All published articles of this journal are available on ScienceDirect.
Analysis of Routine and Integrative Data from Clostridioides difficile Infection Diagnosis and the Consequent Observations
Abstract
Background:
Clostridioides difficile Infection (CDI) is an acute disease that needs a fast proper treatment. Unfortunately, the diagnosis, and above all the understanding of the results, remain arduous.
Objective:
This study analyzed routine and integrative results of all fecal samples from patients over time. Our aim was to understand the dynamics of CDI infection and the meaning of “difficult to interpret” results, to make physicians better understand the various tools they can use.
Methods:
We evaluated routine results obtained from 815 diarrheal stools with Enzyme Immunoassay (EIA) that detects C. difficile Glutamate Dehydrogenase (GDH) antigen and toxin B. We also reanalyzed a part of samples using integrative tests: a Real-time polymerase chain reaction (RT-PCR) for C. difficile toxin B gene (tcdB) and the automated immunoassay VIDAS C. difficile system for GDH and toxins A/B.
Results:
EIA GDH positivity increased through multiple testing over time, with a P value <0.001, depicting a sort of bacterial growth curve. Eighty-five percent of GDH positive/toxin B negative, i.e., discrepant, samples PCR were tcdB positive, 61.5% of discrepant tcdB positive samples were VIDAS toxins A/B positive, and 44.4% of GDH EIA negative stools were VIDAS GDH positive.
Conclusion:
The results confirmed the low sensitivity of the EIA system for C. difficile GDH and toxins, questioned the use of the latter for concluding any CDI diagnostic algorithm, and led us to indicate the algorithm beginning with tcdB molecular research, and continuing in positive cases with VIDAS CD GDH method, as the most effective for CDI.
1. INTRODUCTION
Clostridioides difficile is a Gram-positive, spore forming, strictly anaerobic bacterium. It is the leading cause of the principal antibiotic-associated diarrhea. In humans, C. difficile Infection (CDI) is mediated by cytotoxic enzymes, i.e., toxin A (TcdA), toxin B (TcdB), and, when present, by the binary toxin (CDT), which can potentiate intestinal damage [1].
Given the feature of acute infection, CDI needs a rapid reliable diagnosis, which avoids empirical therapies and consequent further dismicrobism, responsible for most cases [2]. Diagnosis of CDI is performed through the detection of toxins in stools. The Cell Cytotoxicity Neutralization Assay (CCNA) primarily reveals the presence of TcdB through cytopathic effect, whose specificity is assured precisely by neutralization. Given its high sensitivity and specificity, CCNA is recommended as a reference test. Accomplishment of the CCNA procedure is long and burdensome, thus, for routine diagnosis, toxins are revealed by enzyme immunoassays (EIAs), which, though low in sensitivity, are fast and manageable [3].
Little toxin, sufficient to cause intestinal damage, such as in vitro cytopathic effect, yet not sufficient enough to be always detected by the enzyme immunoassay, primarily complicate CDI diagnosis [3]. For this reason, the efficacy of the EIA system has been increased entrusting it with the detection of C. difficile glutamate dehydrogenase (GDH), present during infection in large quantities [3] but endowed with a lower specificity, since GDH is common to toxigenic and non-toxigenic strains [4, 5]. Secondly, in the human bowel, C. difficile can be present in spore and vegetative forms, both sharing tcdB, the gene codifying for TcdB and nowadays used as a diagnostic target [6]. Thanks to the most recent works it is now possible to overcome the old consequent debate, between those authors who feared unrecognized infection and those who feared over-diagnosis [7]. In 2016 and 2018, respectively, the European and American Society of Infectious Diseases updated diagnostic guidance documents for CDI, giving important evidence about laboratory diagnosis [8, 9]. The most interesting evidences are the inappropriateness of any test used as a stand-alone, the evaluation of new available tests and their possible employment in integrating the diagnostic systems and creating a necessary algorithm.
