All published articles of this journal are available on ScienceDirect.
Antibiotic Resistance Patterns and Virulence Determinants of Different SCCmec and Pulsotypes of Staphylococcus Aureus Isolated from a Major Hospital in Ilam, Iran
Abstract
Aims & Objectives:
The aim of this study is to evaluate genetic relatedness, antibiotic resistance pattern, and virulence characteristics of different types of S. aureus isolated from air, surfaces, staff, and patients in a Public hospital in Ilam.
Methods & Materials:
A total of 88 of 140 staphylococci identified as S. aureus by conventional and molecular methods were used in this study. Isolate samples were obtained from surfaces, staff, patients, and hospital indoor air. The sampling from staff and surfaces was done through using swab and air by standard pump. Antimicrobial susceptibility testing and presence different resistant and virulence determinants was assessed. Isolates were then typed by pulsed-field gel electrophoresis (PFGE) and SCCmec typing methods.
Results:
Out of 88isolates, 36 of them (40.9%) were MRSA. Among MRSA isolates, the range of resistance to antibiotic was 0% in vancomycin to 83.3% in gentamycin. The most prevalent resistant genes among gentamicin resistant S. aureus were acc (6')/aph (2”)Ia and aph(3”)IIIa. The most common erythromycin resistant gene was ermC. Surprisingly, SCCmec types I (30.5%), II (25%)were highly distributed. PFGE analysis showed 33 different pulsotypes.
Conclusion:
This study confirms that different isolates of MSSA and MRSA circulate in Ilam which differ in antimicrobial susceptibility, content of resistance, and virulence determinants.
1. INTRODUCTION
Nosocomial infections are among important factors inpatient's mortality in Iran and other countries. Hospital environments have been widely identified as the main source of multidrug-resistant bacteria, most importantly MRSA strains. MRSA identification is vital for controlling and eliminating the outbreaks [1-3].
S. aureus is a Gram-positive, catalase-positive, non-spore forming facultative anaerobic bacteria. This bacterium can colonize in the nostril's anterior section, vagina, armpit, perineum, skin (especially damaged areas), newborns'umbilicus, and oropharynx [1]. These opportunistic pathogens show various virulence factors such as their adhesion, enzymes, and different toxins. The most important transmission route to patients is through the contaminated hands of healthcare staff working in the hospitals [1, 2]. Methicillin resistant strains of Staphylococcus aureus(MRSA) are also commonly resistant to other antibiotics such as erythromycin, beta-lactamases, and cephalosporin and are capable of producing several virulence factors such as exfoliative toxin, hemolysins, and enterotoxins. Due to the increased prevalence of MRSA in Iran, controlling the infection is more important than the past. S. aureus has undergone significant changes in antimicrobial susceptibility patterns during the past years according to the geographical area. It is noteworthy that after Escherichia coli, it is considered as the second leading cause of nosocomial infections [4]. The resistance to antibiotics among S. aureus isolates has increased injuries and mortality rates in hospitalized patients in this city. Nowadays, 30 to 50 percent of S. aureus strains have become resistant to methicillin. The presence of mecA gene in S. aureus genome, normally carried out in cassette chromosome mec or SCCmec, encodes methicillin resistance ability [3-5].
The present study is implemented in order to evaluate the antibiotic resistance pattern and virulence characteristics of different types of susceptible MRSA using PCR, SCCmec typing, and pulsed field gel electrophoresis (PFGE) methods at Mustafa Khomeini Hospital in Ilam in western Iran. The main goal of this study is to evaluate the antibiotic resistance pattern and virulence characteristics of different types of S. aureus exclusively at Mustafa Khomeini Hospital in Ilam City in order to understand the epidemiology of these infections.
2. MATERIAL AND METHODS
2.1. Samples Collection and Primary Identification
A total of 140 isolates were collected from all parts of hospital indoor air, surfaces, staff and patients through using Quick Take 30 Sample Pump, swab and clinical samples at Mustafa Khomeini Hospital in Ilam from July 2014 to January 2015. All isolates were confirmed as S. aureus by conventional methods including gram stain, catalase, coagulase, DNase, and mannitol fermentation. Further confirmations were obtained through the use of molecular methods including PCR of nucA and spa genes. Isolates that were gram-positive, catalase-positive, coagulase-positive, and DNase-positive and also the ones that carried spa and mecA genes were entered into the study [6, 7].
2.2. Antibiotic Susceptibility Test
Antimicrobial susceptibility patterns were determined by the agar disk diffusion methods according to CLSI guidelines [8]. The tested antibiotics included gentamicin (10μg), erythromycin (15μg), clindamycin (2μg), doxycycline (30μg), minocycline (30μg), ciprofloxacin (5μg), trimethoprim-sulfamethoxazole (5μg), rifampin (5μg), quinupristin-dalfopristin (15μg), and linezolid (30μg) [6-8].
The sensitivity and resistance to methicillin were investigated using cefoxitin (30µg) disk. All MRSAs were further identified by agar dilution of oxacillin according to CLSI guidelines and then confirmed by mecA gene detection using PCR after DNA extraction by phenol chloroform based on the past reports [6-8]. Minimum inhibitory concentration (MIC) of vancomycin was also determined by micro-broth dilution. S. aureus ATCC25923 and ATCC 29213 were used as the internal control as well [6, 7].
