γ-PGA was evaluated by cytometry.
Results:
The production of γ-PGA was detected in 40 isolates; all of them were capB gene carriers. Free γ-PGA was detected and in the strain, the amount of released γ-PGA in the supernatant was 67% greater than the cell anchored γ-PGA. Phagocytosis tests performed with one γ-PGA producer isolate showed a significant reduction in neutrophil internalization.
Conclusion:
The adapted methodology was able to detect γ-PGA in the isolates studied. In addition to being found attached to the cell wall, it was demonstrated in this study that γ-PGA can also be found in the culture supernatant. Free γ-PGA did not determine a reduction in the internalization of latex by neutrophils, but cells with anchored γ-PGA showed significant protection against phagocytosis.
1. INTRODUCTION
S. epidermidis was long considered a typically commensal bacterium in humans. However, in the last decades, this species has been progressively recognized as an important opportunistic pathogen, as it is currently the main cause of healthcare-associated infections, as well as infections associated with implanted medical devices [1]. In line with its opportunistic nature, S. epidermidis has a restricted number of factors and mechanisms that can be correlated with its pathogenicity. This particularity differentiates it from S. aureus, a pathogenic species that has a multiplicity of virulence factors [2].
The first described mechanism related to the pathogenicity of S. epidermidis was the ability to produce biofilm. The main component detected in the matrix of this biofilm was the Polysaccharide Intercellular Adhesin (PIA). Biofilm protects the bacterium against the components of immune defense and antimicrobial drugs and explains the close correlation of S. epidermidis with infections associated with internal medical devices [3]. The awareness of the virulence potential of S. epidermidis was amplified with the detection of the capBCAE operon in this species, which is responsible for the production of poly-γ-glutamic acid (γ-PGA) [4].
γ-PGA is a linear anionic polymer, consisting of residues of glutamic acid connected through amide linkages between α-amino and γ-carboxylic acid groups, being, therefore, resistant to proteases. This polymer may consist of L- or D-glutamic acid residues or both isomers. Initially discovered as part of the capsule of Bacillus anthracis, γ-PGA has since been found in many other bacterial species, one archaea, and one eukaryotic organism [5]. In bacteria, it has been observed that only Gram-positive bacteria produce PGA, particularly the members of the order Bacillales, Bacilli class [6].
In S. epidermidis, γ-PGA production has been demonstrated as a factor that facilitates the growth and survival of the microorganism in the human host. It has been suggested that this polymer appears to provide protection against the high concentration of salt present in its natural environment, the human skin. Thus, high osmolarity could be an inducing factor of PGA production in S. epidermidis. Kocianova [4] showed that in most S. epidermidis strains of his collection, γ-PGA production was higher at elevated NaCl concentration. Furthermore, quantitative real-time PCR analysis using a capB probe and immuno-dot blots demonstrated that S. epidermidis upregulates cap expression in response to high NaCl concentration. However, the effect of osmolarity gradients promoted by salts other than NaCl on γ-PGA production has not been investigated. In addition to the role of PGA in osmoprotection, this polymer has been shown to protect the bacteria from the action of antimicrobial peptides and phagocytosis [4].
γ-PGA was also detected in several other coagulase-negative staphylococci, most of which are also opportunistic pathogens [4, 7]. Unlike B. anthracis, where γ-PGA is composed entirely of residues of D-glutamic acid, in S. epidermidis, this polymer has an approximately equal amount of D- and L-glutamic acid [4]. γ-PGA is released in the extracellular medium in some species of Bacillus, while B. anthracis synthesizes γ-PGA that remains bound to the cell wall as a capsule component that enables immune evasion [8]. In S. epidermidis, γ-PGA was found to be bound to the cell wall (anchored PGA) [4], but its release in the extracellular medium has not been investigated.