In our opinion, however, clinicians continue to have difficulty interpreting those diagnostic results which are not fully consistent with each other, or are incomplete. Therefore, the aim of this work was to discover what is behind the so-called ‘difficult to interpret’ results and what can be extrapolated [4]. We report results obtained with C. difficile GDH and toxin B EIA analyses on stools collected during 10 months from patients suspected of suffering from CDI, including repeat samples. When possible, we employed integrative diagnostic systems to reanalyze samples with routine discrepant results.
2. MATERIALS AND METHODS
2.1. Patients and Setting
This study was a cohort study of consecutive patients admitted to the San Martino Hospital in Genoa, Italy, and of subjects in the community from the same regional area, affected by diarrhea, from June 2013 to March 2014. Patient’s stools were analyzed at the microbiology laboratory for TcdB detection. All clinical specimens were collected and stored for the study which was approved by the Institutional Ethics Committee of San Martino University Hospital (n. reg. CEA 13/11-Progetto istMicro1/2011). We considered the total number of patients and analyses, including those repeated for single subjects and changes of results concerning single patients over the time.
2.2. Microbiological Methods
Diarrheic stools were routinely analyzed for the presence of C. difficile GDH and C. difficile toxin B with a Rapid Membrane EIA, C. difficile Quik Chek Complete (Techlab Inc.) within 1 hour from arrival at the laboratory and according to the manufacturer's instructions. Depending on the availability of fecal material, GDH positive, toxin B negative samples, the previously called “discrepant samples” [4], were further analyzed by nucleic acid amplification test for C. difficile toxin B gene (tcdB) and the more sensitive automated immunoassay VIDAS C. difficile Tox A/B (BioMerieux) [10]. For the detection of tcdB, bacterial DNA was extracted from stools with the automated device QIAcube (QIAGEN). Using the extracted DNA, we performed Real-Time Polymerase Chain Reaction (RT-PCR) using RealCycler CDIF (Progenie Molecular), suitable to detect the tcdB gene directly from stools, following the manufacturer’s instructions. Fifty-three EIA GDH negative/toxin B negative samples were reanalyzed with the automated immunoassay VIDAS C. difficile GDH (BioMerieux).
2.3. Statistical Methods
The occurrence of the three possible outcomes from the assay matching GDH and toxin B (positive/positive, negative /negative, positive/negative) with respect to tests repeated over time was tested by means of the multinomial logistic model [11]. This model performed the ratio between the percentage of positive results (GDH positive/toxin B positive and GDH positive/toxin B negative) for the additional samples over time with respect to the first sample, thus estimating Relative Risks (RRs). These RRs were then compared with the analogous relative risks estimated for the double negative GDH negative/toxin B negative outcome, thus obtaining the Relative Risks Ratio (RRR). The standard errors estimated from the multinomial model were adjusted by the clustering effect due to the repeated observations on the same subject. The Kappa test was performed to evaluate the agreement between the different assays. To avoid the influence of repeated measures, the Kappa test was applied to the sum of positivity of each assay [12].
3. RESULTS
3.1. Patients, Stools and Analyses with the Routine EIA System
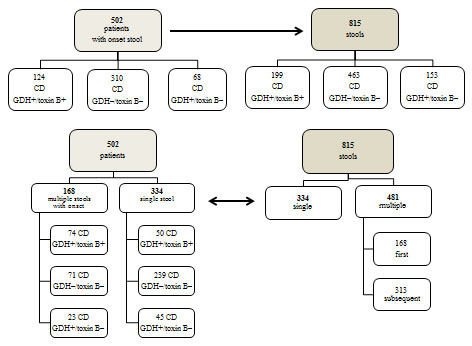
Over the study period, we routinely analyzed 815 diarrheal stools from 502 subjects with EIA systems. Fig. (1) shows total patients, disease onset according to EIA C. difficile GDH/ toxin B results, total stools and the relevant EIA results, grouped according to single or multiple stools. We did not show epidemiological data concerning patients as they were not relevant to our analysis. Table 1 shows that, in addition to the 334 (66.5%) patients with a single analysis, 168 (33.5%) (data not shown) patients with multiple analyses, i.e., 72 (14.4%) patients with multiple analyses always giving the same result, positive, negative or discrepant, and 96 (19.1%) giving different results over time. Sixty of them (11.9%) had positive onset, 19 (3.8%) had negative onset and 17 (3.4%) discrepant (data not shown).
