All published articles of this journal are available on ScienceDirect.
Distribution of Class 1 and 2 Integrons Among Salmonella Enterica Serovars Isolated from Iranian Patients
Abstract
Background:
Salmonella enterica has become increasingly resistant to antimicrobial agents, partly as a result of genes carried by integrons.
Objective:
The aim of the present study was to investigate the prevalence of class 1 and 2 integrons and resistance to antimicrobial agents in clinical isolates of S. enterica.
Methods:
This study included all Salmonella isolates, recovered from patients with salmonellosis, admitted to Medical Children Hospital, Tehran, Iran during 2015-2016. Bacterial isolates were identified using standard biochemical and agglutination tests. Antimicrobial susceptibility testing was performed according to the Clinical and Laboratory Standards Institute guidelines. The presence of class 1 and 2 integrons was investigated by Polymerase Chain Reaction (PCR) assay, using specific primers.
Results:
A total of 138 Salmonella strains were isolated and included in this study. Integrons were detected in 45 (32%) isolates. Class 1 and 2 integrons were detected in 24 (17.3%) and 21 (15.2%) isolates, respectively. All integron-positive isolates showed multidrug-resistant phenotypes. Resistance to more than three antimicrobial agents was observed in integron-positive isolates.
Conclusions:
Our findings showed that integrons were widely distributed among S. enterica isolates in Tehran. Class 1 integrons are more prevalent than class 2 integrons in Salmonella isolates, and there is an association with MDR patterns. Therefore, these integrons are more likely to be involved in the distribution of resistant phenotypes in Salmonella strains.
1. INTRODUCTION
Gastroenteritis and diarrheal diseases are among the most important health problems worldwide, especially in developing countries [1]. Salmonella remains one of the most common causes of acute gastroenteritis and food-borne infections around the world [2]. Different species of this bacterium are frequently encountered in different countries, including Iran [3]. Antibiotic resistance is common among these serovars, and a high prevalence of multidrug-resistance (MDR) isolates has also been reported [4].
There are many mechanisms of antibiotic resistance through which bacteria exhibit resistance to antibiotics. Acquisition of genetic elements is one of the major mechanisms, which frequently occurs in both Gram-positive and Gram-
| Primers | (5′→3′) Sequence | Amplicon size (bp) | References |
|---|---|---|---|
| Int1 | F:5′-GGGTCAAGGATCTGGATTTCG-3′ | 483 | [11] |
| R:5'-ACATGCGTGTAAATCATCGTCG -3' | |||
| Int2 | F:5'-GCAAATGAAGTGCAACGC-3' | ||
| R:5' -ACACGCTTGCTAACGATG -3' | 466 | [11] |
| Antibiotics (µg)* | Group A | Group B | Group C | Group D | Total |
|---|---|---|---|---|---|
| TE (30) | 1 | 9 | 24 | 21 | 55 |
| S (10) | 0 | 9 | 16 | 19 | 44 |
| SXT (1.25+23.75) | 0 | 4 | 12 | 2 | 18 |
| AMP (10) | 0 | 1 | 4 | 4 | 9 |
| GM (10) | 0 | 6 | 8 | 17 | 31 |
negative bacteria. Integrons are DNA elements with major involvement in the spread of antibiotic-resistant genes through horizontal or vertical gene transfer mechanisms [5]. Although at least six classes of integrons have been reported, there is limited data about these integrons classes, and only classes 1, 2, and 3 are well-studied [6].
Integrons are very common in S. enterica serovars and seem to be significantly involved in the advent of antibiotic-resistant phenotypes in the strains [7]. The presence of integrons in the strains increases resistance to fluoroquinolones, which are very effective in the treatment of salmonellosis [8]. In addition to fluoroquinolone resistance, resistance to cephalosporins has been also associated with integron cassettes. It is clear that veb-1 gene, encoding resistance to cephalosporins, is a part of integron cassettes. Fluoroquinolones and cephalosporins are two main classes of antibiotics, frequently used to treat Salmonella infections. The prevalence of resistant genes in association with integron cassettes can be a new challenge in the treatment of salmonellosis and its rate is increasing; therefore, it is necessary to determine different classes of integrons in the strains [9].