In this study, a methodology for the isolation, purification and quantification of γ-PGA in clinical isolates of S. epidermidis was adapted, and the production of γ-PGA was investigated both in the presence and absence of possible inducing agents. Additionally, the fate of the γ-PGA produced, and the ability of neutrophils to phagocyte both PGA-producer S. epidermidis (anchored γ-PGA) and latex beads in the presence of free γ-PGA were determined.
2. MATERIALS AND METHODS
2.1. Bacterial Strains and capB Gene Detection
Fifty clinical isolates of S. epidermidis (46 from blood cultures and 4 from central venous catheters) obtained from infected patients in a hospital in Rio de Janeiro were investigated. Isolate identification was performed with a MicroScan WalkAway-96 system (Dade Behring, Inc.). Definitive identification for S. epidermidis was carried out using Polymerase Chain Reaction (PCR) to detect a species-specific genomic fragment of 705-bp [9]. All the isolates were identified as S. epidermidis. A boiling method was used to extract the DNA of the isolates [10]. After growing the cultures in Tryptic Soy Agar (TSA-10 µL HiMedia), suspensions with turbidity equivalent to 1.0 McFarland standard were prepared in sterile, ultra-purified water. The suspensions were boiled for 5 min and centrifuged at 12,000 x g for 5 min. The supernatants were then removed to be used as DNA templates in PCR. The primers employed to screen for the presence of the capB gene (F 5'-CATGAAGCTGAGAATGCACTTGTATT-3’ and R 5'-CTATCCCTTCTATGAATTCCGCTATT-3') were previously described [4]. The amplification reactions of the gene fragments were performed according to the method descrived by Silva Filho et al. [11], using the PCR Master Mix RED - 2.0X Taq DNA Polymerase Master Mix (Ampliqon A/S). The amplified products were analyzed by agarose (1.5%) gel electrophoresis with GelRed™ and visualized using UV light. Their sizes were estimated by comparison with a 100-bp DNA Ladder (Invitrogen - Life Technologies).
2.2. Extraction, Purification and Quantification of γ-PGA
The extraction of γ-PGA was performed based on the method by Hanby and Rydon [12], with some changes. The strains were grown in 3 mL of Tryptic Soy Broth (TSB-AcuMedia) for 24 h at 35°C and centrifuged at 12,000 x g for 5 min at 4°C. The supernatant was discarded and the cell mass was re-suspended in 3 mL of sterile distilled water. The bacterial suspension was then homogenized and autoclaved at 115°C for 45 min. To extract the γ-PGA bound to the cell surface, the suspension was mixed with 0.5 mL of concentrated HCl and 6 mL of ethanol (95%). After centrifugation at 25,000 x g for 30 min at 4°C, 5 mL of the supernatant was transferred to a new tube, and 1M NaOH solution was added until the pH reached 8.0 to 9.0. Thereafter, 15 mL of ice-cold absolute ethanol was slowly added, and the mixture was centrifuged at 25,000 x g for 30 min at 4oC. The supernatant was then discarded in order to obtain the extract containing γ-PGA (purified extract). For the quantification of γ-PGA, the purified extract was re-solubilized in 3 mL of phosphate buffer (pH 7.0) and mixed with 1 mL of 0.1M solution of cetyltrimethylammonium bromide (CTAB, Sigma-Aldrich) / 1M NaCl [13]. CTAB specifically binds to γ-PGA to form a highly dispersed micelle-like complex. The water insolubility of this complex results in increased turbidity of the suspension. The mixture was homogenized and incubated for 20 min at 30°C. Afterward, γ-PGA concentration was determined by measuring the turbidity of the mixture (OD400nm) [14]. A phosphate buffer of pH 7.0 added with 1 mL of a 0.1 M solution of cetyltrimethylammonium bromide / 1M NaCl was used as blank. The values considered as the γ-PGA amount represent the mean of the results of three independent experiments.
