All published articles of this journal are available on ScienceDirect.
Genetic Investigation of Beta-Lactam Associated Antibiotic Resistance Among Escherichia Coli Strains Isolated from Water Sources
Abstract
Background:
Antimicrobial resistance is an important factor threatening human health. It is widely accepted that antibiotic resistant bacteria such as Escherichia coli (E. coli) released from humans and animals into the water sources, can introduce their resistance genes into the natural bacterial community.
Objective:
The aim of this study was to investigate the prevalence of blaTEM, blaCTX, blaSHV, blaOXA and blaVEB associated-antibiotic resistance among E. coli bacteria isolated from different water resources in Iran.
Methods:
The study contained all E. coli strains segregated from different surface water sources. The Kirby-Bauer method and combined discs method was determined in this study for testing antimicrobial susceptibility and strains that produced Extended-Spectrum Beta Lactamases (ESBL), respectively. DNA extraction kit was applied for genomic and plasmid DNA derivation. Finally the frequency of resistant genes including blaTEM, blaCTX, blaSHV, blaOXA and blaVEB in ESBL producing isolates were studied by PCR.
Results:
One hundred E. coli strains were isolated and entered in the study. The highest antibiotic resistance was observed on clindamycin (96%). Moreover, 38.5% isolates were ESBL producers. The frequency of different ESBLs genes were 37%, 27%, 27%, and 25% for blaTEM, blaCTX, blaSHV, and blaOXA, respectively. The blaVEB wasn’t found in any isolates.
Conclusion:
The study revealed a high prevalence of CTX-M, TEM, SHV and OXA genes among E. coli strains in surface water resources. In conclusion, these results raised a concern regarding the presence and distribution of these threatening factors in surface water sources and its subsequent outcomes.
1. INTRODUCTION
The increase in number of resistant strains of bacteria and transmission of resistance genes from environmental bacteria to human pathogenesis are an important factor threatening human public health, so that become a major concern health authorities worldwide [1, 2]. Extended-spectrum beta-lactamases (ESBL) is responsible for creating resistant strains of bacteria against antibiotics [3-5]. The spread of ESBL-producing strains limited to antimicrobial agents for the treatment of patients effectively and there is a concern on Escherichia coli (E. coli), and most of other pathogen such as, Shigella, Salmonella, and Klebsiella [6-9]. E. coli associated diseases and water-borne outbreaks result in high morbidity and mortality worldwide [10]. Bacterial contamination of surface water resources has long been considered as a water quality issue which can be contribute in disease transmission. Bacteria resistant to antibiotics are one of the most dangerous biological contaminants in surface water sources [1]. Hydrolysis of β-lactam ring structure by ESBL caused antibiotic function impairment. ESBL enzymes are able to target a broad spectrum of antibiotics, such as penicillins and their complexes, monobactams, and new generation cephalosporins [11]. There are hundreds types of β-lactamase enzymes, each ESBL bacterium may possess genes for one or more of these enzymes. Similar bla genes with alike structure are grouped together within these gene families in terms of phylogenetic and by apply PCR are often targeted for detection and identification. TEM and OXA as a kind of ESBL, contain resistance genes against β-lactamase inhibitors, creating such species more of a threat [12]. Transfer of resistance genes between bacteria is easily facilitated because the genes are encoded on mobile vectors (plasmids and transposons) [13, 14].
The use and misuse of antimicrobial drugs in hospitals can lead to development of antibiotic resistant bacteria and the spread of resistant organisms such as E. coli in the environment including swage and surface waters. E.coli is a potential uropathogenic bacterium which may cause a wide range of Urinary Tract Infections (UTIs) including asymptomatic and/or symptomatic bacteriuria, cystitis, and pyelonephritis both in children and adults. Moreover, detection of this organism in water is an implicit indicator of fresh fecal pollution and consequently of the hazard of co-occurrence of enteric pathogens that can reason illness in susceptible populations. Access to adequate amounts of safe and healthy foods is key to supporting life and promoting good health of people [15, 16]. A high incidence of Multiple Antibiotic Resistances (MAR) has been detected in E. coli samples isolated from river and contaminated food or water. MAR in intestinal bacteria is originating from wastewater treatment plants [17-20]. This problem is clinically important in patients with problems associated with E. coli, for example: uropathogenic E. coli strains had the highest resistance to penicillin (100%) and ampicillin (80.20%) while the lowest resistance rate was observed to imipenem (0%) and nitrofurantoin (5.69%) [21-23].
