All published articles of this journal are available on ScienceDirect.
Characterization and Antibiotic Susceptibility Profiles of Pathogenic Escherichia Coli Isolated from Diarrhea Samples within the Buffalo City Metropolitan Municipality, Eastern Cape, South Africa
Abstract
Objectives:
Developing countries like South Africa are still faced with numerous challenges such as poor environmental sanitation, lack of clean drinking water and inadequate hygiene which have contributed largely to diarrheal infections and deaths in children. This study was aimed at investigating the prevalence of pathotypes, antimicrobial resistance and drug resistance determinants among Escherichia coli (E. coli) isolates from diarrhea stool samples within Buffalo City Municipality, Eastern Cape, South Africa.
Methods:
Fresh diarrheal stool samples were collected from 140 patients attending public health centres within the Municipality and presumptive E. coli isolates were obtained from the stool samples using E. coli chromogenic agar while PCR amplification methods were used to confirm the presumptive isolates as well as delineate them into pathotypes based on the presence of certain virulence genes. In addition, antimicrobial susceptibility and screening of some of the antimicrobial resistant determinants were performed on all the confirmed isolates.
Results:
A total of 394 presumptive E. coli isolates from 140 diarrhea stool samples were subjected to polymerase chain reaction amplification, of which 265 were confirmed positive as E. coli. Pathotypes delineation of the positive E. coli isolates validated the presence of ETEC 106 (40%), EAEC 48 (18%), DAEC 37 (14%), and EPEC 31 (11%) while no EIEC pathotype was detected. All E. coli isolates exhibited maximum susceptibility to gentamicin (95%), amikacin (91%), nitrofurantoin (91%), meropenem (90%), chloramphenicol (91%) norfloxacin (84%) and imipenem (83%). However, the isolates showed multidrug resistance to penicillin G, ampicillin, trimethoprim, tetracycline, doxycycline, and erythromycin, with over 71% of the isolates resistant to the drugs. The prevalence and distribution of the five resistance determinants assessed were as follow; sulphonamides; sulII (12%), beta lactams; [ampC (22%); blaTEM, (25%)], and tetracyclines (tetA (35%).
Conclusion:
The results from this study suggest the probable involvement of E. coli pathotypes as an etiologic agent of diarrhea in the study area and revealed high levels of multidrug resistance among the isolates, which could be a major health burden.
1. INTRODUCTION
According to the World Health Organization, diarrhea is characterized as the bowel movement accompanied by three or more loose stools per day or more than it is usually for a person[1]. In 2016, diarrheal disease resulted in an estimated 688 million illnesses and 1.6 million deaths worldwide among children under the age of 5 years [2]. More than a quarter of these deaths occurred in South Asia and sub-Saharan Africa [3]. The fundamental cause of death due to intense diarrhea is dehydration, which comes about with losing electrolytes and water in the stool [1]. Another substantial reason for death is the looseness of the bowels and poor nutrition [1]. Though diarrheal infections are preventable, nonetheless, they are the second foremost cause of death in developing countries [4, 5]. Generally, the transmission of diarrhea initiated by Escherichia coli is through the fecal-oral route, including the eating and/or drinking of contaminated food or water, individual to individual interaction with contaminated excrement due to poor personal hygiene and relaxed environmental sanitation [6].
Most studies have reported that diarrhea is initiated by a wide range of etiologic agents which are bacteria (Campylobacter, Shigella, E. coli, Salmonella spp., Vibrio cholerae,), enteric viruses (Adenovirus, Norwalk virus, Rotavirus, etc.) and parasites (Giardia spp., Cryptosporidium spp. and Entamoeba histolytica [7]. Over the years, E. coli has been reported to be one of the primary communal bacteria that have been isolated and it is the main cause of childhood looseness of bowels [8, 9]. E. coli is found in the gut, but some strains of the organism have virulence factors that trigger them to initiate infections in the gut of both healthy individuals with strong immune system as well as in immuno-compromised patients where these pathogenic strains cause damages in the integrity of the intestinal mucosal once ingested [9, 10]. E. coli pathogenic strains are subdivided into two classes: DEC and the extra-intestinal pathogenic E. coli (ExPEC) that includes UPEC (related to UTI), SEPEC (related to sepsis), and NEMEC (related to neonatal meningitis) [10, 11]. There are six fundamentally recognized strains of diarrheagenic E. coli (DEC) that are related to diarrhea based on their different clinical features, virulence factors and serotypes grouping and they are: entero-aggregative E. coli (EAEC), enteroinvasive E. coli (EIEC), enterohemorrhagic E. coli (EHEC), entero-pathogenic E. coli (EPEC), diffusely adherent E. coli (DAEC), and enterotoxigenic E. coli (ETEC) [12]. These pathotypes are categorised based on their virulence factors and the mechanisms of pathogenicity in the human gut like in diarrhea cases [12]. E. coli possess many virulence factors which could be classified as primary and secondary virulence factors. The primary virulence factors