Table 2 reports the results of the EIA system for GDH and toxin B, stratified by single or multiple stools. Observing the row of percentages, it is clear that the frequency of positivity increases through multiple testing over time. The multinomial logistic model was applied to test this hypothesis. This model shows that the probability of finding a positive result significantly increases by performing up to two or three replicates, at least for +/– (RRR = 2.36; 95%CI = 1.54-3.60 and RRR=3.74; 95%CI = 2.07-6.76 respectively; P <0.001) while for +/+ the result is not significant (P = 0.206) (Table 3).
| Specimen/s | Results |
Patients Number |
% |
|---|---|---|---|
| single | +/+ | 50 | 9.9 |
| single | -/- | 239 | 47.6 |
| single | +/- | 45 | 9.0 |
| total single | 334 | 66.5 | |
| multiple always | +/+ | 14 | 2.8 |
| multiple always | -/- | 52 | 10.4 |
| multiple always | +/- | 6 | 1.2 |
| total multiple always | 72 | 14.4 | |
| multiple variable | 96 | 19.1 | |
| Total | 502 | 100 |
|
Results GDH |
In Stools toxin B |
Multiple Test | – | Total N. |
|---|---|---|---|---|
| Yes No. Yes % | No No. No % | |||
| + | + | 149 74.87 | 50 25.13 | 199 |
| - | - | 224 48.38 | 239 51.6 | 463 |
| + | - | 108 70.59 | 45 29.41 | 153 |
| Total | stools | 481 59.02 | 334 40.98 | 815 |
3.2. Fecal Samples Analyzed with Integrative Tests
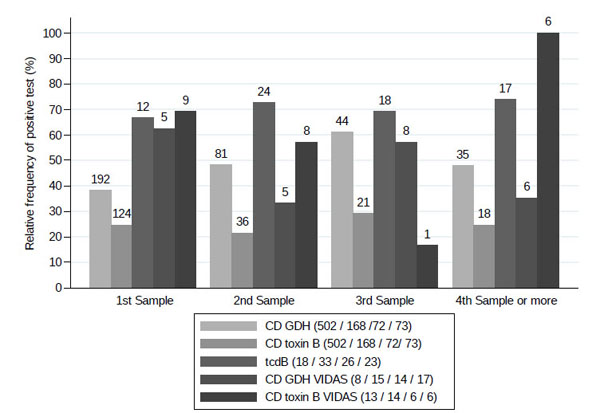
Table 4 shows, for descriptive purposes only, the relative frequencies of the EIA GDH positive/toxin B negative and EIA GDH negative/toxin B negative stools, which were positive using the integrative systems targeting tdB or the VIDAS C. difficile toxin A/B and the VIDAS C. difficile GDH tests, and the provenance according to patients and single or multiple samples. Fig. (2) describes the total positive results for CDI according to the stool sample during the time, obtained with routine analyses and with the integrative tests utilized and shown in Table 4.