2.3. Molecular Detection of Resistant Genes and Virulence Determinants
Molecular detection of resistant and virulence genes was performed through using PCR amplification after DNA extraction through the use of phenol chloroform (PCI) method(Zhang et al., 2005).Specific primers for target genes were conducted using primers sets and cycling conditions that were previously described(Zhang et al., 2005).Antibiotic resistant genes (femA, blaZ, ermA, ermB, ermC, msrA, linA, acc(6')/aph(2”), Ia acc(6')/aph(2”)Ie, aph(3”)IIIa, aph(2”)Ib,aph(2”)Ic, aph(2”)Id, and ant(4')Ia and virulence genes (tst, hlg, etb, sea, hla, hld, see, seb, sec, sed, hlb, eta, pvl) of S. aureus were used in Multiplex-PCR technique according to the past reports [9-11].
Amplification was carried out in a 25µl volume containing 0.5-1mmol of each primer, 0.75µmolof each dNTP, 1.5mM MgCl2, 1 U TaqDNApolymerase, 2.5 µl of PCR buffer, and 3µl of template. The DNA template was replaced with water and then used as negative control. An initial denaturation at 95 ◦C for 5 minutes was followed by 30 cycles of denaturation at 95 ◦C for 60 seconds, and then annealed at 54 ◦C for 120 seconds, and after that extended at 72 ◦C for 120 seconds. Finally, an additional extension was achieved for 5 minutes at 72 ◦C6.For each PCR product, a10 µl aliquot was electrophoresed on a 1.5% agarose gel for 1.5 h at 100 V, stained for 10minutes in ethidium bromide (0·5µg/ml) and then visualized and photographed under UV illumination with Gel Doc apparatus (BioRad. ChemiDoc XRS+ system).
2.4. SCCmec Typing
Multiplex-PCR technique was also used to determine different types of SCCmec with 10 specific primers that were designed by Zhang et al [9]. Moreover, gene amplification was performed with Multiplex-PCR reaction mixture and designed heat cycle by Zhang et al,with a few changes that were optimized by this study [11]. MRSA reference strains including COL (SCCmec type I, ccr1), XU642 (EMRSA-16, SCCmec type II, ccr2), WBG525 (EMRSA-1, SCCmec type III, ccr3), WBG9465 (EMRSA-15, SCCmec type IV, ccr2), and WIS (SCCmec type V, ccrC) were used as controls [12].
2.5. PFGE
(Pulsed-field gel electrophoresis): Analysis of the genetic similarity between S. aureus isolates was performed through using PFGE method in accordance with a previously published protocol [13, 14]. Restriction enzyme digestion was performed with 30 U of SmaI enzyme in Tango buffer (Thermo Fisher Scientific USA) [13]. Electrophoresis was also conducted in a CHEF-Mapper Bio-Rad Laboratories Unit applying parameters including starting pulse (5s), ending pulse (25s), voltage (6V/cm), and run time (19h). After that, Gel-Compar (Applied Maths, Sint-Martens-Latem, Belgium) was used for cluster analysis using the Dice coefficient and unweighted pair group method with arithmetic mean. Isolates clustered ≥ 75% were considered as the same clone [13, 15].
3. RESULTS
3.1. Samples Collection and Primary Identification
A totalof140 gram positive cocci that collect in this period identified as Staphylococcus spp., and 88 of them were then more identified as S. aureus. S. aureus isolate belonged to different samples including surfaces (n=34; 38.9%), staff (23; 26.6%), air (20; 23%) and patients (11;11.9%).
From a total of 36 (40.9%) MRSA isolates that were found,26 (72.2%) were isolated from surfaces, 5 (13.8%) from indoor air, 4 (11.1%) from staff and 1(2.7%) from a patient. All the 36 MRSA isolates were both mecA positive and cefoxitin resistant. The origin of all 36 MRSA isolates was shown in (Fig. 1).
3.2. Antibiotic Susceptibility
The antimicrobial susceptibility patterns of S. aureus (MRAS and MSSA) Isolates to various antibiotics are presented in Table (1). All S. aureus isolates were susceptible to vancomycin, linezolid, and minocycline with resistance rates of 1.1%, 4.5%, and 6.8% that shows good performance against both MRSA and MSSA. As shown in Table (1), resistance to antibiotics was more frequent in MRSA than in MSSA.
