All published articles of this journal are available on ScienceDirect.
Recent Advances in the Ecoepidemiology, Virulence and Diagnosis of Cryptococcus neoformans and Cryptococcus gattii Species Complexes
Abstract
Emerging environmental pathogenic fungal infections, including cryptococcosis, continue to pose a significant threat to humans with compromised immunity and, to some extent, healthy ones. Cryptococcus neoformans was originally identified as the main etiological agent of human cryptococcosis, but recent studies have also identified the occurrence of opportunistic infections caused by Cryptococcus gattii. These two saprophytic facultative yeasts present a paradox as they can infect humans without requiring a host for replication or survival, a phenomenon termed readymade virulence. Many cryptococcal virulence traits appear to have dual effects that provide survival advantages in both animal hosts and the environment. Several molecular techniques have been developed to provide in-depth knowledge of these species complexes. This review will focus on the description of the Cryptococcus neoformans and Cryptococcus gattii (CnCg) species complexes and associated cryptococcal pathogenesis, ecological niches, and virulence factors employed by the pathogens to cause disease.
1. INTRODUCTION
The emergence of pathogenic yeasts continues to threaten humanity, particularly individuals with underlying health conditions [1]. Among these yeast species, the Cryptococcus neoformans and Cryptococcus gattii species (CnCg species) complexes contain the most important and best-characterized pathogens [2]. A study by Park et al. [3] estimated the occurrence of 960,000 new cases of cryptococcal infections predominantly among people infected with HIV. Approximately 60% of people who become infected with these pathogens die due to the lack of effective antifungal treatments [3]. More recent studies report an annual global cryptococcal burden of 223,100 cases, with Sub-Saharan Africa showing the largest burden of the disease, with a reported annual mortality of 181,100 [4].
The pathogenic CnCg species complexes comprise two major species: Cryptococcus neoformans (C. neoformans) and Cryptococcus gattii (C. gattii) [5]. It has been well documented that C. neoformans causes life-threatening meningoencephalitis primarily in immunocompromised patients [4], while C. gattii causes diseases in both immunocompromised and immunocompetent individuals [6]. Cryptococcosis is acquired through the inhalation of sporulated cryptococcal cells from the environment, causing pneumonia in immunosuppressed patients.
In immunocompetent hosts, desiccated yeast cells can be cleared by the immune system or cause asymptomatic latent infection [7]. Traditional virulence factors of CnCg spp. have been well described at the molecular and functional levels and include the ability to synthesize melanin in the presence of a polysaccharide capsule [7, 8]. Clinical detection of CnCg spp. has been achieved using faster and cheaper methods, such as ink staining and antigen kit testing [9, 10]. With regard to the laboratory diagnosis of the pathogen, molecular technologies, such as polymerase chain reaction (PCR), matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), gene chips, and second-generation sequencing technology, present advantages over conventional methods because they can discriminate between C. neoformans and C. gattii [11-13]. Hence, this review will focus on notable discoveries regarding cryptococcal ecology and epidemiology and recent advances in pathogen diagnostics and profiling.
2. CURRENT TAXONOMIC CLASSIFICATION OF THE CNCG SPECIES COMPLEXES
Research on cryptococcosis and the causative agents of the disease dates back to 1894, when pathologist Otto Busse and physician Abraham Buschke isolated Cryptococcus spp. from the tibial bone of a 31-year-old woman diagnosed with chronic granuloma [7]. Studies following this discovery identified a pathogen in peach juice with characteristics similar to that discovered by Busse and termed it Saccharomyces neoformans [14]. Jean-Paul Vuillemin subsequently renamed this yeast Cryptococcus neoformans in 1901 because it did not produce ascospores, which is a distinctive feature of Saccharomyces [15]. Research on cryptococcosis intensified in the 1980s when the disease became a major health threat due to the onset of the AIDS pandemic, and cryptococcal meningitis became one of the AIDS-defining illnesses among individuals with depleted immune systems [16].