| – | GDH-/toxin B – |
GDH+/toxin B – [P<0.001] |
GDH+/toxin B+ [P=0.206] |
||||
|---|---|---|---|---|---|---|---|
| Replicates | Frequency % | Frequency % | RRR | 95% CI | Frequency % | RRR | 95% CI |
| 1 | 61.6 | 13.6 | 1 | ref. | 24.7 | 1 | ref. |
| 2 | 51.8 | 26.8 | 2.36 | 1.54 - 3.60 | 21.4 | 1.03 | 0.69 - 1.54 |
| 3 | 38.9 | 31.9 | 3.74 | 2.07 - 6.76 | 29.2 | 1.88 | 1.04 - 3.38 |
| ≥4 | 52.1 | 23.3 | 2.04 | 0.95 - 4.37 | 24.1 | 1.18 | 0.69 - 2.04 |
| – |
Patients Number |
Stools Numbers |
Patient Trend |
EIA CD GDH |
EIA CD Toxin B |
Integrative | Tests |
|---|---|---|---|---|---|---|---|
| – | – | – | – | – | – | tcdB PCR + | tcdB PCR - |
| – | 44 | 60 | variable | positive | negative | 50 | 10 |
| – | 5 | 8 | always +/- | positive | negative | 8 (85.3)* | 0 (14.7)* |
| – | 23 | 32 | variable | negative | negative | 13 (40.6) | 19 (59.4) |
| Total | – | 100 | – | – | – | 71 (71) | 29 (29) |
| – | – | – | – | – | – | VIDAS toxin B + | VIDAS toxin B - |
| – | 27 | 30 | variable | positive | negative | 17 | 13 |
| – | 2 | 3 | always +/- | positive | negative | 2 | 1 |
| – | 6 | 6 | single +/- | positive | negative | 5 | 1 |
| Total | 35 | 39 | – | – | – | 24 (61.5) | 15 (38.5) |
| – | – | – | – | – | – | VIDAS CD GDH + | VIDAS CD GDH - |
| – | 36 | 54 | variable | negative | negative | 24 (44.4) | 30 (55.6) |
3.2.1. Samples analyzed with PCR for tcdB
We selected 100 fecal samples, 68 EIA discrepant and 32 double negative, where to search for tcdB with the RealCycler CDIF assay. Out of the 68 EIA discrepant samples, 85.3% were tcdB positive, while tcdB positivity decreased to 71% among all one-hundred selected samples. In fact, the tcdB positivity percentage among the 32 negative EIA samples was 40.6% and attributed greater sensitivity to PCR for toxin gene compared to the EIA system for GDH. All eight discrepant stools of patients who had always given the same discrepant result were tcdB positive. This is noteworthy because, assuming that the individual patient had the same intestinal C. difficile strain in a short time, it leads us to consider the role of genetic groups, hyper virulent or not, in the genesis of discrepant results [3]. Thirty-two stools were EIA negative from 23 patients giving variable results over time, half of which also had discrepant EIA results and belonged to the aforementioned group of 44 subjects. Three EIA negative subjects had both positive and negative tcdB. Compared to GDH EIA tests, tcdB assay shows an agreement, i.e., the relative frequency when the two methods are jointly positive or negative, of 69.4%. Applying the Kappa test to check the repeatability between the two methods, the hypothesis of a random agreement, estimated to be 42.1%, was rejected with a P <0.001.
3.2.2. Samples analyzed with the automated immunoassay VIDAS C. difficile Tox A/B
We selected 39 toxin B EIA negative samples where tcdB and GDH EIA positivity ensured, respectively, the presence of toxigenic strains and germinating conditions, to search for toxin B with a test more sensitive than EIA, the VIDAS method [10]. Positive VIDAS toxin B stools were globally 24 out of 39. Out of the 24 VIDAS toxins positive stools, only 9 were onset samples, while 15 from 13 patients were repeats (data not shown). The observed agreement with GDH is 60.0%, quite different from the random one of 46.2%, thus the random agreement hypothesis was rejected by the Kappa test (P <0.001).