| Antibiotic | MRSA n=36 |
MSSA n=52 |
Total n=88 |
||||||
|---|---|---|---|---|---|---|---|---|---|
| No. (%) | No. (%) | No. (%) | |||||||
| R I S | R I S | R I S | |||||||
| Gentamicin | 30(83.3) | 1(2.7) | 5(13.8) | 10(19.2) | 5(9.6) | 37(71.1) | 40(45) | 6(6.8) | 42(47.7) |
| Erythromycin | 30(83.3) | 0(0) | 6(16.6) | 17(32.6) | 2(3.8) | 33(63.4) | 47(53) | 2(2.2) | 39(44.3) |
| Doxycycline | 12(33.3) | 6(16.6) | 18(50) | 4(7.6) | 7(13.4) | 41(78.8) | 16(18) | 13(14.7) | 59(67) |
| Minocycline | 5(13.8%) | 6(16.6%) | 25(69.4%) | 1(1.9%) | 2(3.8) | 49(94.2) | 6(6.8) | 8(9) | 74(84) |
| Ciprofloxacin | 11(30.5) | 6(16.6) | 19(52.7) | 1(1.9) | 2(3.8) | 49(94.2) | 12(13.3) | 8(9) | 68(77) |
| Clindamycin | 19(52.7) | 3(8.3) | 14(38.8) | 7(13.4) | 3(5.7) | 42(80.7) | 26(29.5) | 6(6.8) | 56(63) |
| Trimethoprim Sulfamethoxazole | 14(38.8) | 0(0) | 22(61.1) | 3(5.7) | 0(0) | 49(94.2) | 17(19) | 0(0) | 71(80) |
| Rifampin | 13(36.1) | 0(0) | 23(63.8) | 0(0) | 0(0) | 52(100) | 13(14.7) | 0(0) | 75(85) |
| Quinupristindalfopristin | 1(2.7) | 0(0) | 35(97.2) | 0(0) | 1(1.9) | 51(98) | 1(1.1) | 1(1.1) | 86(97) |
| Linezolid | 1(2.7) | 0(0) | 35(97.2) | 3(5.7) | 0(0) | 49(94.2) | 4(4.5) | 0(0) | 84(95) |
3.3. Frequency of Resistant Genes and Virulence Determinants
All MRSA isolates harbored mecA and femA genes. The predominant AME genes were acc (6')-Ie-aph (2”)Ia (n=25/40; 62.5%), aph(3”)-IIIa (n=12/40; 30%), acc(6')-Ie-aph(2”)Ie (n=11/40; 27.5%), and ant(4')-Ia (n=6/40; 15) among which co-existence of two genes included 35% (n=14/40) of the isolates. According to Table (3), all gentamicin resistant MRSA and MSSA isolates harbored at least one AME gene. However, no other AME gene was detected in the study. The msrA gene was highly distributed among macrolide and lincosamide resistant isolates. Besides, msrA was found in 46.8% (22/47) of erythromycin, 46.6% of clindamycin, and 36.3% of erythromycin and clindamycin resistant isolates.
| Genes | Erythromycin n=47 |
Clindamycin n=15 |
Erythromycin & Clindamycin n=11 |
|||
|---|---|---|---|---|---|---|
| MRSA n=30 |
MSSA n=17 |
MRSA n=9 |
MSSA n=6 |
MRSA n=7 |
MSSA n=4 |
|
| No. (%) | No. (%) | No. (%) | No. (%) | No. (%) | No. (%) | |
| erm A | 5(16.6) | 4(23.5) | 2(22.2) | 2(66.6) | 2(28.5) | 1(25) |
| erm C | 4(13.3) | 1(5.8) | 2(22.2) | 1(16.6) | 1(4.2) | 1(25) |
| erm B | 0(0) | 3(17.6) | 0(0) | 1(16.6) | 0(0) | 1(25) |
| lin A | 0(0) | 2(11.7) | 0(0) | 1(16.6) | 1(0) | 0(2.1) |
| erm A + erm C | 1(3.3) | 0(0) | 0(0) | 0(0) | 0(0) | 0(0) |
| msr A | 7(23.3) | 10(58.8) | 3(33.3) | 1(16.6) | 2(8.5) | 1(25) |
| msr A+ erm C | 3(10) | 0(0) | 2(22.2) | 0(0) | 1(4.2) | 0(0) |
| msr A+ lin A | 0(0) | 2(11.7) | 0(0) | 1(16.6) | 0(0) | 0(0) |
| Total | 20(66.6) | 13(76.4) | 9(100) | 7(100) | 7(100) | 4(100) |
It was also found that 21.2% and 19.1% of erythromycin, 26.6% and 33.3% of clindamycin, and 4.2%of clindamycin and erythromycin resistant isolates contained ermA and ermC.
As shown in Table (2), ermB and linA were found in MSSA but not in MRSA. All clindamycin and also erythromycin and clindamycin co-resistant MRSA and MSSA isolates are carried out at least one resistant gene which was reduced to 66.6% and 76.4% in MRSA and MSSA erythromycin resistant isolates, respectively.
| Gene | MRSA n=36 |
MSSA n=52 |
Total n=88 |
|---|---|---|---|
| No. (%) | No. (%) | No. (%) | |
|
hla hlb hlg hld |
0 (0) | 1(1.9) | 1 (1.1) |
| 0 (0) | 0(0) | 0 (0) | |
| 4(8.3) | 9(17.3) | 13 (14.7) | |
| 0 (0) | 0(0) | 0 (0) | |
|
sea seb sec sed see |
9(25) | 21(40.3) | 30 (34.1) |
| 0 (0) | 0(0) | 0 (0) | |
| 0 (0) | 0(0) | 0 (0) | |
| 0 (0) | 0(0) | 0 (0) | |
| 3(8.3) | 6(11.5) | 9 (10.2) | |
| eta | 8(22.2) | 1(1.9) | 9 (10.2) |
| etb | 1(2.7) | 5(9.6) | 6 (6.8) |
| tst | 11(30.5) | 19(34.6) | 30 (34.1) |
The predominant exotoxins were found to be sea (30/88; 34.1%), tst (30/88; 34.1%), and hlg (113/88; 14.7%). Virulence determinants were revealed to be highly distributed among MSSA isolates. The eta exfoliative toxin was also found in 22.2% of MRSA isolates (Table 4).
