All published articles of this journal are available on ScienceDirect.
Molecular Typing Of Diarrheagenic Escherichia Coli Strains Isolated From Children Under Five Years Using ERIC-PCR in Koula-Moutou, Gabon
Abstract
Introduction
Diarrheagenic Escherichia coli (DEC) are enteric pathogens responsible for diarrhea in children. The aim of this study was to assess the genetic diversity of DEC in children under five years of age in Koula-Moutou, a topic that has not been previously investigated in Gabon.
Methods
A total of 41 DEC strains, previously isolated and characterized, were included in this study. Genetic diversity was assessed using the Enterobacterial Repetitive Intergenic Consensus-Polymerase Chain Reaction (ERIC-PCR) method.
Results
The ERIC-PCR method enabled typing of 34 out of the 41 DEC isolates. These 34 strains were differentiated into 27 distinct ERIC-PCR genetic profiles, with fingerprints consisting of 1 to 6 bands ranging in size from 250 to 1700 bp. The profiles were grouped into 10 clusters with 90% similarity, each cluster containing between 1 and 7 strains. The DEC isolates from children exhibited diverse genetic profiles, indicating a high degree of genotypic polymorphism.
Discussion
27 different and repeated genetic profiles revealed, suggesting a great diversity of circulating clones, which present different resistance profiles, and which may make the treatment ineffective or inappropriate, reflecting the genetic evolution of DEC in this region of Gabon.
Conclusion
ERIC-PCR is a simple and rapid method for assessing the genetic diversity of DEC. This study revealed significant genetic variation among DEC strains and demonstrated that ERIC-PCR is a valuable tool for epidemiological surveillance and health-related research.
1. INTRODUCTION
Diarrhea is one of the most commonly occurring waterborne diseases in developing countries [1]. It mainly affects children and causes morbidity and mortality in infants [2]. This disease is caused by a wide range of viruses, parasites, and bacteria [3]. Among the bacterial pathogens involved, diarrheagenic Escherichia coli (DEC) is an important causative agent of endemic and epidemic diarrhea worldwide [2, 4]. There are many strains of DEC, categorized into six main intestinal pathotypes according to the generated clinical signs and expressed pathogenicity factors. The various pathotypes described are: Enterotoxigenic Escherichia coli (ETEC), Enteropathogenic Escherichia coli (EPEC), Enteroaggregative Escherichia coli (EAggEC), Enterohemorrhagic Escherichia coli (EHEC), Enteroinvasive Escherichia coli (EIEC), and diffuse adherent Escherichia coli (DAEC).
The prevalence of different enteric pathogens, including DEC, varies according to geographical locations [5]. In addition, several studies have suggested that DECs are responsible for about 30-40% of acute diarrheal episodes in children. These studies have shown that DECs were the most predominant pathotypes, playing an important role in diarrheal morbidity [6-9]. In Africa, and particularly in Nigeria, prevalence rates of 18.4% and 35% were recorded in Abuja and Ile-Ife, respectively [10, 11]. In Gabon, a 2021 study highlighted the phenotyping and molecular typing in the city of Koula-Moutou with DEC detection of 63.5% and 68.5%, respectively [12].
The identification by an accurate and rapid diagnosis can help prevent alarming clinical developments and epidemics [13, 14]. Many methods for strain-typing and origin-tracking of infections exist and are essential to help understand the epidemiology of infectious diseases [15]. Among these, the Enterobacterial Repetitive Intergenic Consensus-Polymerase Chain Reaction (ERIC-PCR) offers a clear and reliable discriminatory power, as demonstrated by numerous studies [15-17]. Among all methods that are used to study the genetic diversity of bacteria in community and hospital settings, ERIC-PCR stands out because of its proven strong discriminatory power, reproducibility, accuracy, specificity, speed, and ease of use, making it a useful tool for routine epidemiological investigations and surveillance in developing countries. Consequently, several studies have used ERIC-PCR as a genotyping method to identify the molecular diversity and genetic relatedness of bacterial strains, particularly E. coli [18]. Similar observations related to the effectiveness of ERIC-PCR were reported, and circulating βLSE genes among E. coli isolates were screened in another study [14]. Likewise, in a study conducted in India, 14 clusters of multi-resistant E. coli strains and βLSE producers were reported by Jena et al. [19]. Equally, Ateba and Mbewe's work on monitoring E. coli O157:H7 infections isolated from several origins revealed remarkable similarities (72% to 91%) between obtained ERIC profiles [20]. Thus, ERIC-PCR gives more sensitive typing results than conventional methods and can be used to discriminate among clones or different members of bacterial species [14]. Although most strains of E. coli are non-pathogenic, those that are considered pathogenic, like DEC, can be the causes of some clinical complications.