2.3. Influence of Sodium Chloride and Ethanol in γ-PGA Production
The effect of NaCl on γ-PGA production was studied in 40 isolates of S. epidermidis positive for the capB gene (25 producers and 15 γ-PGA non-producers in TSB). For the ethanol tests, eight capB positive isolates (six producers and two γ-PGA non-producers) were selected. Bacterial growth was performed in TSB with 1 M NaCl or 2% absolute ethanol added. As controls, the respective strains were grown in TSB without the addition of the agents. After growth, OD620nm was determined to assess whether the agents employed interfered with bacterial growth. The extraction and quantification of γ-PGA were performed as described above.
2.4. Influence of Osmolarity on γ-PGA Production
The effect of NaCl or KCl osmotic gradients on the production of γ-PGA was evaluated in three clinical isolates: two with strong γ-PGA production (EP05 and EP61) and one with moderate production (EP07). The gradients were obtained by dilution in sterile TSB to achieve the following concentrations: 0.4 M, 0.8 M, 1.0 M, 1.2 M, 1.6 and 2.0 M.
In order to correlate the amount of γ-PGA produced with the growth of the strains in the osmotic gradient, the “Corrected γ-PGA Production Index” (CPPI) was determined using the following formula: (Amount of γ-PGA (ABS400nm Value) x 100)/ Bacterial Population (ABS620nm) Value.
2.5. Fate of the Produced γ-PGA
γ-PGA, that was both cell-bound (“anchored”) and present in the culture supernatant (“free” or “released”), was investigated in three clinical isolates of S. epidermidis capB+: two strong γ-PGA producers (EP05 and EP61) and one moderate producer (EP07). Three mL of the culture grown in TSB for 24 h at 35°C was centrifuged at 25,000 x g for 25 min at 4°C and then the supernatant obtained was transferred to a new tube. The cell mass was subjected to the extraction, purification, and quantification of γ-PGA as described above, to estimate the amount of anchored γ-PGA. The procedures for the purification and quantification of γ-PGA described above were also used for the supernatant to evaluate the free γ-PGA produced.
2.6. Influence of γ-PGA on Phagocytosis by Peripheral Blood Neutrophils
Peripheral blood neutrophils from healthy subjects were isolated in the Ficoll-Paque Plus density gradient (GE HealthCare Bio-Sciences AB), re-suspended in 1 mL RPMI 1640 medium (Life Technologies-Gibco), and supplemented with 10% fetal bovine serum (RPMI 1640-SFB). The suspension was adjusted to 106 neutrophils in 290 μL of the medium [15]. The purified extract of EP07, a strong γ-PGA producer isolate, was obtained as described above and treated as described by Rhie et al. [16], with slight modifications. The extract was re-solubilized in 3 mL of purified water with the pH adjusted to 1.5 with 6 M HCl. The solution obtained was centrifuged at 10,000 x g for 20 min at 4°C, the supernatant was discarded by tube inversion, and the precipitate was solubilized in 9 mL of 1-propanol at –20°C. After brief shaking, the mixture was centrifuged at 10,000 x g for 20 min at 5°C. The precipitate was then washed twice with 10 mL of acetone and once with 10 mL of ethyl ether. After drying the ether, the purified γ-PGA was stored at –20°C. At the time of use, the dry γ-PGA was weighed and diluted in a concentration gradient (25, 50, 100, and 200 μg/mL) in Phosphate Buffer Saline (PBS), pH 7.2. 10 μL of each γ-PGA concentration was then added to 290 μL of the neutrophil suspension and incubated for 30 min at 37°C. Then, 3.0 x 106 latex beads labeled with 0.01% FITC (Sigma-Aldrich) were added to each tube, and after incubation for 30 min at 37°C, 100 μl of 0.6% trypan blue was added to identify beads that were not internalized by neutrophils. The mixture was kept in an ice bath for 10 min, centrifuged at 4000 x g for 10 min at 4°C, washed twice with ice-cold PBS containing 1% Fetal Bovine Serum (FBS), and re-suspended in 300 μl of PBS containing 1% FBS and 0.4% paraformaldehyde solution. The number of beads internalized by neutrophils was analyzed by flow cytometry (BD Accuri) by measuring the mean fluorescence emission intensity of FITC in channel 1 of the cytometer (neutrophil gate) with the use of FlowJo software V10.1.3 (Tree Star, Inc.). The number of beads adhered to the neutrophil surface was evaluated by the mean fluorescence emission intensity of the FITC after compensation using cells marked with trypan blue in channel three of the cytometer. The effect of S. epidermidis cells on phagocytosis with and without γ-PGA anchored in the cell wall (EP07 and EP60 strains, respectively) was also evaluated. 300 μl of RPMI 1640-FBS medium with 106 neutrophils was added to 1.6 mL PBS pH 7.2 containing 107 colony-forming units of each strain and 0.01% FITC. All the experiments were performed in triplicate and the results were the average of three independent experiments.