| Gene | MRSA n=36 |
MSSA n=52 |
Total n=88 |
|---|---|---|---|
| No. (%) | No. (%) | No. (%) | |
|
hla hlb hlg hld |
0 (0) | 1(1.9) | 1 (1.1) |
| 0 (0) | 0(0) | 0 (0) | |
| 4(8.3) | 9(17.3) | 13 (14.7) | |
| 0 (0) | 0(0) | 0 (0) | |
|
sea seb sec sed see |
9(25) | 21(40.3) | 30 (34.1) |
| 0 (0) | 0(0) | 0 (0) | |
| 0 (0) | 0(0) | 0 (0) | |
| 0 (0) | 0(0) | 0 (0) | |
| 3(8.3) | 6(11.5) | 9 (10.2) | |
| eta | 8(22.2) | 1(1.9) | 9 (10.2) |
| etb | 1(2.7) | 5(9.6) | 6 (6.8) |
| tst | 11(30.5) | 19(34.6) | 30 (34.1) |
3.4. SCCmec Typing
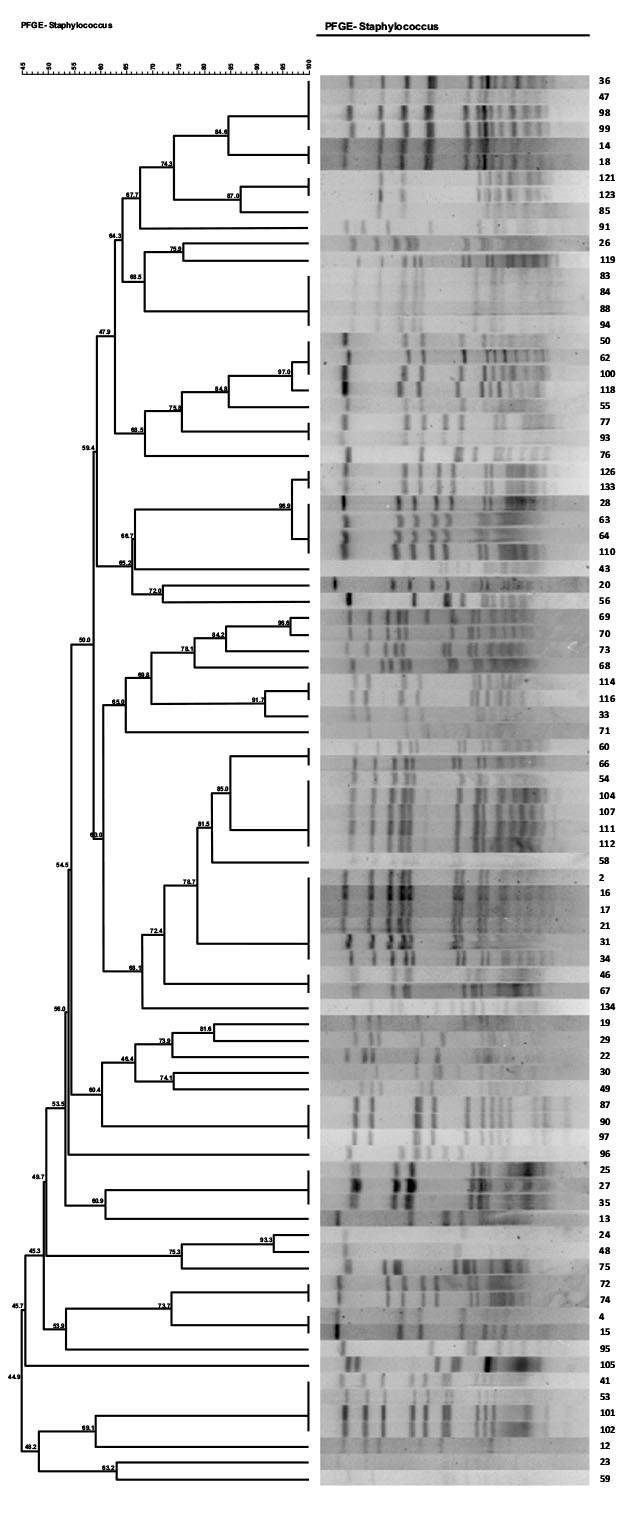
Of 36 MRSA isolates, 11 (30.5%) were identified as SCCmec type I,9 (25%) as SCCmec type II (25%), 8 as SCCmec type IV (22.2%), and 3 (8.3%) as type III (8.3%). Five (13.8%) of the isolates were also not identified as any SCCmec type. The distribution of major SCCmec types according to antibiotic resistance is shown in Dendrogram1 and Table (5). As show in the table SCCmec type I, II and III were more distributed among the isolates and contain more resistant genes and virulence determinant than SCCmec type IV.