This work, which forms part of the study of the genetic relatedness of DEC isolated from a rural city in Gabon, is a pioneering study. Using ERIC-PCR, this study aimed to investigate the genetic diversity of DEC isolates from children under 5 years in Koula-Moutou, a middle-eastern city of Gabon.
2. MATERIALS AND METHODS
2.1. Sample Source and DEC Isolation
The biological material used was supplied and obtained from the collection of strains available at the Laboratory of Molecular and Cellular Biology (LABMC) of the University of Sciences and Technology of Masuku (USTM), Franceville, Gabon. This material was made up of 41 diarrheagenic Escherichia coli (DEC) strains, isolated from childhood diarrheal feces at the Paul Moukambi Regional Hospital Centre in Koula-Moutou, Gabon. The characterization of these DECs has been carried out and described in previous work [21].
2.2. DNA Extraction and Quantification
The extraction of the total DNA was carried out using the heating technique. Young bacterial colonies of 18 h were collected, washed, and homogenized in 300 µL of ultra-pure water. Then, the samples were incubated for 20 min at -20°C and transferred to a dry water bath at 100°C for 10 min. Thereafter, the samples were cooled down for 10 min at -20°C. A volume of 40 µL of protein K at 100 µg/mL was added, and the preparations were incubated at 56°C for 1 h. The resulting lysates were centrifuged at 15,000 rpm for 10 min; the supernatants were transferred to new tubes and kept at -20°C until use.
These supernatants were then quantified using the Invitrogen Qubit™ HS Assay (ThermoFisher, France) with the Qubit Fluorometer. Then, a volume of 1400 µL of the mixture, made of 1393 µL of Buffer Qubit dsDNA HS and 7 µL of Qubit reagents, was freshly prepared and homogenized using a vortex. Previously, 198 and 190 µL of the working solution were collected and allocated into the sample quantification tubes and the standard tubes, respectively. Two microliters (2 µL) of each sample were added to 198 µL of working solution, and 10 µL of standard (S1/S2) to 190 µL of working solution in the appropriate tubes. Then, each sample was shaken for about 2–3 s, and the tubes were incubated for 2 min at room temperature. The sample concentrations were determined on the basis of the calibration curve (S1/S2) and were valid for the volume of 2 µL.
2.3. ERIC-PCR Amplification
Genetic diversity was investigated using the ERIC-PCR method on the 41 DEC strains. The sequences of the different primers used were (5’ATGTAAGCTCCTGGGGA TTCAC3’) and (5’AAGTAAGTGACTGGGGTGAGCG3’) for ERIC-F and -R, respectively.
From total DNA diluted to 1/3, the amplification program was carried out as described in previous studies [22]. The PCR reactions were carried out using a thermocycler (Bio-RAD, T100™, USA) in a final reaction volume of 20 µL containing 2 µL of DNA, 10 µL of master mix at 2X (Golden 360 Taq Amp, ThermoFisher), 2 µL of sense and anti-sense primers (10 pmol), 1.2 µL MgCl2 to 1.5 nM, and 4.8 µL ultra-pure water. The PCR reaction conditions and amplification programs were as follows: initial denaturation at 95°C for 5 min, followed by 30 denaturation cycles at 94°C for 1 min; hybridization at 52°C for 1 min, elongation at 72°C for 2 min, and final elongation at 72°C for 5 min.