2.7. Statistical Analysis
Statistical analyses were performed using GraphPad Prism 5.03 (GraphPad Software, La Jolla, CA). The mean fluorescence intensity (ImedF) results obtained from γ-PGA influence assays on phagocytosis of latex particles were submitted to the Analysis of Variance (ANOVA), followed by Bonferroni post-test and the paired T-test for the influence of PGA on the phagocytosis of S. epidermidis isolates.
3. RESULTS
3.1. capB Gene Detection
The capB gene was detected in 40 of the 50 clinical isolates of S. epidermidis studied. The two reference strains presented results concordant with the genome described in the GenBank® database (http://www.ncbi.nlm.nih.gov/genbank/): A negative result for S. epidermidis ATCC 12228 (NCBI Reference Sequence: NC_004461.1) and a positive one for S. epidermidis ATCC 35984 (NC_002976.3).
3.2. Detection and Quantification of γ -PGA Production
Initially, the original methodology of extraction, purification, and quantification of γ-PGA production was compared with the modified methodology proposed in this study. The objective was to reduce, proportionally, the volume of the bacterial culture and the reagents used. As the difference in the results between the methodologies for five strains tested was small (the average of the difference in experiments performed in triplicate was 5.3%), the modified methodology was used to quantify the γ-PGA produced by all the clinical isolates studied. The production of γ-PGA was detected in 25 of 50 clinical isolates. The classification of γ-PGA production levels was adopted based on the dispersion of the results as follows: weak producer (OD400nm= 0.001 to 0.400), moderate producer (OD400nm= 0.401 to 1.000), and strong producer (OD400nm> 1.000). Considering this classification, 11 of the 25 γ-PGA producers presented weak production, 11 presented moderate production, and 3 presented strong production.
3.3. Correlation Between the Presence of the capB Gene and γ-PGA Production and Influence of NaCl and Ethanol on γ-PGA Production in capB Positive Strains
The production of γ-PGA was detected in 25 of the 40 strains carrying the capB gene. None of the 10 capB negative strains was identified as a γ-PGA producer. There were no significant differences in OD620nm between the TSB (control), TSB + NaCl (1M) and TSB + ethanol (2%) cultures, indicating that both the populations had a similar size. All 15 capB positive / γ-PGA negative isolates produced γ-PGA in the presence of NaCl. Twelve of these were classified as weak producers and three as moderate.
In the group of 25 strains capB positive / γ-PGA producers, NaCl did not determine changes in γ-PGA production levels in 12 strains (10 weak and 2 moderate). One isolate showed increased γ-PGA production in the presence of NaCl, moving from weak to moderate production level, while 12 strains showed a decrease in γ-PGA production levels: 3 strong producers were converted into weak, and 9 moderate became weak producers. In the group of 8 capB positive isolates tested in the presence of ethanol, there was a decrease in the production of γ-PGA in 5 of the 6 strains originally classified as producers: 2 strong producer strains were converted into moderate, and 3 moderate became weak producers. In turn, no change was observed in the sixth strain. Among the 2 non-PGA-producer isolates, 1 was induced to produce this polyanionic polymer at a weak level, while the other remained a non-producer.