Analysis of phenotypic susceptibility to penicillin and cephalosporins can obtain an accurate assessment of β-lactamase producing Enterobacteriaceae [24]. β-lactamases are modifying enzymes (drug-inactivating enzymes) found in all types of bacteria. The efficiency of β-lactam antibiotics in eradicating infection at various body sites can diminish by β-lactamase-producing organisms [25].
By use of DNA-based technologies, a new era seems to be opening in the field of diagnostic and molecular epidemiology of antibiotic resistant bacteria. The application of PCR based techniques has had a revolutionary impact in rapid detection of resistance determinants, such as ESBLs genes [26, 27].
This study was conducted for evaluating the prevalence of blaTEM, blaCTX, blaSHV, blaOXA and blaVEB associated-antibiotic resistance among E. coli strains which are isolated from different water resources in Iran.
2. MATERIALS AND METHODS
A total of 100 water samples which have been taken earlier from different water sources during one year between September 2012 and September 2013 was included for the study [28].Water samples were collected in the sterilized bottles with a capacity of 200 ml. The multiple tube fermentation method was considered for sampling of the coliform group components and presence–absence coliform test. The water samples were chilled on ice packs immediately after collection to prevent any changes in the microbial flora of the samples during transportation to laboratory. Samples were analyzed within 2 h of collection. In order to hold E. coli isolates until the end of collecting samples process, all isolates in culture media of brain-heart infusion (BHI) contained 25% glycerol and were maintained at a temperature of -70°C.
Each water samples were inoculated in Lauryl sulfate broth and PA broth, respectively. All the inoculated tubes were incubated at 35±0.5ºC for 24-48hrs. Following turbidity and gas in the durham tube and create an acidic pH in the samples, the presence of coliform was primarily approved and given to the next-step verification for further consideration. All samples showing production of gas, acid and turbidity were cultured in brilliant green bile broth and incubated at 35±0.5ºC for 24-48hrs. The presence of coliforms in samples confirmed by gas production and acid pH. In the next step of diagnosis process, all samples showing production of gas, acid and turbidity were cultured in EC broth and incubated at 44.5±0.2ºC for 24 hrs. The presence of coliforms in samples that tolerated to heat confirmed by gas production and acid pH. In the next step, the samples that were positive in this phase were cultured on selective culture media of eosin methylene blue agar and incubated at 35±0.5ºC for 24-48 h. The presence of E. coli was confirmed by observation of colonies with green metallic sheen. All colonies containing metallic sheen were tested by IMViC test and if the results were compatible, they would have been isolated and storaged as strains of E. coli.
The dilution and disk-diffusion methods were applied in this study for evaluation of antimicrobial susceptibilities of the E. coli bacteria. Antibiotic discs were placed within a distance of 12 mm from each other and incubated at 37ºC for 18 hrs. To determine the production of ESBL enzymes in accordance with the instructions of CLSI, the screening method of combined discs was used. Antibiotic discs were placed within a distance of 15 mm from each other and incubated at 37ºC for 24 hrs. Antibiotics, including clindamycin, lincomycin, rifampin, ceftazidime, streptomycin, cefazolin, tetracycline, gentamicin, nalidixic acid, kanamycin, trimethoprim, tobramycin, ciprofloxacin, cloramfenicol, imipenem, norfloxacin, amikacin, azithromycin, and nitrofurantoin were tested in this study. The phenotypic confirmatory test was applied for identification of ESBL producers by using cefotaxime and ceftazidime individually and also combined with clavulanic acid [29].
A kit that extract AccuPrep® genomic DNA (Bioneer, South Korea) was applied for extraction of DNA. Absorbance of the samples at 260 nm examined the DNA concentration [30]. The five specific primers [31, 32] were selected to expanding genes under this study and for each one a forward primer and a reverse primer was chosen (Table 1). PCR reaction was performed in a DNA thermocycler as follows: Primary denaturation at 96ºC for 5 min, denaturation by 30 cycles at 95ºC as long as 1 min, annealing at 55ºC within 1 min (based on the type of primer for each gene is different from that given in Table 1), elongation at 72ºC as long as 1 min and extension phase at 72ºC for 7 min, completed by a hold at 4ºC. PCR products were electrophoresed on 1.5% agarose gel containing ethidium bromide at 80 V for 1 h.