2.4. Migration and visualization on Agarose Gel
Amplicons were separated by electrophoresis on a 1.5% (w/v) agarose gel prepared in 0.5X Tris borate EDTA (TBE) solution, containing ethidium bromide (BET). Migration was performed at 200 V and 300 mA for 1 h using the Bio-RAD apparatus (Power Pac™ Basic, USA). In each agarose gel electrophoresis of the amplified products, DNA size markers of 100 bp and 1 kb (Invitrogen, DNA Ladder, ThermoFisher Scientific) were incorporated.
2.5. Data Analysis of Electrophoretic Profiles and Creation of Clusters
The gels were read by the Quantum software (ST4-1100/26MX, France). The manual zero-one method was used to analyze the models using the online data analysis service (http://insilico.ehu.es/dice_upgma/), which uses the Dice index and UPGMA match analysis. It allows comparison of the strains according to their similarities and groups of the most similar individuals, which is to say, with the smallest genetic distance. A dendrogram was then produced according to the obtained clusters.
3. RESULTS
The main objective of this study was to determine the distribution of ERIC sequences within strains of DEC isolated from childhood diarrhea. The analysis of the genetic diversity of the 41 DEC strains by ERIC-PCR revealed a total of 82.9% (34/41) positivity (Fig. 1). Out of the 41 tested isolates, 34 could be successfully typed.
3.1. ERIC-PCR Amplification and Profiling Identification
Fig. 1 presents the analysis of results, revealing DNA fingerprints or profiles of amplified DNA bands ranging from 1 to 7 bands with sizes between 250 and 1700 bp. Common bands were those with sizes of 1450, 1300, 100, 500, and 400 bp, while less frequent bands included those weighing ≥1500, 900, 600, 300, and 250 bp. Seven of the 41 DEC strains did not yield any band, so they could not be genotyped (Fig. 1).