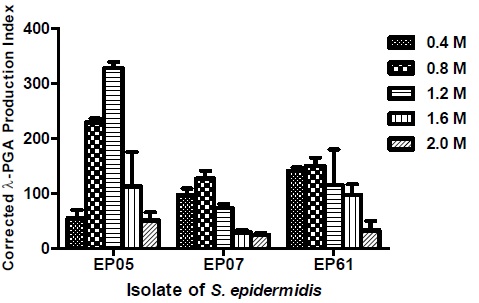
Some strains were studied regarding the influence of an osmotic gradient on the production of PGA. The correlation between the Corrected γ-PGA Production Index (CPPI) and different concentrations of NaCl and KCl are shown in Figs. (1 and 2), respectively. These tests were conducted with three isolates of S. epidermidis: two strong and one moderate γ-PGA producers.
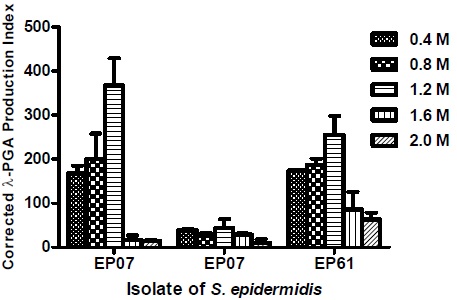
The EP05 isolate, a strong γ-PGA producer, showed the higher CPPI value at concentration of 1.2 M of NaCl and a marked decrease at 1.6 and 2.0 M. Isolate EP61, another strong producer, also showed an increase in CPPI at the concentration of 1.2 M and progressive decrease at 1.6 and 2.0 M. On the other hand, the clinical isolate EP07, classified as a weak producer, showed only small variation in the values of CPPI in the tested concentrations of the NaCl gradient (Fig. 1).
As for KCl gradient (Fig. 2), the EP05 isolate showed a significance increase in CPPI values at concentrations of 0.8 and 1.2 M, whereas the EP07 and EP61 isolates presented only a small increase at the concentration of 0.8 M, followed by a progressive decrease in higher concentrations.
3.4. Fate of the Produced γ-PGA
Table 1 shows the results of γ-PGA anchored in the cell and released in the supernatant from two strong producer isolates (EP05 and EP61) and one weak isolate (EP07). In EP05 and EP61 isolates, the amount of anchored γ-PGA was considerably higher (0.9 and 12.2 x, respectively) than the released one, while EP07 showed an opposite result.
| Isolate | γ-PGA (OD400nm) * | |
|---|---|---|
| Anchored | Released | |
| EP05 | 1.254 | 0.669 |
| EP07 | 0.883 | 1.477 |
| EP61 | 1.183 | 0.090 |
3.5. Influence of γ-PGA on Phagocytosis by Peripheral Blood Neutrophils
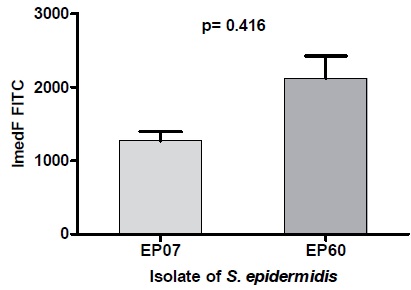
A statistically significant decrease in neutrophil internalization was observed in the γ-PGA-producer strain EP07 (p = 0.0416), in contrast with the non-producer strain EP60 (Fig. 3). For both the strains, no significant differences (p>0.05) were observed in neutrophil adhesion. Contrary to the positive protection against phagocytosis observed in the experiment with anchored γ-PGA S. epidermidis cells, none of the free PGA concentrations tested significantly altered the adhesion or internalization of latex beads.
