All published articles of this journal are available on ScienceDirect.
Adaptation of Mycobacterium smegmatis to an Industrial Scale Medium and Isolation of the Mycobacterial PorinMspA
Abstract
The adaptation of the organism to a simple and cost-effective growth medium is mandatory in developing a process for large scale production of the octamericporinMspA, which is isolated from Mycobacterium smegmatis. A fermentation optimization with the minimal nutrients required for growth has been performed. During the fermentation, the iron- and ammonium chloride concentrations in the medium were varied to determine their impact on the observed growth rates and cell mass yields. Common antibiotics to control contamination were eliminated in favor of copper sulfate to reduce costs. MspA has been successfully isolated from the harvested M. smegmatisusing aqueous nOPOE (n-octyloligooxyethylene) at 65°C. Because of the extraordinary stability of MspA, it is possible to denature and precipitate virtually all other proteins and contaminants by following this approach. To further purify the product, acetone is used for precipitation. Gel electrophoresis confirmed the presence and purity of MspA. A maximum of 840µg (via Bradford assay) of pure MspA per liter of the optimized simple growth medium has been obtained. This is a 40% increase with respect to the previously reported culture medium for MspA.
INTRODUCTION
MspA is an octamericporin in the outer membrane of M. smegmatis [1-3].The inner pore of MspA is the major general diffusion pathway for the organism and provides a passage for hydrophilic solutes. The crystal structure of MspA shows the assembly of eight monomers to a goblet shaped pore [4]. Each monomer contains two consecutive 16-stranded β-Barrels with hydrophobic outer surfaces (the so-called constriction zone, which is marked with an arrow in Fig. 1).
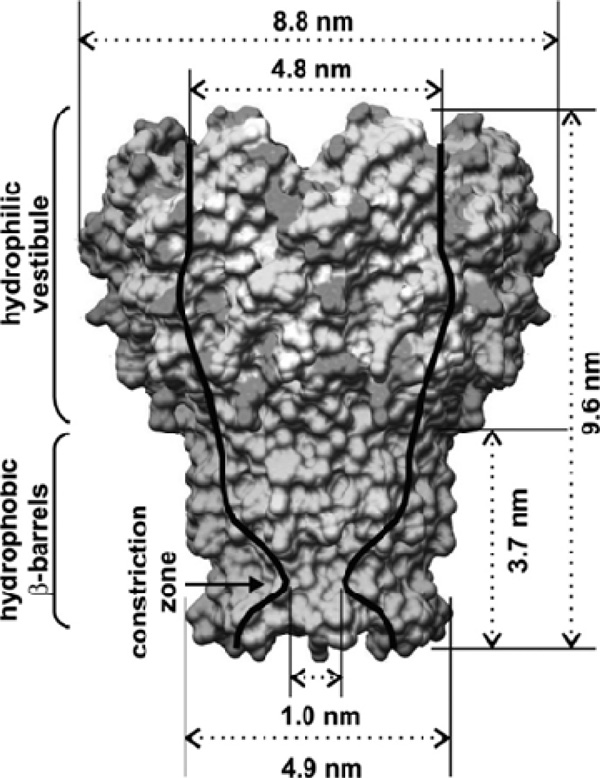
Crystal Structure of MspA of M. smegmatis.Surface representation: light/dark gray: hydrophilic amino acids; mid-gray: hydrophobic amino acids. Note the constriction zone consisting of only hydrophobic amino acids.
With permission from the Journal of Physical Chemistry C, copyright American Chemical Society, 2010.5)
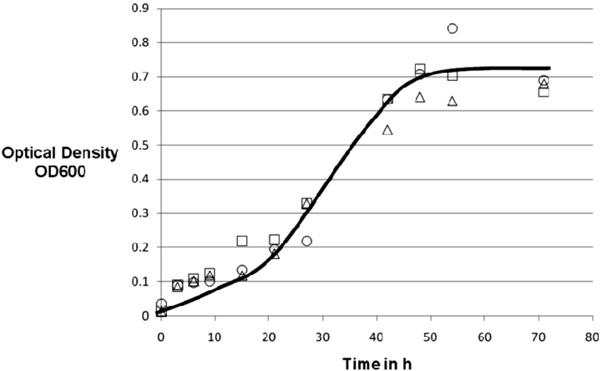
Standard recipe for minimal medium, all cultures grown under the same conditions (37°C, 75 rpm). 20g Glucose, 0.5g sodium nitrate, 1g monobasic sodium phosphate, 1.2g sodium chloride, 0.05g zinc chloride, 15ml liquid minerals, 2ml Tween 80, 2ml copper complex solution, line drawn to guide the eye only.