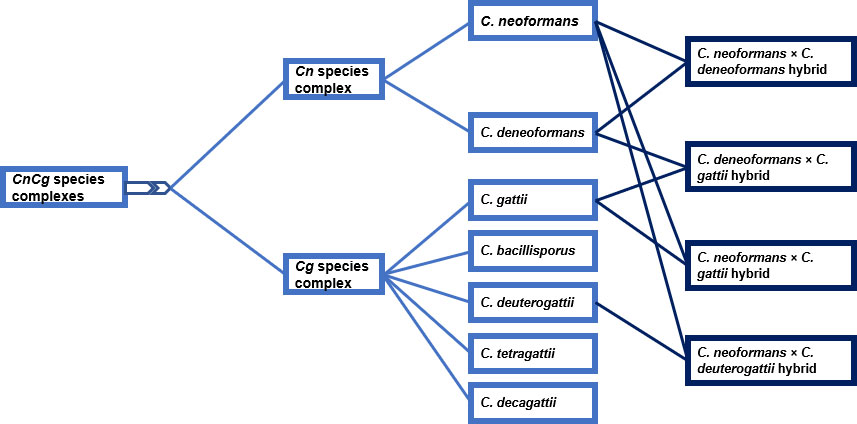
Previous studies have divided the CnCg species complexes into Serotype A and Serotype D based on biochemical tests or capsular epitopes [17] (Fig. 1). Serotype A comprises C. neoformans, which is further divided into C. neoformans var. grubii and C. neoformans var. neoformans. The molecular genotypes of the latter include the VNI, VNII, and VNB strains, and the former includes VNIV strains. The endemic VNB strain was first discovered in Botswana by Litvintseva et al. [18], who associated the strain with native African trees, especially the Colophospermum mopane (Co. mopane) tree that occurs abundantly in Botswana. Phylogenetic analysis shows that the VNB and VNI strains are closely related and share a common phylogenetic history, which is consistent with parsimonious conclusions that the VNI population originates in Africa [18]. Hybrids of the two varieties are serotype AD strains, constituting the VNIII strains [19]. C. gattii was recognized as a distinct group and classified as serotypes B and C based on basidiospore morphology, morphological features of the yeast cells, limited molecular identity (55 to 61% relatedness of DNA), and inefficient cross-species mating, which led to the production of sterile hybrids with no recombination. The group was further subdivided into the VGI, VGII, VGIII, and VGIV strains [20].
In recent molecular phylogenetic studies, researchers have identified significant diversity and genetic divergence within the CnCg species complexes. In the new taxonomic classification, the former C. neoformans varieties have been reclassified into C. neoformans and C. deneoformans. The reclassification was further extended to C. gattii (Serotype B/C), where the following names have been allocated to its strains: C. gattii (VGI), C. bacillisporus (VGIII), C. deuterogattii (VGII), C. tetragattii (VGIV) and C. decagattii (VGIV/VGIIIc). In addition, the hybrids that formed as a result of interspecies mating were classified as C. neoformans x C. deneoformans, C. deneoformans x C. gattii, C. neoformans x C. gattii, and C. neoformans x C. deuterogattii (Fig. 1).
3. HYBRIDIZATION IN THE CNCG SPECIES COMPLEXES AND ITS ROLE IN SPECIATION
While hybridization was once thought to be rare or nonexistent in nature, there is evidence that hybridization occurs pervasively in some fungal phyla and some oomycetes. Through the process of hybridization, genes and alleles that have diverged significantly from each other over time are brought together in individual cells, creating novel interactions among genes and genomes [23]. The hybrids that form may be haploid or dikaryotic in many fungi or diploid in oomycetes. Normally, the genetic integrity of the species is maintained by natural reproductive barriers and by processes preventing the introduction of genetic material through mating with nonconspecific individuals. However, when reproductive barriers become permissive, mating will occur between individuals or different species, leading to the formation of hybrids [24]. Mating in CnCg spp. is well-defined and is known to be controlled by genetic information encoded in the mating type genes within the MAT locus. This region is unique such that homologous chromosomes contain nonhomologous information that confers genetic differences between cell types [25].
In nature, CnCg spp. exist as two mating types, designated α (MATα) and a (MATa), where the former is the most dominant in both environmental and clinical isolates [9]. Both C. neoformans and C. gattii show two forms of sexual reproduction: (i) bisexual reproduction, which takes place between two cells possessing compatible mating types (i.e., MATα and MATa), and (2) unisexual reproduction, also known as haploid fruiting, which involves cells with the same mating type (i.e., MATα cells) [26]. Sequencing of the MAT locus revealed that it is unique, spans > 100 kb in size, and comprises more than 20 genes. The gene products encoded by the MAT locus include divergent components of a MAP kinase cascade, regulators of sexual reproduction, pheromone and pheromone receptors, and other proteins that play no role in mating [27]. The mating type locus in CnCg spp. significantly differs from other known fungal mating type loci, but it shares features with self-incompatibility systems and sex chromosomes of plant algae and animals [27]. For the mating process to occur, cells secrete either a- or α-pheromones that bind to G-protein-coupled receptors on cells of the opposite mating type.
Mating is initiated when the mating pheromone binds to its cognate receptor, eventually activating the heterotrimeric G protein and subsequently the MAP kinase cascade. Haploid yeast cells of the opposite mating type (MATa and MATα) will then encounter each other, leading to the dissociation of the α-subunit of the trimeric G-protein. Eventually, βγ heterodimers recruit the scaffolds Ste5 and Far1 to the plasma membrane by binding to their RING-H2 domains [28]. Then, Far1 binds various effector proteins, including Bem1 and Cdc24, a Guanine nucleotide exchange factor (GEF) for the Rho-type GTPase Cdc42, which collectively mediate polarized growth toward the compatible mating partner [29]. Similarly, activated Cdc42 and Ste5 recruit Ste20 and Ste11, the MEK Ste7 and the MAPK Fus3 [30]. Once activated, Fus3 phosphorylates multiple substrates regulating various cellular activities, such as the induction of a transcriptional program, cell cycle arrest in G1, oriented polarized growth, and cell-cell and nuclear fusion, which are critical in the complex mating process [28].