| Antibiotic Resistant Phenotypes | No. (%) | Major SCCmec Types No. (%) | Major PulsotypesNo. (%) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | N | 1 | 6 | 12 | 15 | 31 | ||
| CefoxitinC | 36(100) | 9(25) | 10(27) | 7(19.4) | 6(16.6) | 4(11.1) | 4(12.1) | 6(18.1) | 1(3) | 6(18.1) | 2(6) |
| Gentamicin | 30(83.3) | 9(25) | 7(19.4) | 7(19.4) | 5(13.8) | 2(5.5) | 3(9) | 6(18.1) | 1(3) | 5(15.1) | 1(3) |
| Erythromycin | 30(83.3) | 8(22.2) | 7(19.4) | 6(16.6) | 5(13.8) | 4(11.1) | 3(9) | 6(18.1) | 0(0) | 5(15.1) | 0(0) |
| Doxycycline | 12(33.3) | 3(8.3) | 2(5.5) | 4(11.1) | 1(2.7) | 2(5.5) | 1(3) | 2(6) | 0(0) | 3(9) | 0(0) |
| Minocycline | 5(13.8) | 3(8.3) | 1(2.7) | 1(2.7) | 0(0) | 0(0) | 1(3) | 2(6) | 0(0) | 2(6) | 0(0) |
| Ciprofloxacin | 11(30.5) | 5(13.8) | 3(8.3) | 2(5.5) | 1(2.7) | 0(0) | 3(9) | 2(6) | 1(3) | 1(3) | 0(0) |
| Clindamycin | 19(52.7) | 5(13.8) | 3(8.3) | 6(16.6) | 3(8.3) | 2(5.5) | 3(9) | 3(9) | 1(3) | 2(6) | 1(3) |
| Trimethoprim | 14(38.8) | 3(8.3) | 2(5.5) | 3(8.3) | 4(11.1) | 2(5.5) | 2(6) | 2(6) | 1(3) | 2(6) | 1(3) |
| Rifampin | 13(36.1) | 6(16.6) | 2(5.5) | 4(11.1) | 0(0) | 1(2.7) | 1(3) | 2(6) | 1(3) | 1(3) | 1(3) |
| Quinupristindalfopristin | 1(2.7) | 0(0) | 0() | 0(0) | 0(0) | 1(2.7) | 0(0) | 0(0) | 0(0) | 1(3) | 0(0) |
| Linezolid | 1(2.7) | 0(0) | 0() | 1(2.7) | 0(0) | 0(0) | 0(0) | 0(0) | 0(0) | 0(0) | 1(3) |
3.5. PFGE
(Pulsed-field gel electrophoresis). PFGE results showed 33 different pulsotype patterns. The major pulsotypes among MRSA isolates were 15 (6/33;18.1%), 6 (6/33; 18.1%), and 1(4/33; 12.1%). The distribution of major pulsotypes according to antibiotic resistance was shown in Table (5). Pulsotype patterns 15, 6, 8, and 1 had the highest abundance among all MRSA and MSSA isolates. The pulsotype patterns with the lowest abundance included patterns 3, 7, 9, 10, 11, 14, 17, 13, 20, 21, 25, 23, 22, and 30 were singleton. Also, 8 pulsotypes in 11 samples of patients, 17 pulsotypes in 30 isolates of staff, 16 pulsotypes in 24 different isolates of air, and 40 pulsotypes in 16 isolates of surfaces were identified. According to Fig. (1) same pulsotypes such pulsotype 15 were identified among different ward.
| Sample No | Ward | Origin |
Resistance Phenotype |
Pulsotypes | SCC mec | Resistant Genes | Virulence Genes |
|---|---|---|---|---|---|---|---|
| 55 | Women, internal | Bed | FOX, GM,E, D,MI, CIP, RA | 6 | I | blaZ, aph(6)-IIIa | sea, tst |
| 62 | Women, surgery | Computer | FOX, GM,E, CIP,DA, TMP, RA | 6 | I | blaZ | tst |
| 77 | NICU | Air | FOX, GM,E | 6 | II | blaZ, acc(6')-Ie-aph(2”)Ia, ermA | - |
| 93 | Men, internal | Tralee | FOX, GM,E, DA, TMP | 6 | IV | aac(6)Ie-aph(2)-Ie, blaZ, acc(6')-Ie-aph(2”)Ia, aph(6)-IIIa, ermC, msrA | see |
| 100 | ICU | Sink | FOX, GM,E, DA | 6 | I | blaZ, acc(6')-Ie-aph(2”)Ia, ermA | etb, sea, see, |
| 118 | CCU | Bed | FOX, GM,E,MI, CIP, DA, RA | 6 | III | blaZ, acc(6')-Ie-aph(2”)Ia | tst |
| 31 | CCU | Station | FOX, GM,E, D,MI, RA | 15 | II | blaZ | eta |
| 34 | Women, surgery | Mobile staff | FOX | 15 | N | blaZ | eta |
| 54 | ICU | IPhone | FOX, GM,E, D, CIP,DA, TMP | 15 | III | blaZ | sea, tst .eta |
| 104 | Men, Post CCU | Air | FOX, GM,E,MI | 15 | I | blaZ, linA, msrA | - |
| 111 | Women, internal | Station | FOX, GM,E | 15 | IV | blaZ, ermB | tst |
| 14 | CCU | Ventilator | FOX, GM,E, CIP, DA, RA | 1 | I | aac(6)Ie-aph(2)-Ie, blaZ, acc(6')-Ie-aph(2”)Ia, aph(6)-IIIa, ermA | see, tst, sea |
| 36 | CCU | Air | FOX, GM | 1 | III | blaZ,, ermA, ermC | tst, eta |
| 47 | CCU | Ventilator | FOX,E, CIP, DA, TMP | 1 | N | blaZ, acc(6')-Ie-aph(2”)Ia, aph(6)-IIIa, msrA | sea .eta |
| 99 | NICU | Incubator | FOX, GM,E,MI, CIP, DA, TMP | 1 | II | blaZ, acc(6')-Ie-aph(2”)Ia | see, hlg, sea |
| 25 | CCU | Staff=Men | FOX, GM,E, CIP, DA, RA | 24 | N | blaZ, acc(6')-Ie-aph(2”)Ia | hlb |
| 27 | CCU | Monitor | FOX, | 24 | I | blaZ | tst, hlg |
| Continue Table 6. Phenotypic and genotypic features of MRSA isolates in this study. | |||||||
| Sample No | Ward | Origin |
Resistance phenotype |
Pulsotypes | SCC mec | Resistant Genes | Virulence Genes |
| 35 | CCU | Air | FOX, E, DA, RA | 24 | II | blaZ | tst, hlg, eta |
| 53 | NICU | Station | FOX | 31 | II | blaZ | tst, seb, sea |
| 102 | Emergency department | Ventilator | FOX, GM,E, DA, TMP, RA, LNZ | 31 | III | blaZ | - |
| 24 | CCU | Staff=Men | FOX, GM,E, CIP, DA, RA | 26 | II | blaZ, acc(6')-Ie-aph(2”)Ia | tst, hlg |
| 75 | Operating room | Medicine cabinet | FOX, GM,E | 26 | I | blaZ | hlg, tst |