4. DISCUSSION
In addition to biofilm, Kocianova et al. [4] pointed out the production of γ-PGA as another important mechanism of pathogenicity in S. epidermidis. This helps to explain the participation of non-biofilm producer isolates in infections, reinforcing arguments that frame this species as an important opportunistic pathogen.
In our study, the presence of capB gene was detected in 80% of the S. epidermidis clinical isolates analyzed, differing from the omnipresence of this gene detected by Kocianova et al. [4]. However, it should be noted that these authors did not find the genes of the cap operon in all the reference strains of other species of coagulase-negative staphylococci.
Extraction, purification, and dosing techniques for γ-PGA have been developed for B. anthracis and other species [12]. For S. epidermidis, however, only electronic immunomicroscopy and dot-blot techniques were reported for the detection of this polymer on the cell surface [4].
In order to develop a methodology for the determination of γ-PGA production in S. epidermidis, the PGA-extraction step used for the “anchored” γ-PGA in B. anthracis [12] was combined with a purification process based on the method described by Kanno and Takamatsu [13]. This process separates γ-PGA from the constituents of the culture medium, as well as from bacterial proteins and polysaccharides. In addition to the objective of detecting γ-PGA anchored on the surface of S. epidermidis, our study also aimed to evaluate differences between the strains in relation to their levels of γ-PGA production, as well as to estimate the production of non-anchored γ-PGA (free PGA).
Modifications in the original method included a procedure for the precipitation of γ-PGA with ice-cold absolute ethanol. After purification, the spectrophotometric amount of γ-PGA was determined using Cetyltrimethylammonium Bromide (CTAB) [13]. The principle of this method is the binding of CTAB specifically to γ-PGA to form a water-insoluble compound. This complex is highly dispersed and similar to micelles, generating turbidity in the purified extract, which is quantified by absorption in a spectrophotometer. In this method, ABS400nm values show good linearity of results and high correspondence with known concentrations of γ-PGA and can be used to estimate the amount of γ-PGA present in the purified extract [14].
The adaptations implemented in the purification step were also intended to allow a reduction in the volumes of the reagents used since the original methodology is used in processes with an industrial focus. The changes introduced were acceptable for the quantification of γ-PGA, since the difference in the results compared to the original methodology was only 5.3% on average.
In addition, all 25 γ-PGA-producer clinical isolates showed the presence of the capB gene (capB positive / γ-PGA producers), indicating that the adaptations in the methodology resulted in high specificity and did not generate false-positive results. As for the 15 capB positive / γ-PGA non-producer isolates, one hypothesis would be the lack of transcription of the cap operon in these strains under the culture conditions used. In fact, when these strains were grown in TSB with 1 M NaCl, this methodology was able to detect γ-PGA production in all the isolates that had the capB gene, indicating that the proposed method presented a total correlation with the genotype of the studied isolates.
In contrast to our expectations, in 12 of the 25 capB positive / γ-PGA producer isolates, the addition of NaCl resulted in a reduction of the production levels.
Considering the widespread use of ethanol as an antiseptic in the hospital environment, and based on the fact that low concentrations of this alcohol may affect the virulence characteristics of S. epidermidis, particularly biofilm production [17], the effect of subinhibitory ethanol concentration on γ-PGA production was investigated.
Ethanol (2%) did not determine an increase in γ-PGA production in the producer strains, but in one of the two capB positive / non γ-PGA producer strains tested, induction was observed.
The differences observed in the effect of NaCl and ethanol among the isolates tested may be due to specific mechanisms in the regulatory pathways of γ-PGA production, which may be related to particular strains and/or dependent on the conditions of the culture used.
The exposure to a NaCl gradient determined increases in CPPI values in two of the three isolates, particularly at a concentration of 1.2 M. However, CPPI values decreased at higher concentrations, contrary to what was observed by Kocianova et al. [4], who reported higher γ-PGA production at the concentration of 2 M. For the KCl gradient, one isolate showed a significance increase in CPPI values at concentrations of 0.8 and 1.2 M, while the other two presented a small increase that was restricted to the concentration of 0.8 M.