The most frequently encountered CnCg hybrid strains found in both clinical and environmental settings thus far are AD hybrids, probably because serotypes A and D occupy similar environmental niches and cause most cases of cryptococcosis [31]. Upon germination, these hybrids produce diploid or aneuploid products, suggesting that the sexual cycle is not efficient in crosses between C. neoformans var. neoformans and C. neoformans var. deneoformans. Previous laboratory studies have reported the presence of αADα hybrids possessing two MATα locus alleles originating from serotype A and serotype D parental strains [31, 32]. This finding provided evidence that the same-sex mating cycle also occurs in nature. The presence of αAD and ADα hybrids could arise via mating between cells of either the opposite or the same mating type, followed by the loss of one of the MAT loci [31].
Previous reports provide evidence of a close relationship between mating type, sexual reproduction, and virulence in CnCg spp [32, 33]. Sexual reproduction favors the virulence of yeast via the production of infectious spores, as α isolates can be more virulent than congenic a-isolates [32]. Genes located outside of the MAT locus interact with genes within the α locus to direct pathogenesis. Spores and desiccated CnCg cells are very small, making it easier for them to fit into sites deep within alveoli, thus enhancing the virulence of the species complexes. The mating types also differ in their interactions with the mammalian host; for instance, C. neoformans var. grubii α isolates disseminate through the central nervous system more easily than congenic a-type isolates [33]. Furthermore, hybridization provides an additional selective advantage to CnCg spp., as hyphal forms that are induced during mating enable nutrient foraging over an increased area [33].
4. EPIDEMIOLOGY AND ECOLOGICAL NICHE OF CRYPTOCOCCOSIS ETIOLOGICAL AGENTS
The epidemiological profile of cryptococcosis has not been well established because the routine differentiation of C. gattii infections from those caused by C. neoformans is not standard practice in certain parts of the world. Previous studies on CnCg species report that C. gattii is geographically restricted to tropical and subtropical regions around the world [22, 34]. However, increased surveillance has now identified the emergence of C. gattii in the northern part of the United States, Canada, and northern Europe, indicating that this pathogen may occupy an ecological niche that has not been previously recognized [7]. This hypothesis is supported by reported outbreaks of this saprophytic yeast in Canada [35] and the United States [36].
In nature, the CnCg species complexes exist in diverse ecological niches, and the most preferred habitats of C. neoformans are the droppings of birds (e.g., pigeons) and unicellular organisms (e.g., amoebae and sowbugs) [7]. The ability of C. neoformans to elude soil amoebae has played a critical role in the evolution of yeast as a facultative intracellular pathogen. It has been discovered that C. neoformans can utilize survival strategies acquired through interaction with amoebae to evade human macrophages. The predatory instincts of amoebae have helped the CnCg species complexes to acquire virulence factors that enhance pathogenicity in humans [37]. Rare cases of infection have been reported in pigeons because of their elevated body temperatures of 41 - 42 °C, which exceed the optimum growth temperature of C. neoformans; however, the isolation of this species from bird beaks, crops, or feet indicates that feed ingested by the birds could be contaminated with saprophytic yeast [38]. The linkage between the CnCg species complexes and natural habitats has not been well defined despite the global consistency in reports of the relationship between C. neoformans and avian hosts [7].
Unlike its sister pathogen, C. gattii has been isolated from Eucalyptus camaldulensis and E. tereticornis, and these trees represent its preferred ecological niche [39-41]. However, it has been revealed that Eucalyptus is not the only environmental habitat of C. gattii, as the pathogen has been isolated from other tree species and from geographic regions that do not include Eucalyptus [42]. A previous study by Jo-Marie Vreulink and colleagues isolated members of the CnCg species complexes from trees in two recreational areas in South Africa [43]. Litvintseva et al. [18] described mopane trees as a novel ecological niche of C. neoformans in Botswana and other Southern African countries where this tree is endemic. Furthermore, they discovered endemic strains of VNB and VNI that are found only in Africa and described them as highly genetically diverse and potentially virulent strains that may cause major disease outbreaks. Their findings indicate a public health concern because communities in areas highly populated with mopane trees benefit culturally and economically from this tree species. For instance, some parts of the tree, such as mopane bark, wood, leaves, and seeds, can serve as traditional medicines [44, 45]; debarked mopane poles are used for the construction of mud huts [44]; and more importantly, mopane trees are the sole substrate of mopane worms, which are a component of the diet and are considered a culinary delicacy in Angola, Botswana, Namibia, South Africa, and Zimbabwe [46]. These areas report the highest burden of HIV/AIDS, and continuous exposure to and human interactions with mopane trees may therefore represent a tremendous health hazard in these areas. Further investigations are needed to reveal the interactions among humans, mopane trees, and the CnCg species complexes (Fig. 2).