| Target Type of β-Lactamase Genes | Primer Sequence |
Size of Product (bp) |
AT (ºC) |
|---|---|---|---|
| blaSHV | 5´-TTA ACT CCC TGT TAG CCA-3´ (F) | 860 | 51.2 |
| 5´-GAT TTG CTG ATT TCG CCC-3´ (R) | |||
| blaTEM | 5´-ATG AGT ATT CAA CAT TTC CG-3´ (F) | 867 | 52.2 |
| 5´-CTG ACA GTT ACC AAT GCT TA-3´ (R) | |||
| blaOXA | 5´-ACA CAA TAC ATA TCA ACT TCG -3´ (F) | 885 | 60 |
| 5´-AGT GTG TTT AGA ATG GTG ATC-3´ (R) | |||
| blaVEB | 5´-CGA CTT CCA TTT CCC GAT GC-3´ (F) | 1014 | 55 |
| 5´-GGA CTC TGC AAC AAA TAC GC-3´ (R) | |||
| blaCTX | 5´-TTT GCG ATG TGC AGT ACC AGT AA- 3´ (F) | 560 | 60 |
| 5´-CTC CGC TGC CGG TTT TAT -3´ (R) |
3. RESULTS
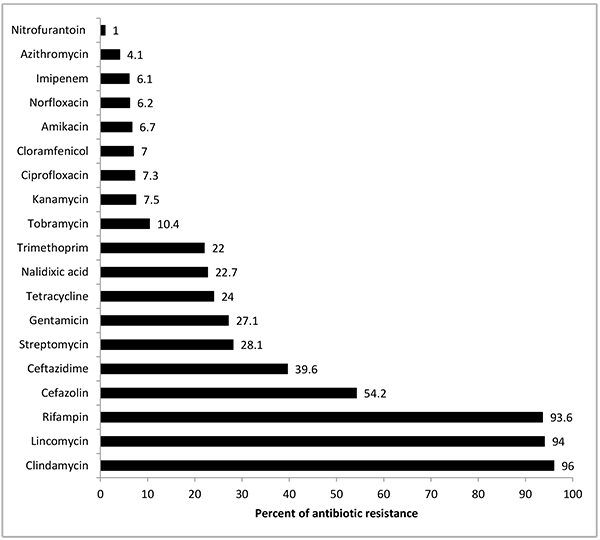
Totally, 100 E. coli isolates were isolated. In the present study, the antibiotic resistance rates to different antibiotics is given in Fig. (1). The highest and lowest rates of antibiotic resistance were related to clindamycin (96%) and nitrofurantoin (1%), respectively.
The frequency of resistance genes were 37, 27, 27, and 25 into different strains based on the detection of the various resistance genes including blaTEM, blaCTX, blaSHV, and blaOXA, respectively. The blaVEB gene was not distinguished at all items. Also, at the same time, 37% of the samples showed resistance to more than one gene (11, 12, and 14 percent of isolates showed resistance genes to two, three and four ones).
4. DISCUSSION
E. coli bacteria through water plays a role in the transmission of different types of diseases. Presence of this bacteria in the water, proceeds to the release of these bacteria in the environment which involved in antibiotic resistance. This resistance can be due to the presence of specific genes of ESBL such as, blaTEM, blaCTX, blaSHV, blaOXA and blaVEB. Knowing the types and frequency of these genes made a decision for the treatment process of patients effectively.
The present study investigated the genetic study of beta lactam associated antibiotic resistance among E. coli bacteria from water resources and reported rate of resistance to antibiotics. High level of resistance in 100 isolates of this bacteria isolated from different water resources mostly shown to clindamycin (96%), lincomycin (94%), rifampin (93.6), cefazolin (54.2) and ceftazidime (39.6). ESBL genes and the frequency of blaTEM, blaCTX, blaSHV and blaOXA genes were apprehended, while Kim et al. [33] detected sixteen strains of E. coli that were highly resistant to about 10 antibiotics. Resistance to cephalothin, ampicillin, tetracycline, streptomycin, sulfixosazole, trimethoprim, kanamycin, and gentamicin have been shown in all ESBL-producing E. coli strains. Further, Kim et al. [33] only reported blaTEM and blaCTX as ESBL. Jang et al. showed that with the dominance of the blaCTX, beta-lactamase gene had more than one gene in 67% of cases [34]. In another study that was conducted on the Thames river water samples, the greatest resistance mainly caused by blaCTX-M14 gene for E. coli that used ciprofloxacin and ampicillin [35].