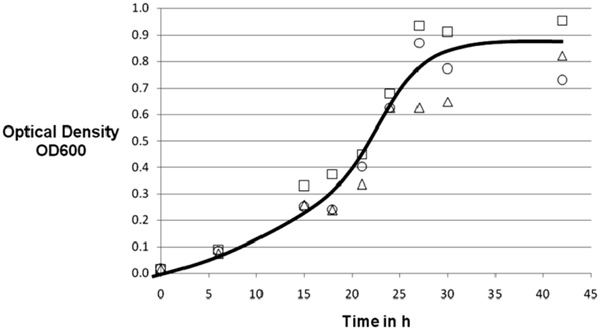
Growth experiment with various iron contents. 20g Glucose, 0.5g sodium nitrate, 1g monobasic sodium phosphate, 1.2g sodium chloride, 0.05g zinc chloride, 15ml liquid minerals, 2ml Tween 80, 2ml copper complex solution. ∆: 20mg FeCl3/l; □: 50mg FeCl3/l; ○: 100mg FeCl3/l, line drawn to guide the eye only.
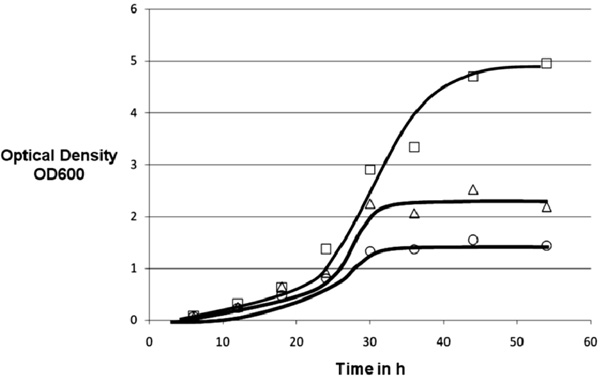
Growth experiment with various amounts of ammonium chloride as secondary nitrogen source. 20g Glucose, 0.5g sodium nitrate, 1g monobasic sodium phosphate, 1.2g sodium chloride, 0.05g zinc chloride, 15ml liquid minerals, 2ml Tween 80, 2ml copper complex solution ○: 2g/l Ammonium Chloride; □: 1g/l Ammonium Chloride; ∆: 0.5g/l Ammonium Chloride, line drawn to guide the eye only.
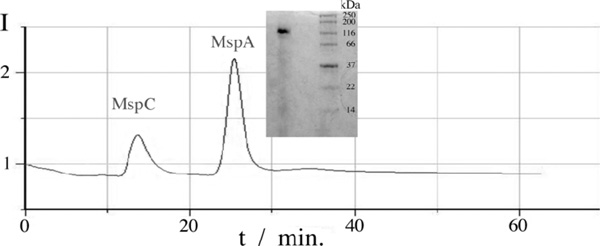
HPLC analysis of an extraction sample from M. smegmatis and gel from MspA analysis with molecular marker.
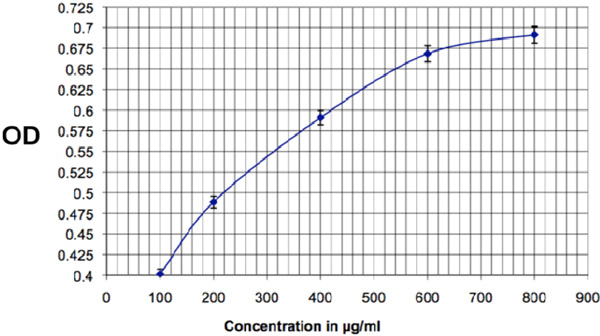
Calibration for the Bradford assay to determine the protein concentration in the extracted product. Optical Density (OD) is measured at 595nm, pathlength: 1 cm.