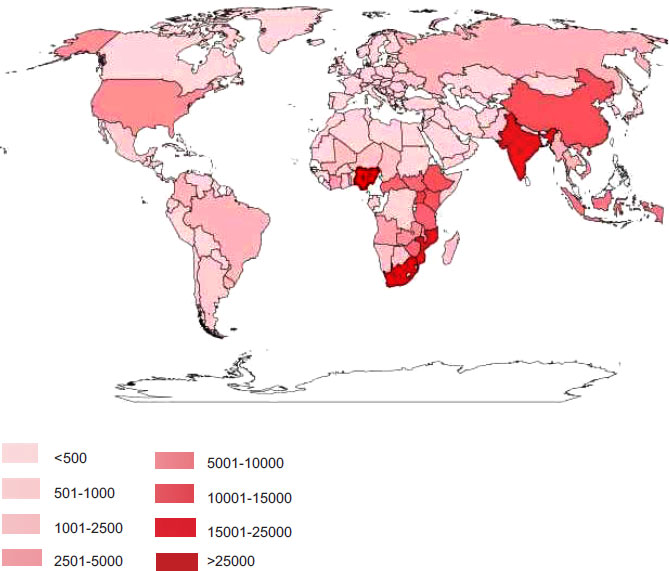
The global annual incidence of cryptococcal meningitis was estimated to be 223,100 [4]. The largest burden of the disease is experienced by the sub-Saharan region, with Europe and America reporting the lowest numbers of infections.
5. CRYPTOCOCCAL PATHOGENESIS
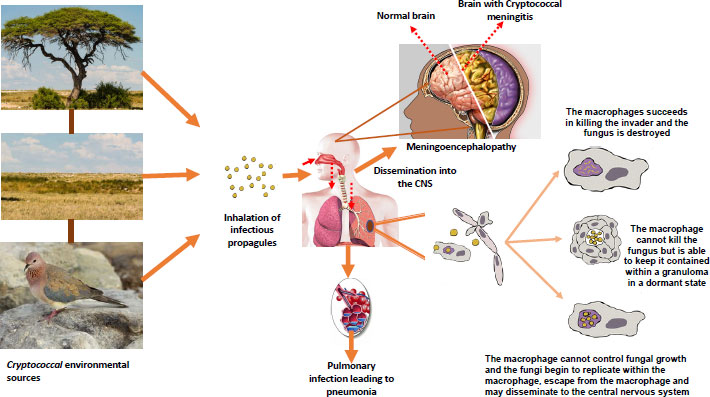
The cryptococcal infection starts with the inhalation of either small yeast cells or desiccated basidiospores [47] generated through heterosexual or unisexual reproduction [48] (Fig. 3). Host defense mechanisms, such as airflow turbulence, mucociliary transport, and mucus trapping, do not stop these small infectious propagules from penetrating the distal part of the lungs [49]. At this initial stage of infection, the pathogen battles against macrophages in the lungs by employing antiphagocytic factors, such as capsules and melanin pigment [50]. Upon entering the lungs, the pathogen can cause pneumonia in immunosuppressed patients, but in immunocompetent hosts, yeast cells are either cleared by alveolar macrophages or may establish an asymptomatic latent infection [7].
Shibuya et al. [51] revealed that macrophages cannot clear these yeasts on their own; instead, they require an adaptive immune response that leads to a self-resolving granuloma. These findings are supported by lesions observed in athymic mice when the mice are experimentally infected with C. neoformans. The response of mural macrophages to infection was confirmed even though they were not aggregated, and cryptococcal cells of both intra- and extra-cytoplasmic yeasts were observed [51]. The conclusions derived from the work of Shibuya et al. [51] were that necrosis occurred as a result of an ischemic state produced by the enlargement of the granuloma and that cryptococcal proliferation did not cause necrosis. On the other hand, cryptococcal infections in patients with compromised immunity can enter the respiratory tract and initially cause pneumonia or eventually disseminate into the brain via the bloodstream, where it will manifest as meningoencephalitis [52]. Meningoencephalitis occurs when blood‐borne CnCg species, either in the blood or contained within phagocytic cells, are arrested in the brain vasculature and make contact with brain endothelial cells before transmigrating into the brain parenchyma [53].