| 4 | CCU | Staff=Men | FOX, GM,E | 28 | II | blaZ, acc(6')-Ie-aph(2”)Ia, ant (4')-Ia, ermC, msrA | tst, hlg, sea |
| 20 | Men, Post CCU | Staff=Men | FOX, GM, | 10 | IV | blaZ | tst |
| 28 | NICU | Medicine cabinet | FOX, GM,E, CIP, DA, RA | 8 | II | blaZ, ant (4')-Ia, msrA | sea |
| 29 | Women, surgery | Bed | FOX, GM,E, D, TMP | 18 | N | blaZ, acc(6')-Ie-aph(2”)Ia,, ermA | tst |
| 30 | NICU | Station | FOX, GM,E, D, TMP | 20 | I | blaZ, acc(6')-Ie-aph(2”)Ia, aph(6)-IIIa | - |
| 43 | ICU | patient=BC | FOX, GM,E, DA, RA | 9 | IV | blaZ, ant (4')-Ia | tst, eta |
| 43 | ICU | patient=BC | FOX, GM,E, DA, RA | 9 | IV | blaZ, ant (4')-Ia | tst, eta |
| 56 | Women, internal | Refrigerator | FOX, GM,E,DA, TMP | 11 | IV | blaZ | sea |
| 68 | Men, internal | Refrigerator | FOX, GM | 12 | IV | blaZ | sea, tst |
| 72 | NICU | Station | FOX, GM,E, DA | 27 | III | blaZ | tst, hlg |
| 83 | Men, Post CCU | Air | FOX,E, TMP | 5 | II | blaZ, ermA | - |
| 90 | ICU | Cover records | FOX, GM,E, D, CIP, DA, RA | 22 | III | blaZ | - |
| 91 | Women Surgery | Floor | FOX, GM,E, CIP, DA, TMP, RA | 3 | I | aac(6)Ie-aph(2)-Ie, blaZ, acc(6')-Ie-aph(2”)Ia, ermC, msrA | sea |
| 96 | CCU | Tralee | FOX, GM,E, D, TMP | 23 | III | blaZ | see |
4. DISCUSSION
Nosocomial infections are among the important factors of patient's mortality in Iran and other countries around the world [16, 17].
Bacterial identification and their antibiotic resistance are significant for the prevention and treatment of bacterial infections. Hospital environments have been widely identified as the main source of multidrug-resistant bacteria most importantly MRSA strains [18]. The development of bacterial resistance was linked with antibiotic use and hence selective pressure which was specific for the type of antibiotic and the bacterial species [18]. Antibiotics arguably constitute the most concentrated selective pressure ever brought to bear on S. aureus in its long co-evolutionary history with mankind. The consequences of this selective pressure in conjunction with horizontal and vertical gene transfer are the subject of this review. Given their critical importance as therapeutic agents, the story will focus on resistance to penicillin’s and the structurally related beta-lactam antibiotics [19].
The resistance to antibiotics is one of the main threats to human life in the present century [20]. Bacterial contamination of different parts of hospitals is due to various factors such as physical structure, devices and medical equipment multiplicity, inefficient purification systems, presence of certain diseases, and etc. Based on the guidelines of Infection Control Committee of the hospital, hospitalization of patients more than fifteen days in different units of hospital including intensive care units, enhances the risk of nosocomial infections especially by MRSA strains [21]. These clonal complexes were widespread prior to emergence of methicillin resistance indicating that superior epideictic preceded acquisition of drug resistance and that the adaptations and innovations that make clones successful also may favor their adaptation to antibiotic selective pressures [22].
The reasons for the disparity in rates of quinolone resistance between MSSA and MRSA strains are uncertain. One contributing factor is likely antibiotic selective pressure, especially in the hospital setting, resulting in the selection and spread of the more antibiotic-resistant MRSA strains [10].
Several studies have been conducted worldwide on toxic genes of MRSA from various dimensions. In similar clinical studies, the frequency of sea gene of MRSA was 58.8% in Gorgan-Iran, 74.4% in Tehran-Iran, 33% in China, and 12% in Germany, while the frequency of sea gene in MRSA isolates from various specimens obtained in this study was 25% which is similar to that of the study conducted in Korea [10, 23-26]. Moreover, the frequency of sub gene has been investigated in many studies. The frequency of this gene in Gorgan, Tehran, China, Canada, Korea, Czech, and Colombia was 61.3%, 73.58%,5%, 15.78%, 5.6%, 3%, and 7%, respectively [27-29]. In this study, the frequency of sea gene was found to be 8.3%. Another resistance mechanism that was evaluated was macrolide resistance among MRSA isolates. In accordance with an earlier report from Iran [21], resistance rates to erythromycin and clindamycin in this study were discovered to be 50% and 27%, respectively. Also, the resistance rates to erythromycin and clindamycin were 57% and 17%, respectively which were lower than those reported in other studies [30, 31].