These surfaces are thought to induce the anchorage of the porin in the lipid bi-layer [1]. Infrared and circular dichroism spectroscopy revealed that heating to 92°C and 112 °C is required to dissociate the MspAoctamer and to unfold the β-sheet domain in the monomer, respectively [6]. The thermal stability of the MspA octamer exceeds even the remarkable stability of the porins of Gram-negative bacteria for every condition tested and is not diminished in the presence of 2 % SDS or within the pH-range from 2 to 11. Due to its superior stability, anisotropy, and the ability to form a hydrophilic homopore, MspA is suited for many and surprisingly different applications. It has been deposited on HOPG-surfaces to form nanopores [5] and complex protein-networks [7], and letter- or star-shaped microstructures when deposited together with PMMA [8].MspA reconstitutes in artificial [9,10]and natural cell membranes [11], as well as polymer-layers [12]. It forms cation-selective ion channels that show voltage gating [12]. The probably most intriguing discovery is that MspA can freely stand with its axis of symmetry perpendicular to a MICA surface without the support of a self assembled monolayer or polymer-layer [13]. A self assembled monolayer is a layer of self-positioning molecules connected to a surface.
Due to its versatility, MspA represents a high-value product (currently $35 per 1µg of purified protein). Our goal is the efficient production of MspA from minimal medium and its efficient purification, so that a market price of less than $10 per 1µg could be reached. This would permit the broad application of MspA as a biological nanotool. One possible applicationmay be the use of MspA as a template for copper nanoparticles in the research for nano-sized digital storage devices [10]. Furthermore MspAmay be applied in the investigation of mycobacterial channel blocking agents,which represent a new strategy in the treatment of tuberculosis and the supply of M. tuberculosis with hydrophilic nutrients [14].The Niederweis group at the University of Alabama at Birmingham has developed a method to selectively extract Mspporins out of M. smegmatis grown in Middlebrook 7H9 medium [15]. This procedure exploits the extreme thermal stability of MspA by heating M. smegmatis cells to 100 °C in the presence of 0.5% of the non-ionic detergent n-octylpolyoxyethylene and yields mainly Mspporins with very little contamination by other proteins [15]. The detergent extracts of the strain M. smegmatis ML10 (ΔMspAΔMspC) do not show any porin band in Coomassie-stained protein gels. A background expression of Mspporins is still detectable in immunoblots using an Msp-specific antiserum [15].It will be demonstrated here that efficient production of MspA can be attained in a relatively simple and inexpensive medium, with the total cost for the fermentation medium estimated at $20 per µg purified protein and with a productivity of 12 µg protein per liter of medium and hour.
MATERIALS & METHODOLOGY
Materials
- SDS (Sodium Dodecyl Sulfate)
- Centrifuge Thermo Scientific Sorvall Legend RT+, Asheville, NC
- Gel electrophoresis equipment: Power Pac Basic, Mini PROTEAN Tetra cell, Bio Rad, Hercules, CA
- Gel electrophoresis chemicals: TEMED (Tetramethylethylenediamine), APS (Ammonium persulfate), Acrylamide / N,N’-Methylenbisacrylamide solution (30:1), Bio Rad, Hercules, CA
- Gel buffer 3x concentrated: 1l distilled water, 3M TRIS (tris-hydroxymethyl-aminomethane), 0.3 w% SDS, pH 8.45 adjusted with NaOH / HCl
- Cathode Buffer 10x concentrated: 1M TRIS, 1M Tricine(N-(2-Hydroxy-1,1-bis(hydroxymethyl)ethyl)glycine), 1w% SDS, pH 8.25 adjusted with NaOH / HCl
- Anode Buffer 10x concentrated: 1M TRIS, pH8.9 adjusted with NaOH / HCl
- Controlled environment incubation shaker, New Brunswick Scientific Co. Inc, New Brunswick, NJ
- 14ml polystyrene round bottom tube Falcon, Becton Dickinson Labware, Franklin Lakes NJ
- KIMAX 2l culture flasks, 125ml culture flasks, KIMAX, Janesville WI
- Phosphate buffered saline (PBS buffer) 10x concentrated: 1l distilled water, 1.36M NaCl, 45mM KCl, 100mM Na2HPO4, 18mM KH2PO4, pH 7.4 adjusted with NaOH / HCl
- PEN buffer 3x concentrated: 750ml distilled water, 300mM Na2HPO4, 0,3mM Na2-EDTA, 450 mMNaCl
- The mc2155 strain of Mycobacterium smegmatis was used in all experiments.