The aim of the current study was to investigate the prevalence of integrons and resistance to antimicrobial agents in clinical isolates of S. enterica, isolated from patients admitted to the Medical Children Hospital of Tehran, Iran.
2. MATERIALS AND METHODS
2.1. Bacterial Isolates
This study included all recovered Salmonella isolates from patients with Salmonella infections, who were admitted to the Children Medical Center of Tehran, Iran during 2015-2016. The isolates were identified by standard biochemical and agglutination tests using commercial antisera (Staten Serum Institute, Copenhagen, Denmark), according to the manufacturer’s instructions.
Antimicrobial susceptibility testing was performed according to the Clinical and Laboratory Standards Institute (CLSI) guidelines and Kirby-Bauer disc diffusion method. The tested antibiotics included cotrimoxazole (1.25/23.75 μg), gentamicin (10 μg), ampicillin (10 μg), streptomycin (10 μg), and tetracycline (30 μg).
2.2. DNA Extraction
DNA extraction was carried out using a previously described method, with some modifications [10]. Briefly, overnight bacterial cultures were separately suspended in 250 μL of lysis buffer (10 mM tris-HCl, 0.1 M NaCl, 1 mM EDTA, and 5% [v/v] Triton X100; pH, 8.0) and boiled at 100°C for three minutes. The suspensions were centrifuged at 10 000 rpm for 10 minutes, and the supernatants were transferred to new sterile microtubes. Then, 250 μL of ethanol 95% was added to each microtube and kept at −20°C for 30 minutes. The solutions were centrifuged at 14 000 rpm for 20 minutes, and then, the sediments were dissolved in 100 μL of sterile distilled water and stored at −20°C. DNA concentration was measured using Thermo Scientific NanoDrop spectrophotometers.
2.3. Polymerase Chain Reaction (PCR) Assay
The PCR assay was carried out for the reaction mixture in a final volume of 25 uL, containing 12.5 μL of 2X PCR Master Mix (CinnaGen, Iran), 10 pmol of forward and reverse primers (Bioneer, Korea) (Table 1), and 50 ng DNA. The PCR conditions were as follows: initial denaturation at 94°C for five minutes; 30 cycles of 94°C for 50 seconds; annealing at 60°C and 72°C for one minute; and a final extension at 72°C for 10 minutes using a thermocycler (Eppendorf Thermal Cycler, Germany). The PCR products were electrophoresed on 1% (W/V) Agarose gel (Sigma-Aldrich, USA) and visualized under UV light after staining with a DNA Safe Stain (CinnaGen, Iran).
3. RESULTS
Among 138 Salmonella isolates, 1, 18, 41, and 78 isolates belonged to serogroups A, B, C, and D, respectively. The results of the antimicrobial susceptibility test, based on the serogroup categories, are presented in Table (2). As shown in this Table, 55 isolates were resistant to tetracycline, while only nine isolates had resistant phenotypes to ampicillin.
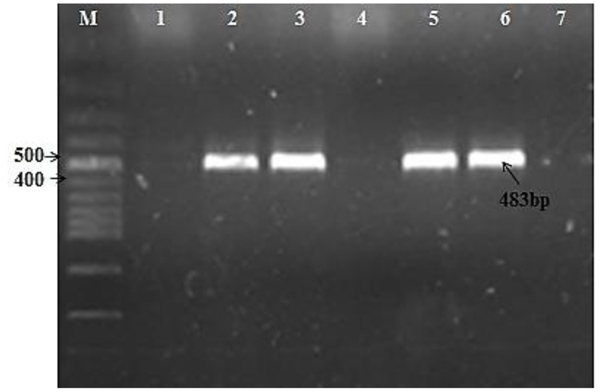
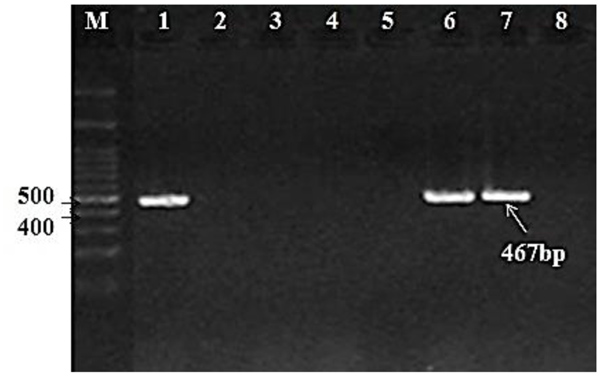
A total of 138 isolates were screened for int1 and int2 genes. Integron classes 1 and 2 were only detected in 45 isolates (class 1: 24 isolates (17.3%); class 2: 21 isolates (15.2%)). Figs. (1 and 2) demonstrate the amplicons of class 1 and 2 integrons, respectively.