Toroglu et al. [36] isolated 67 strains from Aksu river which 67,2% were E. coli organism. Multiple antibiotic resistances were determined in 27 isolates (40%) in which they were resistant to five or more antibiotics and 49,3% of the isolates were found to be Beta-lactamase producing bacteria. A high antibiotic resistance of E. coli to antibiotics was indicated in this study.
ESBLs and carbapenemases considered as important mechanisms in Enterobacteriaceae that cause modern broaded-spectrum cephalosporin and carbapenem resistance. A great concern is the extension of ESBL and carbapenemase producers into the environment. ESBL-producing Enterobacteriaceae were detected in 21 (36.2%) of samples. The expression of an ESBL phenotype was e shown (74 strains confirmed). SHV-12 ESBL producers were identified in three different strains, the genes encoding CTX-M ESBLs were presented in 71 strains [37].
The presence of circumstance of CTX-M (84.2%) and TEM (47.4%) genes in isolates from urban surface waters in Malaysian were mostly E. coli and K. pneumoniae [38].
Change of antibiotic resistance following the change in water quality is leading to negative effects on Public health. High frequency of resistant bacteria against amoxicillin and ampicillin has been reported by Muleta T. et al. [39]. Haque et al. examined surface water in Dhaka, revealing E. coli as the most prevalent isolated among Enterobacteriaceae. The spread of multidrug resistant bacteria may be affected by CTX-M and SHV types ESBL genes [40].
From the 767 E. coli isolates in wastewater treatment plants, the highest resistance rates were found for ampicillin and piperacillin; cefalothin and cefuroxime-axetil; nalidixic acid; trimethoprim/sulfamethoxazole and tetracycline [41].
Chatterjee et al. revealed that susceptibility rate was high (i.e. 85%) in case of Cefixime. It was 75% and 30% against Streptomycin and Ciprofloxacin among the twenty E.coli isolates, respectively. High level of resistance was reported over Doxycycline, since all isolates were found to be resistant against this antibiotic. Cefixime was detected as the most impressive antimicrobials against E. coli isolates [42].
In addition, the results of 27,886 isolates of E. coli from Minjiang river in China showed most resistant to cefoperazone, cefazolin, ciprofloxacin, norfloxacin and rifampicin [43].
To the best of our knowledge, this is the first study on evaluation of the prevalence of blaTEM, blaCTX, blaSHV, blaOXA and blaVEB associated antibiotic-resistance among E. coli bacteria isolated from different water resources in Alborz province of Iran. This study revealed high levels of antibiotic resistance and prevalence of blaTEM, blaCTX, blaSHV and blaOXA genes that were detected in E. coli isolations from water samples. Increasingly, the risk of antibiotic resistance gene transfer rises among bacteria in the water sources. This is a serious warning, because ESBLs are an eminent threat to the efficacy of antibiotics that are available in current medical uses.
E. coli producer isolates have now risen to prominence, mainly associated with community-acquired infections. Iran, like many other regions of the world, has experienced a significant increase in the numbers of ESBL-producing bacteria in the new millennium. The high prevalence of ESBL genes in Iran means that the empirical treatment of serious infections with beta-lactam antibiotics is seriously compromised. In addition to human and animal health risks from infectious microorganisms, potential hazards associated with the application of sewage sludge to agricultural land include contamination of surface and ground waters. In this paper, we presented evidence of a greater frequency of fecal coliforms and drug-resistant bacteria in surface waters.
Our results revealed a high prevalence of ESBLs genes among E. coli strains in surface water resources. The antimicrobial drug resistance that is increasingly compromising the usefulness of the b-lactam antibiotics and other previously life-saving antibacterial drugs.
CONCLUSION
In conclusion, these results raised a concern regarding the presence and distribution of these threatening factors in the surface water sources and its subsequent outcomes. Prevention and control strategies should be urgently implemented to stop further spreading of these strains.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies that are base of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We would like to thank from the “Clinical Research Development Center of Baqiyatallah hospital” for their kindly cooperation. This study was supported in part by a grant from “Clinical Research Development Center of Baqiyatallah hospital”.