Profile identification showed 27 genotypes with 21 non-repeated and 6 repeated profiles (Table 1). These 27 profiles represented 34 of the 41 DEC strains.
The distribution of profiles according to the pathovars and environment indicate the existence of different genetic profiles (Fig. 2).
| Stains | Diversity of bands | Minimum bands | Maximum bands | Dominant bands (%) | Number of profiles | Unrepeated profiles | Repeated profiles | Number of clusters |
|---|---|---|---|---|---|---|---|---|
| 41 | 13 | 250 | 1700 | 1450 (73.2) | 27 | 21 | 6 | 10 |
| 1000 (39.0) |
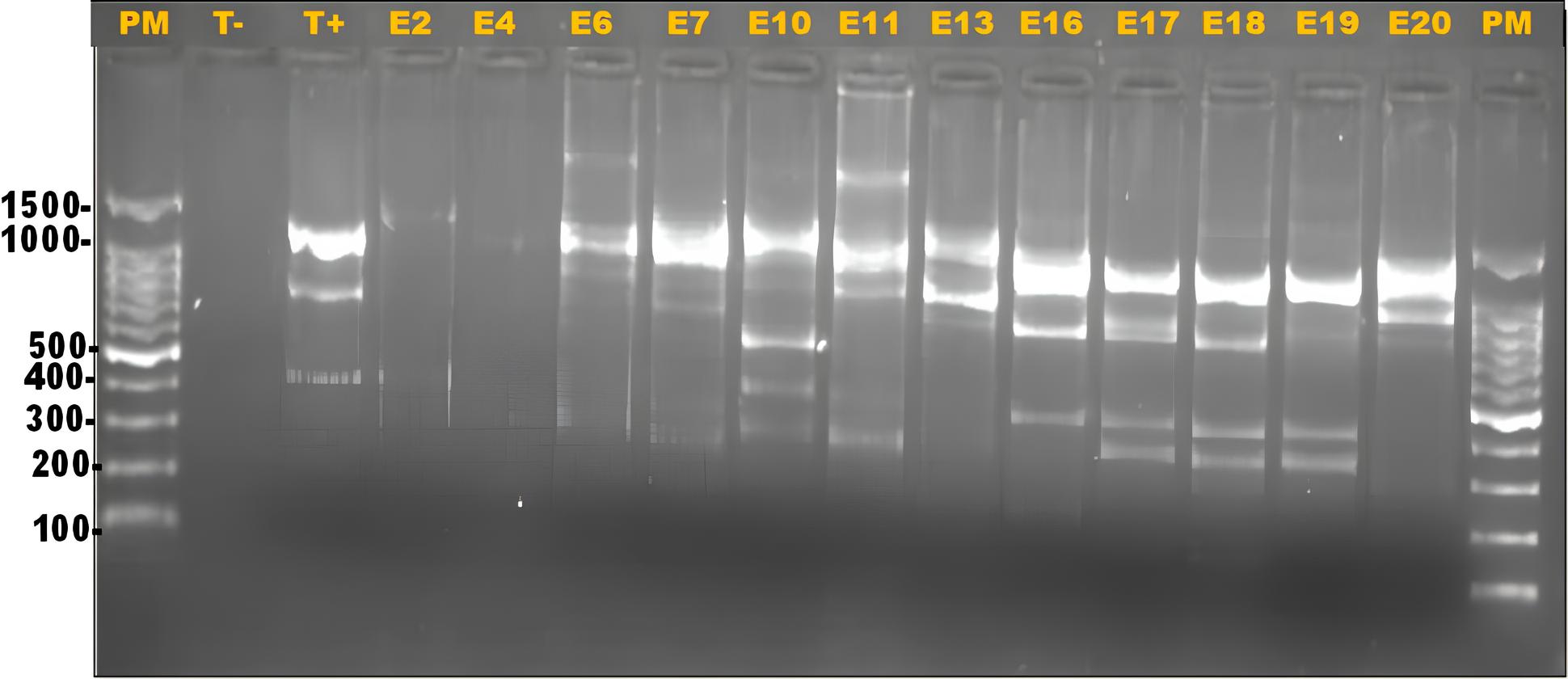
Electrophoretic gel presenting ERIC-PCR profiles of DEC from Koula-Moutou, Gabon. PM: 100 bp DNA ladder; T-: negative control (UltraPure™ DNase/RNase-Free Distilled Water) T+: positive control (E. coli35218); E2-E20: some amplified DEC isolates.
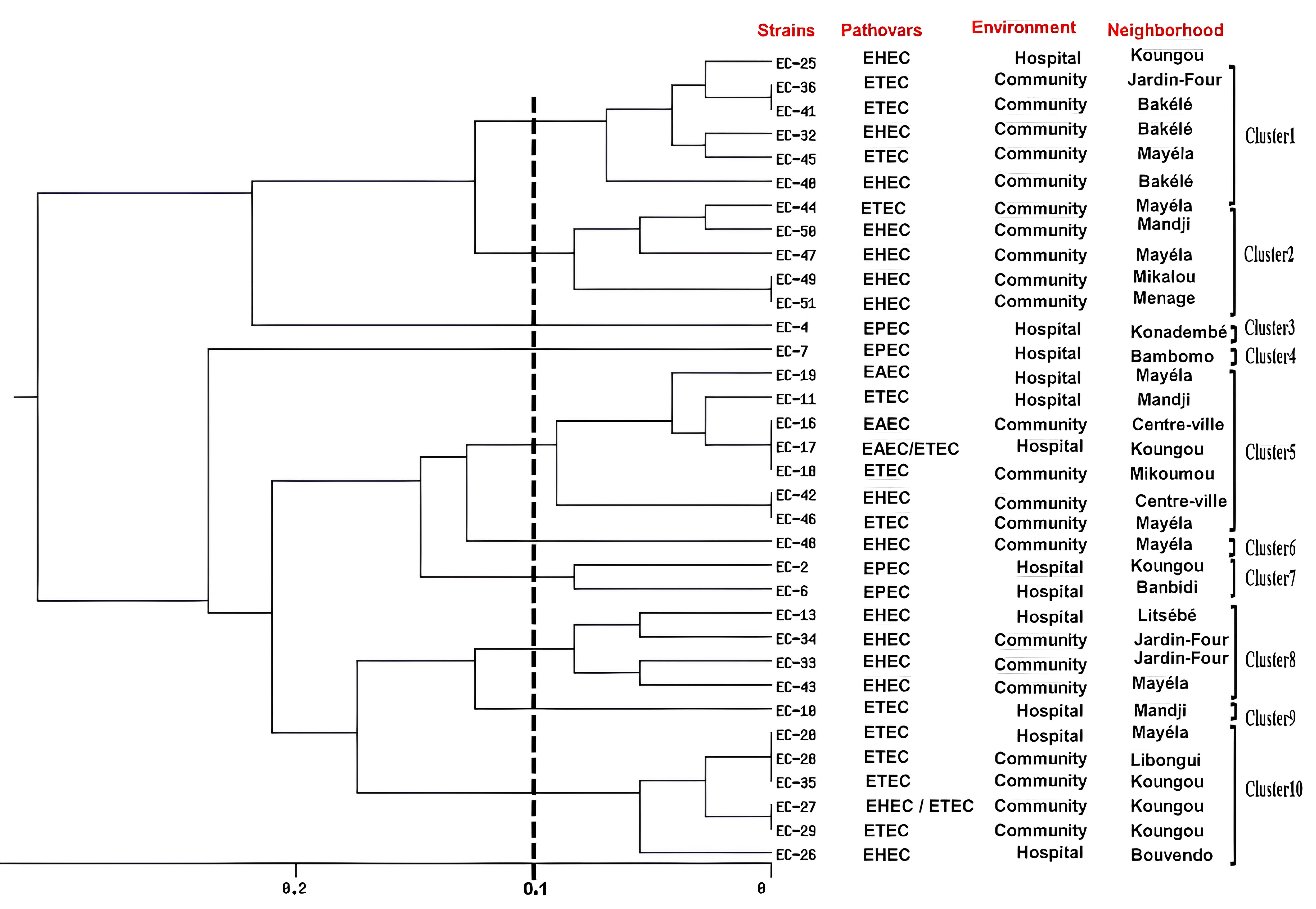
UPMA dendrogram image obtained from cluster analysis showing the relationship of 41 DEC strains isolated from Koula-Moutou, Gabon, using the ERIC-PCR technique.
3.2. Profile and Cluster Analysis
Dendrogram analysis showed that ERIC-PCR differentiated between 34 DEC isolates and categorized them into 10 clusters (C1–C10) (Fig. 2). Results also showed a high degree of similarity among the isolates, with similarity indices ranging from 90% for most isolates to 100% for identical isolates. Depending on their specificity, clusters 1, 2, and 10 had the same type of pathovars (EHEC and ETEC) but in different proportions. Clusters 3, 4, and 7 consisted of the same pathovar type (EPEC); clusters 6 and 8 contained the pathovar EHEC, while cluster 5 contained three different pathovars (EAEC, EHEC, and ETEC).