6. GENOMIC DESCRIPTION OF C. NEOFORMANS AND C. GATTII AND ASSORTED VIRULENCE FACTORS
The availability of genomic sequencing platforms for the CnCg strains and the recent technological improvements in molecular assays have greatly advanced our understanding of these pathogenic yeasts. Researchers utilized multi-locus sequence typing (MLST) to reveal the genetic diversity of C. neoformans in Africa and how phenotypic heterogeneity impacts the virulence of the pathogen in immunocompromised individuals [54]. Their findings reported high levels of genetic diversity and locus variability in South African genotypes relative to globally distributed types. The authors discovered that there is a correlation between fungal genotypes and phenotypic characteristics (phagocytosis, laccase activity, and cerebrospinal fluid (CSF) survival) with regard to virulence [54]. Moreover, when they compared MLST genotyping data with clinical parameters, Beale and other authors identified a relationship between genetic lineages and clinical outcomes, with indications of the worst survival in patients infected with the VNB lineage relative to other lineages. A similar study by Day et al. [55] utilized MLST coupled with genomic sequencing to investigate why a C. neoformans var. grubii molecular genotype they named VNIγ was causing infections among healthy individuals in Vietnam. Their study identified a consortium of proteins (phenolic acid decarboxylase, DEAD-box ATP-dependent RNA helicase 26, oxoprolinases, a taurine catabolism dioxygenase, a zinc finger protein, membrane transport proteins, and various drug transporters) that enhanced the pathogenicity of C. neoformans var. grubii in infected healthy people [55].
With the cost of sequencing becoming reasonably affordable, researchers have expanded the knowledge of the genomic diversity of CnCg species complexes. Most recent sequence studies have characterized CnCg species into traditional lineages of VNI, VNII, and VNB and supported the splitting of VNB into two lineages, VNBI and VNBII, a pattern that has been noted in previous AFLP and MLST studies [42, 56, 57]. Further genotypic studies revealed that the VNI and VNII molecular types, which represent C. neoformans var. grubii, are closely related to the VNIV molecular type, which represents C. neoformans var. neoformans. Furthermore, the clades that have represented molecular types VGI, VGIII, and VGIV in this work were more closely related to one another than to the sibling VGII clade [58]. Molecular characterization of the two strains of VNB showed that environmental VNBI isolates were more resistant to oxidative stress and more heavily melanized than clinical VNBI isolates. Sequencing studies associated mutations in virulence factors and stress response genes with clinical isolates rather than environmental isolates [56]. These findings imply that genomic sequencing can be used to directly test and identify a host of potential virulence factors and mutations in the laboratory setting.
A study by Lee et al. [59] analyzed the function of C. neoformans phosphatase, kinase, and its transcriptional factors (TFs) network to get a better understanding of its pathobiological signaling. They constructed a high-quality library of 219 signature-tagged gene deletion mutant strains representing 109 out of 139 putative phosphatases in C. neoformans. Their in vivo expression studies revealed certain phosphatases (PPU1, DPP101, and AKP1) to be markedly upregulated during host infection [59]. This study discovered 31 pathogenicity-related phosphatases that affect a series of biological processes, such as growth, stress response, virulence factor production, carbohydrate metabolism, protein sorting, cell signaling, and vesicular trafficking [59]. Some of their findings, including the role of PTP2, CAC1, CNA1, CCR4, and HAD1 in the virulence of C. neoformans, have been corroborated by other previous researches [60, 61].
Another study performed by Vogan et al. [62] revealed genomic regions that significantly contribute to the differences in virulence-associated traits between C. neoformans and C. deneoformans. Using Quantitative Trait Locus (QTL) mapping, which is regarded as a powerful tool used in the identification of genomic regions associated with phenotypic differences between strains, researchers identified 23 QTL associated with differences in the expression of virulence factors (melanin production, cell size, cell wall thickness, and capsule production) and susceptibility to the antifungal drug fluconazole. Among these identified QTL, six QTL were involved in resistance to fluconazole. Interestingly, all of the six traits identified possessed QTL on chromosome A, which is the largest chromosome, containing approximately 13% of the genes in the cryptococcal genome [62].
7. CRYPTOCOCCAL VIRULENCE FACTORS
Traditional determinants of virulence factors produced by the CnCg species complexes have been extensively studied in C. neoformans than in C. gattii, in part, because C. neoformans’ H99 strain is popularly used as a reference strain to study the pathogenic Cryptococcus species [5, 7, 37, 63]. Nevertheless, several studies have shown that traits responsible for virulence in these two cryptococcal species are similar [64-66]. These traits, which include capsular polysaccharide production, melanin pigment secretion, cellular signaling pathways, thermal tolerance, strategies for parasitizing host phagocytes, and mechanisms for breaching the blood–brain barrier, are all crucial for the virulence of the CnCg species complexes [63]. Notably, cell aging plays a pivotal role in the pathogenesis of C. neoformans. It has been shown that cells that get older as a result of undergoing many replicative cycles have been shown to accumulate during chronic cryptococcal infections and manifest distinct phenotypic traits associated with virulence [67]. For example, older yeast cells, also known as founder cells, manifest a distinct phenotype, such as an enlarged cell size, thicker cell wall, resistance to fungicides, or resistance to hydrogen peroxide-mediated killing [67]. It has been hypothesized that increased resistance to antifungals and phagocytosis is caused by changes in cell wall structure [68]; consequentially, this results in the accumulation of more founder cells in the brain than young cells [69].