Earlier studies have indicated that ermA (13.8%) has the most important role in macrolide resistance [16]. However, this study showed ermC (11.1%) to be the most prevalent gene among macrolide-resistant MRSA in West of Iran and that ermB didn’t play an important role in macrolide-resistant MRSA which is contradictory to the previous reports from Iran [32].
Similar to earlier studies, no ermB gene was found among macrolide-resistant MRSA isolates in the current study. In similar studies, the prevalence rate of ermB was seen in just a few erythromycin-resistant staphylococci [33]. The results also showed that the resistance to erythromycin was due to the existence of ermA among MRSA isolates in Ilam.
The study of antibiotic resistance identified the abundance of several macrolide resistant genes including ermC, ermA, ermC, ermB and linA in this work. However, the frequency of these genes appeared to be lower than that in the previous study by Aktas et al [34]. After ermC gene with 10.2% abundance, ermA, ermB, and linA genes with respective 5.7%, 4.5%, and 2.3% abundances, were identified as the most common macrolide resistance genes in this study which were lower than the obtained results by Aktas et al [34]. Gene aac (6')-Ie-aph (2”)-Ia was the most common aminoglycoside resistance gene with 22.7% abundance. Subsequently, the aph (3”)-IIIa gene was the second most common one with12.5% abundance. In the current study, as confirmed in many reports [32-34], acc (6')-Ie-aph (2”)Ia was the most prevalent AME gene that was encountered in more than 75% of all aminoglycoside-resistant isolates. However, reports from Kuwait and Japan indicated that the prevalence of ant (4')-Ia gene is much higher than that of the other three AME genes found in the present study). The most frequently encountered AME in staphylococci is acc(6')-Ie-aph (2”)Ia which deactivates the wide range of medically important aminoglycosides. Moreover, 46.6% of MRSA isolates in this study were also positive for acc(6')-Ie-aph (2”)Ia. The majority of these isolates (95%) was resistant to all tested aminoglycoside antibiotics [35]. The second most detected AME gene in this study is aph(3”)-IIIa and the enzyme encoded by it confers resistance to amikacin, but not gentamicin. The rate for ant(4')-Ia in this study (10%) was lower than those reported from Kuwait (87%) and European countries (53%) [36]. However, the rates of aph(3”)-IIIa and ant(4')-Ia differ greatly among different countries. Except for 7 aminoglycoside-resistant strains, the rest contained AME genes in this study revealing the importance of these enzymes in the modification of aminoglycoside antibiotics among staphylococci in Ilam.
Compared to other studies, a much higher frequency of tst gene was found in this study which raised a significant concern for its sanitation and hygienic practices [36]. Immediate regulatory action should be taken to reduce its nosocomial infection rate and prevent the spread of S. aureus among patients and staff [35, 36]. Given that most isolated strains of this gene existed on the surfaces, regulatory actions of authorities are necessary to be taken in order to control the environmental health in hospitals and prevent the spread of these strains among patients and staff. The frequency rates of eta gene reported from Germany, Czech Republic, Turkey, and Colombia were 2%, 10%, 19.2%, and 3%, respectively [36-38] while it was equal to 22.2%in this study. The frequency of etb gene in a study carried out in Turkey was 9.2%. In Colombia, however, no etb gene was detected among 30 MRSA [38]. The frequency of the etb in this study was also obtained 6.8%.
The most predominant hemolysin gene among MRSA and MSSA isolates was hlg gene. However, the frequencies obtained in this study (84.24% abundance) were lower than those obtained in the study by Kim et al (93.15% abundance) [39]. The frequency of hla gene, in a study by Hoseini Alfatemi et al. in Shiraz was reported to be 93.1% [11].In this study however, hla gene with a frequency of 93.15% was the most abundant one. According to studies conducted in other parts of the world, we can conclude that this gene is comparatively much more frequent in MRSA isolates [11].
In the present study, four types of SCCmec were identified among 36 strains of MRSA. However, SCCmec type V was not detected in isolated strains. The strains with the highest abundance were SCCmec type I with 30.5% and then types II, IV, and III with 25.0%, 22.2%, and 8.3%, respectively.
Similar results were reported in Spain by Pereze et al (Shukla et al., 2010). (1998-2000) and 2.9% of 375 MRSA strains were not typed in this study. In the reports of Chung et al. (2004) and Zhang et al., among all types, type IV has the highest abundance (50%). Kim et al. (2006) in South Korea reported the abundance of type IV to be 68.8% which is higher than the obtained results in this study [40].
In another study, it has been reported [41] that type III with 73% abundance was the most detected SCCmec type [42].
In this study, the most abundant SCCmec type was related to CCU with 36.3%. All these strains were isolated from shock devices, monitors, and patient’s urine. Accordingly, various hospital surfaces and patients contributed to circulation of these strains. Therefore, proper routines sanitation of surfaces should be immediately implemented [40, 42].
The highest abundance of SCCmec type II and IV was seen in the CCU and the lowest abundance was observed in obstetrics surgery and emergency units. The presence of these similar strain types in different units of a hospital strongly suggests the contamination and circulation of these strains between surfaces and patients [42].