Of the 34 positive ERIC-PCR strains and the distribution of pathovars by cluster (C), 7 were contained in C5, 6 in C1 and C10, 5 in C2, 4 in C8, 2 in C7 and 1 in C3, C4, C6 and C9, respectively (Fig. 2).
4. DISCUSSION
Several molecular typing studies have been conducted on strains of Escherichia coli, but fewer studies have been done on molecular typing of DEC strains by ERIC-PCR [13, 17, 22, 23]. Using the ERIC-PCR method, the purpose of this study was to determine the genetic diversity of DEC strains isolated in children in Koula-Moutou, Gabon. This genetic prospection of DEC was done on a collection of 41 previously screened and characterized DEC strains [12].
The typing of DEC strains in this study revealed 27 different and repeated genetic profiles, suggesting a wide diversity of circulating clones, which were similar to those reported in the study by Roy et al. [24]. These 27 different genetic profiles can have different resistance profiles and make the treatment ineffective or unsuitable. As a result, patient care can be complicated. In addition, these different clones can be spread through pathways such as low sanitation, unprotected water sources, and non-compliance with good healthcare practices.
Although the band sizes greatly varied from 250 to 1700 bp between strains, the 1450 bp band was present in 30 of the 34 (88.24%) strains. Previous work on the typing of E. coli isolated from avian strains recorded electrophoretic band sizes ranging from 232 to 2690 bp [13]. The difference in sizes would be the result of deletions or acquisitions of sequences at the ERIC sequence level [25].