The composition of the different types of media is shown in Table 1.
Different Media and Their Components
| Chemicals | Minimal Medium “First Approach” | Iron Impact Medium | Nitrogen Source Medium |
|---|---|---|---|
| distilled water | 960 ml | 960 ml | 960 ml |
| Tween 80 | 2 ml | 2 ml | 2 ml |
| glucose | 20.0 g | 20.0 g | 20.0 g |
| sodium nitrate | 0.5 g | 0.5 g | 0.5 g |
| Liquid Minerals | 15.0 ml | 15.0 ml | 15.0 ml |
| monobasic sodium phosphate | 1.0 g | 1.0 g | 1.0 g |
| sodium chloride | 1.2 g | 1.2 g | 1.2 g |
| zinc chloride | 0.05 g | 0.05 g | 0.05 g |
| Copper Complex (to replace the antibiotic hygromycin) | 2.0 ml | 2.0 ml | 2.0 ml |
| Ferric chloride | 0 mg | 20, 50, 100 mg | 0 mg |
| Ammonium chloride | 0 g | 0 g | 0.5, 1, 2 g |
7H9 Middlebrook Medium Composition (Fluka Analytical)
| 0.5 g/l Ammonium sulfate |
| 2.5 g/l Disodium phosphate |
| 1 g/l Monopotassium phosphate |
| 0.1 g/l Sodium chloride |
| 0.05 g/l Magnesium sulfate |
| 0.0005 g/l Calcium chloride |
| 0.001 g/l Zinc sulfate |
| 0.001 g/l Copper sulfate |
| 0.04 g/l Ferric ammonium citrate |
| 0.5 g/l L-Glutamic acid |
| 0.001 g/l Pyridoxine |
| 0.0005 g/l Biotin |
Obtained Cell-Mass for the Additional Nitrogen Source Experiment
| Amount of Ammonium Chloride in g/l | 0.5 | 1 | 2 |
|---|---|---|---|
| Cell mass in g | 2.7 | 3 | 2.1 |
The composition of the 7H9 Middlebrook is shown in Table 2. 4ml Glycerol, 2ml Tween 80, 300µl Hygromycin-were added additionally.
All chemicals mentioned are purchased from Sigma Aldrich if not otherwise noted.
Medium Preparation
All chemicals were purchased from Sigma-Aldrich. A 1l Nalgene bottle is filled with 400ml distilled H2O. Potassium phosphate, sodium chloride, sodium nitrate and ferric chloride were added (amounts see Table 2) and the pH is adjusted to 7.2 with hydrochloric acid. A second 1l Nalgene bottle is filled with 560ml H2O and Zinc Chloride and Glucose were added and pH is adjusted to pH 7.2 (HCl). Both bottles were autoclaved and the contents were then combined. 15ml mineral solution, 2ml Tween 80, and 2ml of a 0.01% Copper(II) sulfate-Malachite green solution were added.
Growth Cycle
The Growth cycle is structured in three steps: Maintenance-, medium size and large cultures. To start a new maintenance culture, 4.5ml medium and 0.5ml inoculum were combined in a 14 ml polystyrene round bottom Falcon tube allowing 2-3 days of growth time, using the New Brunswick Scientific PsycroTherm Incubator Shaker at 75 rpm, 37°C. The medium cultures grow in KIMAX 125ml shaking flasks. A sterile flask was under filled with 36ml medium on the clean bench and inoculated with 4ml cell suspension from a maintenance culture without further shaking. The flask was covered with metal lids and placed in an orbital shaker for two days.