Secondly, nutrient deprivation in cells induces several virulence-associated determinants, such as increasing the gene expression and activity of the laccase enzyme (Lac1), which propels virulence by initiating pigment formation [70]. Melanin is a naturally occurring negatively charged hydrophobic pigment of high molecular weight formed by the oxidative polymerization of phenolic and/or indolic compounds. This pigment is required for the pathogenicity of many fungal pathogens; however, the mechanisms that play a compressive regulatory role remain unidentified. In more recent research that aimed at analyzing melanin regulatory signaling pathways in C. neoformans, Lee et al. [59] identified four melanin-regulating core transcription factors (Bzp4, Hob1, Usv101, and Mbs1) and two kinases (Gsk3 and Kic1) responsible for the regulation of melanin synthesis and other steps that are very important in melanization processes. These melanin-regulating core transcription factors (MRC-TFs) play a critical role in the regulation of LAC1 in a distinctive way and act in mediating interconnections with cAMP, regulation of Ace2 and morphogenesis (RAM), and HOG signaling pathways [70]. Furthermore, they demonstrated Hob1 and Mbs1 to have pleiotropic roles that affect hundreds of genes, whereas Bzp4 and Usv101 demonstrated a defined range of action in terms of the number of genes that they regulate [59]. The findings laid out by Lee et al. [59] present new insights into how C. neoformans species regulate melanization, and subsequently, the role played by melanin in the virulence of these pathogenic yeasts.
Similar to other microbial pathogens, the CnCg species complexes produce several degrading enzymes, such as proteases and lipases, which can confer virulence in their hosts [71]. For C. neoformans, urease has been profiled as a degradative enzyme that plays an important role during infection. This yeast produces large quantities of urease that catalyze the degradation of urea into CO2 and ammonia, and the enzyme is required for brain invasion during infection [72]. Ammonia produced may be required for the adhesion of C. neoformans either by promoting the expression of adhesins on the endothelial layer or by inducing a cytotoxic effect on the integrity of tight junctions of the blood-brain barrier, thereby facilitating the invasion of the brain [73].
Further, experiments have shown that superoxide dismutase (SOD1p) protects the functional integrity of several virulence factors in C. gattii. This prominent antioxidant has been shown to be essential for infection in an animal model and protect yeast from phagocytosis by human macrophages [74]. An equally important virulence determinant enzyme is mitochondrial superoxide dismutase (SOD2p), which allows the growth of both C. neoformans and C. gattii at 37 °C [75]. Another important antioxidant and stress protectant in fungi is trehalose. A study revealed that homologous proteins encoding the key biosynthetic enzyme trehalose-6-phosphate (T6P) synthase (TPS1p and TPS2p) regulate the thermotolerance, virulence, and integrity of other virulence determinants, such as the capsule and melanin [58].
8. PATHOGEN DIAGNOSIS AND CHARACTERIZATION
The clinical detection of cryptococcosis is performed by isolating the CnCg species from a clinical specimen (e.g., CSF, sputum, or skin biopsies) or by direct detection of the fungus by means of India ink staining of body fluids. The latter is the most rapid diagnostic method and involves the visualization of cryptococcal yeast by direct microscopy after staining CSF with India ink [76]. In culture-dependent methods, positive samples are characterized by the presence of opaque or white-to-cream colonies that may eventually change color to orange-tan or brown after prolonged incubation [77]. Other rapid clinical laboratory methods include serology, cytology, and histopathology. Molecular methods are not currently applied in routine clinical practice, although they are extensively used for research purposes [9].
8.1. Current Trends in Rapid Diagnostics and Profiling of CnCg Species Complexes
Recent advances in fungal genomics and high-throughput sequencing techniques have provided rapid and highly discriminatory protocols to circumvent the disadvantages of conventional methods. Most of these techniques are nucleic acid or protein-based, including whole genome sequencing (WGS), MLST, and PCR strategies, such as digital droplet PCR, Illumina MiSeq, ion torrent, and nanopore technologies, and terminal restriction fragment length polymorphism (T-RFLP) analysis [78-80]. Other assays, such as MALDI-TOF MS, are based on proteomic profiles.
8.1.1. MALDI-TOF MS
MALDI-TOF MS has been developed for the rapid and accurate identification of most pathogenic bacteria and fungal isolates in clinical settings. Several studies have used this platform to correctly identify CnCg species complexes members and their genotypes based on characteristic protein spectra obtained from whole cells, allowing the highly discriminatory identification of pure cultures rapidly and at a minimal cost depending on the sample preparatory method (Table 1) [22, 81-88]. The efficacy of the platform was tested by Hagen et al. [22], who confirmed the positivity of 423 well-characterized CnCg isolates.