3.2.3. Samples analyzed with the automated immunoassay VIDAS C. difficile GDH
We analyzed 54 EIA GDH negative/toxin B negative samples with the VIDAS C. difficile GDH test, in order to confirm the greater sensitivity of this system on the GDH target with respect to EIA. We selected these samples as likely to be positive, in that coming from 36 patients with variable results. We obtained 24 VIDAS C. difficile GDH positive results and 30 negative. Out of 24 VIDAS GDH positive stools, only 5 were onset samples, while 19 from 17 patients were repeats, and 11 preempted those obtained with the EIA system (data not shown). Since the observed and the expected agreement with EIA GDH were only slightly different (50.0 and 47.2 respectively) the Kappa test suggested that the concordance between the two assays was not significant (P = 0.138). Out of 24 EIA GDH negative stools retested with both VIDAS for GDH and PCR for tcdB, 15 were GDH positive and 9 were negative, while 12 were tcdB positive and 12 were negative (data not shown).
4. DISCUSSION
In the last years, a better knowledge of C. difficile infection clarified some misleading points, which previously had increased the intrinsic limits of diagnostic tools. The main achievement is represented by the fact that over diagnosis is no longer contested with evidence of cases of underdiagnoses, and vice versa [13-16]. Actually, the diagnostic guidance recommendation for using a diagnostic algorithm just arises from awareness that various pathogenic frameworks, due to the same bacterium, are possibly present behind various unsatisfying diagnoses or fragments of results, and cannot be distinguished by a single test alone [8, 9]. Nevertheless, CDI diagnosis remains complex, more than that of most bacterial infections [3]. In our opinion, some viewpoints do remain unconvincing, such as the idea of possible toxins’ degradation, and prevent clinicians from having a vision clear enough to extrapolate what is best for the individual patient from diagnostic results.
We evaluated the first and the additional analyses of all subjects suspected of being affected by CDI, to know the dynamic of the infection, to select samples for analyses with integrative tests and to frame the relevant results in the right context.
One third of our cohort was composed of subjects who gave subsequent samples over time. More than one third of the stools were repeats. This aspect primarily shows a lack of confidence in the tests for CDI detection, which led to the disregard, in our institution, of the recommendation not to repeat tests [8].
Data from routine analyses show that, by performing up to three or four new EIA analyses, the probability of detecting GDH positivity significantly increases, while that of finding toxin B positivity increases less clearly. In the past, the possible benefit deriving from repeated testing using CCNA, PCR for tcdB and the EIA system for toxins was excluded [17-19]. It is possible that the most sensitive analyses, as well as the least sensitive one, were equally indifferent to possible changes in bacterial amount during infection. Different to the EIA GDH performance observed here, CCNA and PCR for tcdB may not require peaks in quantity to detect their targets, and EIA toxin B sensitivity may never reach the detection threshold.
Previous studies showed that C. difficile bacterial load in infection, expressed as CFU/g or tcdB DNA copies/mL, positively correlates with stools positive for tcdB and even for toxins’ products detected with the EIA system [20, 21]. Other works reported the same correlation indirectly, through the degree of severity of the cases [16, 22, 23]. Therefore, results varying over time within single subjects could indirectly indicate variable fecal loads of C. difficile, that, probably as in other infections, follows a sort of growth curve. We did not consider the gain in positivity of the repeats obtained with integrative tests, even if apparently evident, in that we performed a precise selection of stools eligible for further analyses, distorting our epidemiology.
We absolutely do not propose additional samples to achieve a diagnostic certainty, in that this would contradict the need for fast results. Moreover, EIA GDH multiple analyses would not be exhaustive, yielding a gain in positivity resulting from the intermediate sensitivity, while recovery of positivity for EIA toxins would not be significant. We analyzed results from repetitions just to confirm the bacterial dynamics we had hypothesized, which in turn is partly responsible for the complexity of CDI diagnosis, and which, in our opinion, must be taken into account.