The growth of the large cultures was carried out in 2000ml KIMAX shaking flasks. The flasks were filled with 960ml medium and inoculated with a complete medium size culture,bringing the total volume in the flask to 1000ml. The flasks were sealed with aluminum foil to prevent contamination and placed in the shaker for 3-5 days (see medium size cultures for conditions).
MspA Purification Process
A large culture (about 1 l) was centrifuged for one hour at 3,700g using 800 g capacity bottles. The supernatant was decanted while the cells were suspended in 50ml PBS buffer and the resulting suspension was divided in 50ml conical tubes for centrifugation at 10,000g for one hour using a fixed rotor. 10ml PEN buffer was added to the cell pellet to disperse the cells in the 50ml conical tubes after the supernatant was removed. The dispersed cells are collected in a 200 ml Nalgene bottle and kept at 60-65°C in a water bath under agitation (magnetic stir bar, 200rpm, 2cm length). The temperature range is very important for optimum results. 10 µl nOPOE was added and the cell suspension was kept in the water bath for one hour. The suspension was then transferred into a 50 ml conical tube and again centrifuged at 10,000 g for one hour. A 50 ml conical tube with water was used as a counterweight. The supernatant was collected in a 100ml nalgene bottle and 10 ml pre-cooled acetone (-20°C) was added before storing the bottle in a freezer (-20 °C) overnight. After centrifuging the cold suspension for 30 minutes at 10,000g the supernatant was discarded and the cell pellets were dissolved in 10ml PBS buffer before ultrafiltration (Millipore Centrifugal Filter Units MWCO 3000 Dalton, Millipore, Billerica, MA). Centrifuging at 10000g for 30 minutes concentrates the product further.
Product Analysis: Gel Electrophoresis
To prepare an acryl amide separation gel for the gel electrophoresis 3.30ml acryl amide-BIS solution (30:1) was added into a 15 ml conical tube. Also 4.50ml gel buffer, 3.30ml bidest. H2O and 2.25ml glycerol were added and mixed well. Next, 0.015ml TEMED and 0.135ml 10%APS were added and the solution was filled into the glass chamber and the top was covered with a layer of water. For the collection gel, 0.80ml acrylamide-BIS (30:1), 2.60ml gel buffer and 4.20ml bidistilledH2Owere added to a 15ml conical tube. Then 0.015 TEMED and 0.160 10% APS were added and mixed. The covering water from the separation gel was removed and the collection gel solution was added. A comb is placed for pocket formation.
The gel was placed after 30 minutes in the electrophoresis cartridge and the combwas removed. After loading the samples and molecular marker into pockets, the electrophoresis process was started using 125V constant voltage.
Electric power was supplied by BioRadPowerPac Basic and the gel cartridge was a BioRad Mini-Protean Tetra Cell.
HPLC
Protein-containing solutions were analyzed using a Gilson/Hewlett Packard HPLC workstation (Gilson 322 pump, Gilson, Middleton, WI, Hewlett-Packard series 1100 detector, Agilent, Santa Clara, CA) employing a POROS HQ anion exchange column (4.6mm diameter, Applied Biosystems, Carlsbad, CA). The mobile phase consisted of a gradient of two buffers (5mM SDS, composition and 5mM SDS, 1M NaCl, composition). The HPLC procedure has been adapted from reference 14. Flow rate was 0.50 ml/min and the injection volume was 50 µl.
Sterilization
Autoclaving (Yamato Sterilizer SM 200, Santa Clara, CA 95050, USA) 110°C, 30 min holding time, was used for sterilization. 70vol% ethanol in distilled water was used to sterilize equipment that could not be autoclaved.
RESULTS AND DISCUSSION
Growth Experiments
M. smegmatis can be grown efficiently in a simple and well-defined medium as shown in Fig. 2). The effectiveness of the copper complex in suppressing competing microorganisms was demonstrated by the fact that none of the fermentations were contaminated without any great measures to maintain sterility.
The experiment was stopped after 70 hours of total growth-time with a maximum optical density of 0.84. A wet cell-mass result of 2.0g ±0.1g was harvested which equals a yield of 10wt% on the carbohydrates in the medium.