The platform produces results within a relatively shorter period of time, uses fewer reagents, and has few experimental procedures, when compared to conventional identification methods [89]. MALDI-TOF MS has thus been shown to be an alternative technology for CnCg species identification that can replace time-consuming conventional methods. The adoption of this technique in the clinical setting will significantly contribute as a useful rapid tool for routine and accurate identification of CnCg species, which will eventually lead to decreased hospitalization times and reduced mortality [11].
| Source | Pathogen Identified | Identification at Species Or Genus Level (%) | Molecular Genotypes Identified | Identification by other Methods (%) | References | ||
|---|---|---|---|---|---|---|---|
| C. neoformans/C. gattii species complexes | Clinical, veterinary, environmental | 20/20 (100%) | VNI, VNII, VNIV, VGI, VGII, VGIII, VGIV, AD hybrids | URA 5 RFLP (100%) | [86] | ||
| C. neoformans/C. gattii species complexes | Clinical | 136/136 (100%) | NI | ITS/IGS sequencing (100%) | [81] | ||
| C. neoformans/C. deneoformans species | Clinical |
Cn=18/22 (81.8%) Cd= (33.3%) |
NI | AFLP1 (49%) AFLP3 (41%) AFLP2 (10%) |
[82] | ||
| C. neoformans/C. gattii species | Clinical | 423/423 (100%) | VNI-VNIV, C. gattii, C. bacillisporus, C. decagattii, C. deuterogattii | AFLP | [24] | ||
| C. neoformans/C. gattii species | Clinical | 494/498 (99%) | NI | Microscan, AFLP | [89] | ||
| C. neoformans/C. gattii species | Clinical | 129/129 (100%) | VNI-aA, VNI-aA, VNII-aA, VNIV-aD, VNIII-aADa | AFLP | [84] | ||
| C. neoformans/C. gattii species | Clinical, veterinary | 70/70 (100%) | CnCg | ITS sequencing | [88] | ||
| Cryptococcus species complexes | Veterinary | 62/62 (100%) | NI | ITS sequencing | [90] | ||
| C. gattii species complex | Clinical | 254/254 (100%) | VGI, VGII, VGIII, VGIV | MLST (100%) | [91, 92] | ||
| C. neoformans/C. gattii species complex | Clinical | 32/32 (100%) | VNI, VNII, VNIV, VGI, VGII, VGIII | URA 5 RFLP (100%) | [93] | ||
8.1.2. Multilocus Sequence Typing (MLST)
MLST has been adopted as the first-choice typing approach for epidemiological investigations of CnCg species by the International Society for Human and Animal Mycology (ISHAM) [90]. The platform was originally developed for typing bacteria and has proven to be highly discriminatory for several human pathogenic fungi, including Candida glabrata, Candida tropicalis, Candida albicans, Coccidioides spp. and Histoplasma capsulatum [91]. Using seven unlinked genetic loci (CAP59, GPD1, LAC1, PLB 1, SOD 1, URA5 and IGS1), the technique was employed to identify major cryptococcal isolates corresponding to VNI, VNII, and VNB, a Botswana-specific genotype closely related to VNI [91]. Other previous studies have successfully used MLST to reveal subpopulations within C. neoformans var. grubii and C. gattii [93, 94].
Chen et al. [95] showed that the VNB molecular type occurs widely in arboreal environments, and it has been specifically associated with the mopane tree belt across Botswana. In the years since, the ISHAM consensus MLST scheme was used to reveal a highly clonal population structure with less variability among Brazilian populations relative to populations from other continents [96]. A study by Vanhove et al. [42] recovered high concentrations of C. neoformans VNB from mopane trees across Zambia, further confirming the Zambezi Mopane Woodland ecoregion as the main niche for this genotype [42]. To further improve the MLST approach to make it highly discriminatory with highly reproducible results, Cogliati et al. [97] designed multilocus sequence typing primers to enable the genotyping of AD hybrids within the C. neoformans species complex. This step makes the evaluation and identification of emerging highly virulent genotypes possible. Moreover, the study provides a basis for studying the molecular epidemiology of AD hybrids, hence revealing important new insights about the species complexes.
8.1.3. Genome Sequencing
Whole-genome sequencing typing (WGST) has been used to study the epidemiology of different organisms, including CnCg species. This platform can capture all types of genetic variation, including single-gene copy number variations, nucleotide polymorphisms, insertions and deletions, and genome structural rearrangements. Currently, microbial epidemiological studies using whole-genome sequencing rely on short read sequences from the Illumina MiSeq or HiSeq platform for robust identification of SNPs for the inference of epidemiological patterns and strain relationships [54, 98]. Whole-genome sequencing has been used to recapitulate the three traditional serotype A molecular genotypes (VNI, VNII, and VNB) [98]. In the Pacific Northwest, WGS has been used to understand the isolates of VGII that caused an outbreak of cryptococcal infections in the region. Additionally, VGIII isolates were deeply sequenced to characterize an increasing number of infections in the USA, South and Central America.