According to the results obtained in this study, SCCmec type II had the highest rate (62.5%, 8/5) among various SCCmec types. Strains isolated from staff also had the highest SCCmec type III (60%, 5/3).Similarly, the most frequent strain in air and surfaces was SCCmec type IV (50%, 2/1).It has been already shown that SCCmec types I to III are frequently isolated from hospital-acquired MRSA and often the mecA gene is one of the additional genes playing a role in resistance to several beta-lactam antibiotics [42]. Several reasons for gaining additional resistant genes by these isolates have been reported in medical centers that could explain the high antibiotic resistance of SCCmec type I-III isolates in this study. Moreover, the significant antibiotic susceptibility pattern of certain SCCmec type isolates obtained in this study can prompt us to consider these antibiotics as an option for the treatment of infection caused by such isolates. Most type IV isolates in this study were susceptible to the majority of the tested antibiotics. However, similar to some studies, a few isolates containing type IV cassette were also multidrug-resistant in this study. This suggests that since type IV isolates are often exposed to antibiotics, they could gain antibiotic resistant genes other than mecA gene to survive in medical centers [7]. The frequency of SCCmec type I isolates in the present study shows the emergence of this type in the studied medical centers. As was previously mentioned in the results section, due to the significant clinical sources and the antimicrobial susceptibility patterns of different SCCmec type isolates, optimizing institutional infection control policies for preventing MRSA transmission among hospitalized patients must be considered [7, 12]. Several molecular typing methods such as DNA fingerprint by PFGE and MLST are best known for typing MRSA isolates. As a gold standard typing technique, it has been used for its reliability, discriminatory, and reproducibility in regional studies [12, 40, 42, 43]. It has also been stated that PFGE of S. aureus is the most discriminant genotyping tool and a good method to resolve clonal relationships. Javidnia et al reported 4 different types of S. aureus isolated from intensive care units (ICU) of three different hospitals in Tehran. In their study, PFGE typing method revealed differences in pulsotypes similar to the results obtained in this study [40]. Moreover, in another similar study conducted by Emaneini et al using PFGE in Tehran hospitals, different pulsotypes were isolated with7 types being detected in only one isolated S. aureus [18].
Rahimi et al also reported that diverse pulsotypes consisting of 13 common types and 18 single types along with seven common PFGE types were found among the MRSA strains [33]. Lastly, in a study by Aris de Sousa in Portugal, 271 MRSA isolates were typed by the PFGE typing method and 13 types and 63 subtypes were detected, 5 types of which were typed at least in ten isolates [44]. The results of these studies showed that the origin of all isolates was not identical and there was no significant relationship between them which are consistent with the results obtained in the present study. Moreover, the results of this study indicated that the abundance of pulsotype patterns was different in isolated strains of different samples taken from the hospital. The obtained results also showed that there was no significant correlation between pulsotype patterns of isolated strains in patients and isolated strains of staff, hospital-air, and hospital surfaces. Moreover, different isolates of S. aureus were not of the same clonal origin(Aires de Sousa de Lencastre. 2004). Pulsotype patterns of S. aureus showed that 75% of pulsotype patterns was different. However, different pulsotypes were observed in this study with no significant relationship between them. Based on the obtained results, pulsotypes1, 2, 11, and 16 were observed in all the understudy parts of the hospital which expressed a relationship between S. aureus isolates in different populations. These findings reflect the urgent need to control and eliminate S. aureus infection. Accordingly, rigorous regulations and policies should be adopted for sanitation or disinfection programs. Proper indoor air ventilation is also needed and minimal antibiotic therapy should be administered among patients [6, 7, 18, 44].
Due to their high discriminatory power, PFGE techniques are considered as a useful tool for determining the prevalence of S. aureus infections particularly in local hospitals. In the past, it was difficult to analyze and compare the obtained information of PFGE from different hospitals and evaluation of the obtained results for each hospital was only used for itself.
CONCLUSION
The obtained results in this study, like those of other studies conducted in Iran and other parts of the world, show high prevalence of S. aureus isolates in various units of hospitals, equipment, staff and patients. The PFGE typing technique is used as a standard molecular typing method to determine the relationship between isolated S. aureus of different parts in the hospital.
The obtained results indicated that the highest and lowest abundance of S. aureus was found on hospital surfaces and in patients, respectively. These findings confirm two points. First, surfaces are the main sources of contamination and infection in the hospitals as they transfer the center of S. aureus isolates into different parts of a hospital. Second, in Mostafa Khomeini Hospital in Ilam, it was more likely for bacteria to be transferred from the hospital environment to patients than the other way around. Different pulsotypes were also identified in this study with some having greater abundance than the others which describe the possibility of a common genetic origin of the isolates. In order to improve and complete the treatment and reduce the incidence of MRSA and also to limit the spread resistance and accurate diagnosis, the following suggestions are offered. Antibiotic resistance and virulence factors should be annually surveyed and clone resistance should be identified. Moreover, hospital entries and exits should be controlled. We also found that the hospital staff and surfaces have very important role in the transmission of infection into the hospitals and can be a way to transfer S. aureus strains among different wards of hospitals.
FUNDING
No funding.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
This study received ethical approval from Ilam University of Medical Sciences code number 919004/42.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies that are base of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors announce that the research was conducted in the absence of any commercial or financial relationship that could be construed as a potential conflict of interest.
ACKNOWLEDGEMENTS
The authors would like to thank the dedicated laboratory personnel at the Ilam University of Medical Sciences and Lorestan University of Medical Sciences who helped collect specimens for this study.