To analyze the impact of iron on the growth of M. smegmatisa second approach with various iron contents was started. Three cultures with an addition of 20, 50 and 100 mg ferric chloride per liter were investigated. The experiment was stopped after 40 hours of total growth-time with a maximum optical density of 0.96. Again a cell-mass result of 2.0g ±0.1g was achieved.
Surprisingly, no conclusive influence of iron in the range tested on the growth of M. smegmatiswas observed (Fig. 3) although iron has been reported to be of some importance in mycobacterial protein synthesis [19]. The absence of a consistent impact of iron concentration in the range tested points perhaps towards a very effective collection mechanism of iron in mycobacteria [20].
The third approach focuses on the nitrogen source. Besides sodium nitrate, we introduced ammonium chloride as a second nitrogen source to further promote growth in our cultures (Fig. 4). The attempt led to extended growth of the M. smegmatis cultures. Maximum growth times, as well as maximum OD600 and cell mass yield were increased:
The cell mass harvested is shown in Table 3, along with the initial ammonium chloride concentration.
Therefore the maximum cell mass yield relative to the carbon source was 15wt%.
The addition of a different nitrogen source shows a major impact on growth and yield of M. smegmatis.The addition of 0.5g NH4Cl led to an increased biomass yield of 35% in comparison to the first approach. These results were exceeded by the addition of 1.0 g NH4Cl. Here the comparison with the first approach results in a 50 % cell mass gain. However, the growth did not reach the level discussed above when 2.0g NH4Cl were added. Apparently a somewhat toxic level was reached, resulting in partial growth inhibition. The harvested cell mass of M. smegmatis was decreased to the results of the basic approach and its growth showed only minor improvements.
The same trend as above has also been observed in the carbon-conversion yield to cell mass. The yield relative to the amount of the consumed carbohydrate substrate was increased from 10wt% in the reference experiments and the iron supplement experiments to approximately 15wt%.
MspA Extraction and Analysis
The extracted product MspA was analyzed in three steps. The first step, gel electrophoresis, shows the purity of the product and verifies the presence of MspA. Our results were identical with earlier findings by Niederweis et al. [6,15]. Fig. (5) shows the MspA band on the left side and the molecular marker on the right. The marker indicates a size of approximately 160kD and the single band present demonstrates the high purity of the extraction product. However the slight haze down the left lane indicates that there were likely some denatured protein fragments in the sample. Fig. (5) also shows the HPLC analysis of our extraction product. Two peaks at retention times of about 14 and 25 minutes are clearly discernible. Both fractions were collected and a gel electrophoresis experiment with the second fraction shows the presence of MspA. Comparison with the literature indicates that the first peak is MspC, a porin quite similar to MspA regarding size and structure [15].
The results of the Bradford assay show a concentration of about 600mg protein per ml (±5.6%), at a yield of 1.4ml per extraction. Fig. (6) shows the calibration standard for the assay.
Assay
The Bradford assay was carried out according to reference 21.See Fig. (6) for the calibration results.
CONCLUSION
In the experiments described above, we show that M. smegmatiscan be cultivated in a medium providing only very basic nutrients and that the cell mass yields can compete with the results obtained from the standard medium (7H9 Middlebrook). This new method results not only in high yields and a pure product (the high-value porinMspA), but also a low risk of contamination. No contaminations were detected during the whole series of experiments described here. As a result from using the less expensive growth medium and the enhanced yield of purified MspA (up by 40% compared to 7H9), we estimate that we were able to lower the price of MspA to approx. $20 per 1µg of purified protein. A next step could be the investigation of the impact of agitation and aeration on the growth of M. smegmatis for scaleup to a multi-liter size bioreactor. Furthermore a possible replacement of glucose as carbon source by an industrial waste product, for example molasses, would be interesting to further reduce costs.
CONFLICT OF INTEREST
The authors confirm that this article content has no conflicts of interest.
ACKNOWLEDGEMENTS
This material is based upon work supported by the National Science Foundation under Award No. EPS-0903806 and matching support from the State of Kansas through Kansas Technology Enterprise Corporation. The authors thank Dr. Michael Niederweis, Department of Microbiology, University of Alabama at Birmingham for generously providing the mc2 155 strain of Mycobacterium smegmatis.