While γ-PGA is an important factor for biofilm formation in some strains of B. subtilis [6], Kocianova et al. [4] observed that this did not occur in S. epidermidis. However, studies on gene expression have indicated that the cap locus is upregulated in biofilm producers [18]. Thus, in addition to the presence of capB gene and γ-PGA production, the correlation between biofilm and γ-PGA production was evaluated using the microtiter plate technique [19] (results not shown). Among 24 biofilm-negative strains, only 6 were PGA producers in TSB, but inversely, among 26 biofilm-positive strains, only 6 were non-PGA producers.
It is also worth noting that all the 8 strains that produced biofilm with a protein matrix were positive for γ-PGA production, whereas 10 of the 15 polysaccharide biofilm-producing strains were γ-PGA-positive and 5 negative.
The fate of the γ-PGA produced has a close correlation with its function for the producing microorganism. For example, B. anthracis anchors its produced γ-PGA to peptidoglycan, forming a capsule with anti-phagocytic action [20]. Some Archaea, such as Natrialba aegyptiaca, produce and release L-PGA to the external environment, where it will exert functions such as protecting cells against drastic dehydration [21]. Different species of Bacillus release γ-PGA to sequester ions of toxic metals from the environment [22] or even to be degraded under conditions of nutrient shortage, providing glutamic acid [23]. Unlike CapD, a γ-glutamyltranspeptidase that promotes covalent anchoring of γ-PGA in B. anthracis cell wall, in B. subtilis, a γ-glutamylhydrolase (PgsS) has been identified, which allows free polymer accumulation in the culture media [24].
CONCLUSION
In S. epidermidis, only anchored γ-PGA were previously reported [4]. However, in addition to anchored PGA, our study also detected PGA in the supernatant of the three isolates that were tested for free PGA. In one of them, the amount of free γ-PGA was 67% greater than that of the anchored one, while in the other two, the amount of anchored γ-PGA was higher. This result suggests that the amount of anchored or free γ-PGA produced in S. epidermidis may be related to particular physiological characteristics of the producing strain. Regarding the mechanism of release of PGA to the extracellular medium, it may involve some enzymatic action similar to that occurring in B. subtilis, but acting partially, since anchored γ-PGA was also found in the isolates studied. Another hypothesis is that at the stage of growth of the microorganism, γ-PGA production exceeded its binding capacity to the peptidoglycan and was then released into the culture medium. S. epidermidis also produces CapD, which is associated with the anchoring of PGA on the bacterial surface. However, it differs considerably from B. anthracisCapD, possibly reflecting differences in γ-PGA acceptors in the peptidoglycan [24]. Indeed, the S. epidermidis peptidoglycan harbors a lysine with a pentaglycine peptide to which the γ-PGA can be anchored, in place of the meso-diaminopimelate present in the B. anthracis peptidoglycan [24]. Although different concentrations of free γ-PGA did not determine a reduction in the internalization of latex beads by neutrophils, cells with anchored γ-PGA showed significant protection against phagocytosis compared to the non-producing strain tested. It is important to note that this γ-PGA negative strain was characterized as a strong producer of polysaccharide biofilm (results not shown). This result suggests that γ-PGA anchored at the cell surface may confer greater anti-phagocytic protection than PIA, the major component of the polysaccharide biofilm matrix, which can also be found on the cell surface of the bacterium in the planktonic form [25].
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study was approved by the Human Research Ethics Committee (HFSE), Brazil with approval number 000.417.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Written informed consent was obtained from all the participants.
AVAILABILITY OF DATA AND MATERIALS
The authors confirm that the data supporting the findings of this research are available within the article.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We gratefully acknowledged the Federal University of the State of Rio de Janeiro (UNIRIO), Rio de Janeiro, Brazil.