4. DISCUSSION
Different levels of antimicrobial resistance to several antibiotics are common among bacteria worldwide [12]. Integrons are genetic elements associated with resistance to antimicrobial agents. They are categorized into more than six classes, based on the intI gene sequence [13]. Transferability of integrons, especially class 1 integrons, has been found to be common among bacteria through the horizontal gene transfer mechanism [14]; our findings are consistent with the literature.
In the present study, the prevalence of antibiotic resistance among Salmonella isolates was determined. The results showed that most isolates were resistant and susceptible to tetracycline and ampicillin, respectively. In accordance with our findings, Jackson et al. showed that S. typhimurium isolates are resistant to tetracycline, ampicillin, and cotrimoxazole [15]. In a previous study, we revealed that Salmonella isolates had high levels of resistance to doxycycline, tetracycline, and strepto mycin [16]. Our findings, in line with previous studies, indicate that resistance to tetracycline is common among isolates, and this agent seems to be ineffective in the treatment of Salmonella infections.
In this study, resistance to antimicrobial agents varied among Salmonella serogroups; this finding is predictable due to the presence of different strains in these groups. These serogroups were formerly classified as Salmonella species. Also, different patterns of antibiotic resistance have been reported among Salmonella species by other researchers [17, 18].
In the present study, we characterized two classes of integrons, and 45 out of 138 isolates harbored class 1 (17.3%) and class 2 (15.2%) integrons. In a previous study, Asgharpour et al. showed that 36% of S. infantis isolates were positive for class 1 integrons, and all these isolates were also resistant to nalidixic acid, tetracycline, and streptomycin [19]. In comparison with our study, the higher prevalence of class 1 integrons in the mentioned study shows that application of antibiotics in the poultry industry may increase antibiotic resistance [20]. In another study, 45% of S. enterica serotypes, including Mbandaka, Brandenburg, Agona, Alachua, and Panama serotypes, harbored class 1 integron genes [21].
Up to 17.39% of isolates harboring class 1 integrons were resistant to ampicillin (2.89%), gentamicin and trimethoprim / sulfamethoxazole (5.79%), tetracycline (10.14%), and streptomycin (7.97%), whereas resistance to these antibiotics in integron-negative isolates was 14.49%, 7.97%, 7.24%, 26.08%, and 22.46%, respectively. The low level of ampicillin resistance among integron-negative isolates suggests that resistance to this antibiotic is not associated with integrons and is commonly associated with TEM β-lactamases, as described in the literature [22].
The prevalence of class 2 integrons was found to be 15.21% among the studied isolates, indicating the low prevalence of this class of integrons. In studies by other researchers, the prevalence of class 2 integrons was also lower than our study [23, 24]. In this regard, in a study by Jin et al., class 2 integrons were not detected even among 834 Salmonella isolates [25].
CONCLUSION
According to our findings, the prevalence of antibiotic resistance is high among Salmonella isolates, and screening for integrons, as likely indicators of antibiotic resistance, may be useful in detecting Salmonella organisms with antibiotic-resistance phenotypes. Although our study showed that integrons are associated with antibiotic-resistance phenotypes, further studies are needed to determine the exact relationship between classes of integrons and resistance to different antibiotics.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study was approved in Ethical Committee No.33 in Baqiyatallah University of Medical Sciences, Tehran.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Informed consent was obtaind from all the participants.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENT
We would like to thank the “Clinical Research Development Center of Baqiyatallah hospital” for their kind cooperation. This study was financially supported in part by “Clinical Research Development Center of Baqiyatallah hospital”.