The high level of DNA profiles of DECs (82.9%) could be explained by the discriminatory power of two ERIC primers (ERIC1R, ERIC2, and ERIC1R+ERIC2). Indeed, it has been previously shown that the ERIC1R and ERIC2 used in our study have a greater ability to discriminate within the E. coli populations [26, 27]. These results are similar to those on 63 enteroaggregative Escherichia coli isolates from Saudi Arabia and revealed 63 unique profiles after typing by ERIC-PCR [23]. Ranjbar et al. [17] identified 115 strains of E. coli from 120 strains isolated from animal feces using ERIC-PCR. Strong discriminatory powers have also been observed with uropathogens [13, 22]. All these results show the existence of considerable variations between different DEC strains, particularly within the same pathovar type, correlated by the presence of one or more elements in the inter-gene region. As a result, the EPEC strains identified had different ERIC profiles, which were grouped in different clusters and originated from different areas of the city of Koula-Moutou. All these data seem to converge on the hypothesis that these strains originally had a common ancestor and then evolved over time by retaining some genetic material. Furthermore, some genetic fragments appear to have been preserved since the divergence of DEC [25]. This conservation of ERIC sequences would undoubtedly be linked to the acquisition of a function certainly related to the stability of the mRNA or a functional role in the bacterial cell, as suggested by other authors [25, 28]. Three DEC pathovars (EHEC, ETEC, EAEC) had similar repeated ERIC profiles, which were grouped into clusters sharing the type of pathovar and the type of patient (hospitalized or community), suggesting the existence of an endemic clone predominant in the city of Koula-Moutou. These results also reflect the genetic evolution of DEC in this region of Gabon. Moreover, the discriminatory power, reproducibility, precision, specificity, speed, and ease of use of ERIC-PCR make it a useful tool for routine epidemiological surveys and surveillance in this country as well as elsewhere.
STUDY LIMITATIONS
Despite the fact that this study showed relevant results on the relatedness of DEC in community and hospital settings, the sample size (41 strains tested) could be a limitation to this study. In addition, although molecular typing methods are commonly used to investigate epidemiological relationships among isolates and sources of infection, it would be advisable, before being used for those purposes, PCR methods for molecular typing require careful in-house validation of typeability, reproducibility, repeatability, stability, discriminatory power and epidemiologic concordance, which were not considered in this study. Also, it would have been interesting to use the MLST, pulsed field gel electrophoresis (PFGE) method or whole-genome sequencing (WGS), which are the gold-standard techniques and seem to be more discriminatory than the ERICPCR used. The uniqueness of these profiles could demonstrate that horizontal transmission and clonal dissemination can occur from clones of different or the same origin and over a greater distance [28].
CONCLUSION
This study showed a genetic difference and relatedness of DEC strains, and the ERIC-PCR method can be a tool for epidemiological health surveillance in Gabon. The genetic diversity revealed through techniques like ERIC-PCR further emphasizes the complexity of challenges in managing DEC infections, as similar clones circulate across different environments. This is the first work reporting on the genetic diversity of DECs in Gabon and revealing different genetic profiles after analyzing the feces of children in the city of Koula-Moutou.
AUTHORS’ CONTRIBUTIONS
The authors confirm their contribution to the paper as follows: J.F.Y., R.M.M., and F.M.: Coordinated the study and drafted the manuscript; R.B.M. and O.Z.M.: Carried out the molecular analyses; J.F.Y., R.M., and A.S:. Revised the article after it was drafted. All authors read and approved the final manuscript.
LIST OF ABBREVIATIONS
| DAEC | = Diffuse adhesion Escherichia coli |
| DEC | = Diarrheagenic Echerichia coli |
| EAggEC | = Enteroaggregative Escherichia coli |
| EHEC | = Enterohemorrhagic Escherichia coli |
| EIEC | = Enteroinvasive Escherichia coli |
| EPEC | = Enteropathogenic Escherichia coli |
| ERIC-PCR | = Enterobacterial Repetitive Intergenic Consensus-Polymerase Chain Reaction |
| ETEC | = Enterotoxinogenic Escherichia coli |
| LABMC | = Laboratory of Molecular and Cellular Biology |
| USTM | = University of Sciences and Technology of Masuku |
ACKNOWLEDGEMENTS
The authors are grateful to the bacteriology immunology team at LABMC, Gabon for providing the bacterial materials necessary for this study and their availability.