Out of the thirty-nine EIA discrepant/tcdB positive samples, which indicated the presence of vegetative and toxigenic strains, twenty-four were positive for C. difficile toxin B with the VIDAS ToxinA/B test. Given the low quantity of many feces, these results are partial, when compared to the total number. However, we can assert that, in these samples, CD toxin was present, detectable with a test more sensitive than the routine one, and not degraded, as instead emerged from other works [21, 24-26]. In our opinion, toxin quantity under the detection threshold of the EIA, and more rarely under the threshold of VIDAS system, whose higher sensitivity compared to EIA is also confirmed on GDH target, is the most plausible explanation for GDH positive, toxins negative, tcdB positive results. The toxins’ instability had already been considered responsible for the greater sensitivity of Toxigenic Cultures (TC) compared to CCNA [7]. It is now ascertained that the greater sensitivity of TC, compared to detection toxin products’ detection, is illusory and due to the low specificity of the method that, like PCR for tcdB, also detects spore forms [16].
Our results, as well as those of authors here cited, lead us to disagree with the specific proposal to follow with EIA systems for CD toxins the positivity of CDI algorithms started with EIA GDH or with PCR for tcdB [8, 9, 27]. In fact, EIA systems for toxins detection are not sensitive enough to conclude the algorithm in case of its own negativity. Furthermore, given the current availability of EIA tests complete for both GDH and toxins A/B, the choice of GDH alone as the first step of the algorithm is not clear, and testing for toxins in sequence would be a step backwards. Companies improved EIA systems for CDI with GDH to recover samples that would be negative when evaluated for toxins alone, and to address the search for tcdB only where it is worthwhile. That is where GDH indicates CD germination but the toxins’ products are not detected. However, the present data on the greater sensitivity of VIDAS and on its ability to anticipate GDH detection compared to EIA, although few, show the possibility that EIA does not detect a GDH positivity that would otherwise be detectable. Since the VIDAS system is not manageable as EIA, the same VIDAS for GDH is not suitable as a starting point for a routine algorithm. Moreover, here, we confirmed the higher sensitivity of PCR for tcdB compared to both EIA and VIDAS for GDH, due in part to the molecular method itself, which is indeed amplification, and partly to the different target [3].
VIDAS C. difficile GDH. On top of the columns are the absolute numbers of positive results. The frame containing the legend of tests utilized, shows the total number of 1st samples (onset assay) and those of the 2nd, 3rd, and 4th samples (repeat assays), in brackets.
The bacterial dynamic we observed by following a large number of subjects over time, and the variables it can contribute to create, confirm the great complexity of the disease. This imposes, in turn, an unequivocal diagnostics, only achievable through an algorithm. We think that, in the current state of technology, the beginning of CDI algorithm should be entrusted to the search for tcdB, nowadays manageable. Positive cases, in which it is worth continuing the investigation, in small quantities, should be followed by VIDAS GDH, to exclude spore forms. In fact, in our opinion, it is preferable to begin any diagnostic process with the more sensitive method, especially when it is possible to correct its poor specificity, rather than underestimating a disease.
CONCLUSION
The above mentioned algorithm is quite feasible with the current tools, although these are not available in all diagnostic laboratories. Anyway, even incomplete results, and those somewhat discordant, can be correctly interpreted and utilized by physicians. Faecal CD GDH alone, not supported by toxins or tcdB positivity, not found or not searched, represents the risk of germination for any new toxigenic strains. The tcdB positivity alone, not supported by the presence of GHD and toxins’ products, not found or not searched, indicates the faecal presence of C. difficile spore forms that can germinate when the environment is suitable, condition underlying CDI recurrence [13, 28]. The algorithm itself must be clearly interpreted as a diagnostic path that distinguishes the low amount of toxin from the non-toxigenic C. difficile, and germination or not of toxigenic strains.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by the Institutional Ethics Committee of San Martino University Hospital (n. reg. CEA 13/11-Progetto istMicro1/2011), Italy.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Informed consent was obtained from each participant.
STANDARD OF REPORTING
STROBE guidelines and methodology were followed.
AVAILABILITY OF DATA AND MATERIALS
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENT
We thank Dr. Jennifer McDermott for English editing