Another recent population genomic study identified a highly clonal cluster of VNI isolates capable of infecting immunocompetent individuals in East Asia [55]. Preliminary studies by Ramachandran et al. [99] suggested the potential application of metagenomic next-generation sequencing (mNGS) for the detection of CnCg species in the central nervous system, even at very low amounts undetectable by traditional microbiologic testing. The suggestion was made following the detection of Mycobacterium tuberculosis and C. neoformans in the CSF of 3 patients with symptomatic cryptococcal antigenemia and symptoms of meningitis. Recent findings by Xing et al. [100] revealed mNGS to be more effective than conventional methods in the diagnosis of infectious encephalitis and/or meningitis, especially concerning the identification of species. However, they acknowledge that this novel approach has shortcomings and should be used in conjunction with conventional microbiological testing, and that interpretation of data should differ depending on the pathogen involved [100].
8.1.4. Surface-enhanced Raman Scattering (SERS)
Raman microspectroscopy is a very powerful tool that can be used to analyze biological materials. This technique is rapid and non-invasive with a high spatial resolution in the acquisition of biochemical and structural data by generating point spectra or spectral images [101]. A recent study employed a surface-enhanced Raman scattering platform for the detection and discrimination of C. neoformans and C. gattii. In contrast to other molecular techniques used to detect C. neoformans and C. gattii, SERS exhibits high specificity with a relatively low cost and fast analysis times [13].
The application of SERS in fungal identification offers several advantages that can make up for shortcomings of other methods of fungal detection. For example, spectral fingerprints offered by SERS are unique to the constituent molecules. Furthermore, SERS provides qualitative and quantitative detection of the analyte because of high specificity [102]. The spectral bands of SERS are narrow with high resolution of spectral peaks, thus minimizing interferences on the fluorescence background [103]. The characteristic spectra allow the synchronized detection of various components. The application of SERS can be utilized for biological testing as it is relatively not expensive and portable [104].
A major challenge encountered when using SERS biosensors is the quantitative detection of pathogens in the sample. This phenomenon originates from the nature of the SERS platform, in which enhancement of Raman signals highly depends on the nanoscale roughness of the metal surface [105]. However, researchers have focused on acquiring highly controlled SERS signals by combining cucurbit [n] urils with Au colloids to immediately bridge adjacent NPs, hence creating uniform gap distances [106]; by use of atomically smoothened nanowires (NW) with Å roughness [105, 107] as an active substrate for SERS; by using silicon substrates coated with Au; and finally, by using an Ag nanorod array as a SERS active substrate [108].
CONCLUSION AND FUTURE STUDIES
Cryptococcosis continues to cause high morbidity and mortality among individuals with underlying ailments because of failure to detect the causative agents sufficiently early. First, it is imperative to fully understand the ecological niche of the species complexes in its natural environment and within human hosts, its adaptations, and its associations with other microbiome residents. In this regard, ecological studies will lead to an in-depth understanding of the pathogen’s epidemiology. There should be an increased focus on epidemiology, ecological modeling, and host–pathogen interactions to acquire a better understanding of this enigmatic yeast and ultimately identify better measures for its control. Moreover, technologies, such as NGS, SERS, and gene editing, should be employed to effectively study microorganisms and the biome with which they coexist collectively as a holistic approach that may reveal previously unknown intricate associations of CnCg species and other organisms.
LIST OF ABBREVIATIONS
| T-RFLP | = Terminal restriction fragment length polymorphism |
| ISHAM | = International Society for Human and Animal Mycology |
| WGST | = Whole-genome sequencing typing |
| SERS | = Surface-enhanced Raman scattering |
| NW | = Nanowires |
AUTHORS’ CONTRIBUTION
Kenosi Kebabonye and Mosimanegape Jongman contributed to conceptualization; Mosimanegape Jongman, Ishmael Kasvosve, and Daniel Loeto supervised the study; Mosimanegape Jongman contributed to validation; Kenosi Kebabonye and Mosimanegape Jongman wrote the original draft; Kenosi Kebabonye, Mosimanegape Jongman, Ishmael Kasvosve, and Daniel Loeto contributed to writing, review and editing.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
This study was funded by University of Botswana, funder ID: 501100022089 ( Grant number: UBR/RES/ROUND OF 39).
CONFLICT OF INTEREST
The authors declare that they have no conflict of interest concerning this article.
AVAILABILITY OF DATA AND MATERIALS
The data supporting the findings of the study is available with in the article. Fig. (3) was generated with authors using photographic images from unpublished works.
ACKNOWLEDGEMENTS
Declared none.