are Shiga toxins (Stx1, Stx2), Heat labile toxins (LTI, LTII), Type III secretion system(s) (Sep), cytotoxic necrotizing factors (CNF1, CNF2), haemolysins (Hly, Ehx/Ely), intimin (Eae) while the secondary virulence factors are Adherence factors/Fimbriae (P-, S-, F1Cfimbriae, Bfp, AAF), Colonization factors (CFA/CS), heat stable toxins (STa, STb, EAST), invasion factors (Ipa), iron transport systems (aerobactin/Iuc) and capsule [13, 14]. Antimicrobial resistance hampers the effective treatment of diseases caused by pathogenic microorganisms. New antimicrobial resistance mechanisms are emerging and spreading globally, thus impeding humans’ ability to treat common infectious diseases, resulting in prolonged illness, disability, and death [15]. It increases the cost of health care as it lengthens patients' stay in hospitals, thus leading to higher global disease burden. The indiscriminate and non prudent use of antibiotics is the major reason for the rapid development of bacterial resistance to antimicrobials [15]. Carbapenems, cephalosporine, monobac- tams, and penicillins which are members of the beta-lactam antibiotics, fluoroquinolones, sulfanamides, streptogramins like quinupristin and dalfopristin, tetracyclines, glycopeptides and lipoglycopeptides such as vancomycin, macrolides such as erythromycin and azithromycin, oazolidines (linezolid and tedizolid), polypeptides and rifamycins are commonly avai- lable antibiotics for treating bacterial infections in humans. Ceftolozane/tazobactam, ceftazidime, and cefiderocol, which are all cephalosporin, Carbapenem (imipenem, meropenem) Quinolone (delafloacin), Omadacycline which is a synthetic tetracycline derivative and lefamulin, a pleuromutilin anti- biotic, are the newest antibiotic approved by FDA in recent years. Other older antibiotics such as the penicillin and its derivatives, chloramphenicol, tetracycline, colistin, clindamy- cin, daptomycin, fosfomycin, nitrofurantoin, and tigecyclne are equally used in the management of bacterial infectious diseases.
The use of antibiotics in the management of acute gastroenteritis in children is quite challenging. This is because the pattern of pathogens causing diarrhea does not only vary from one region to another but also according to age, season, immunization status and symptoms [16]. Also complicating treatment is that diarrhea infections could be caused by multiple pathogens which are usually the norm among children with the ailment. The prevalence of antibiotic resistance in any given locality is also a huge factor in considering the choice of empiric antibiotic for treatment in order to minimize the rates of treatment failure and further resistance development. Generally, the selection of antibiotic is usually based on two major considerations: the chance of obtaining microbiological results timeously, including resistance pattern, and the severity of clinical conditions [16]. The WHO recommends treating all episodes of blood in the stools with antibiotics and to use ciprofloxacin as the first-line drug while other alternatives are azithromycin, ceftriaxone, metronidazole, co-trimoxazole, rifaximin, and vancomycin [17]. Children or adults who have diarrhea can be prescribed with oral rehydration treatment and also antibiotics are suggested to patients who have blood in their feces [18]. Among diarrheagenic E. coli isolates, resistance to the prescribed drugs is likely to occur due to intrinsic factors as well as possession of resistance factors mediated by genetic determinants such as efflux pump, enzyme modification of the drug, alteration on the target size and metabolic pathway and poor stewardship in the use of these drugs in the community could exacerbate the problem. Therefore, this study was intended to characterize the pathotypes, assess the antibiotic susceptibility patterns and profile some resistance determinants of E. coli isolates from diarrheagenic stools of patients visiting public medical facilities within the Buffalo City Metropolitan Municipality in Eastern Cape of South Africa.
2. MATERIALS AND METHODS
2.1. Ethical Approval and Consent to Participate
Before the study commenced, the ethical clearance was granted by the University of Fort Hare research ethics board (Protocol number IWE001), while permission to collect samples was sought from the department of health. Verbal informed consent was granted by study participants and sampling was a prospective once off collection with the aid of nurses in the health facilities used as study sites.
2.2. Description of Study Area and Sample Collection
The Buffalo City Metropolitan Municipality lies on the eastern side of the Eastern Cape Province with a geographical co-ordinates of 32˚59’S 27˚52’E. Local clinics and other health-care centers in Zwelitsha and Mdantsane were used as sampling locations. Sample collection was a once-off prospective sampling from diarrheal patients with the help of nurses of the health centres within the study site. Fresh diarrheal stool was collected into sterile screw stool sample containers and transported in ice boxes to the Applied and Environmental Microbiology Research Group (AEMREG) laboratory at the University of Fort Hare, Alice and were processed immediately upon arrival. Stool samples were only collected from patients who had symptoms of diarrhea for at least two consecutive days and no other criteria apart from age and having diarrhea were considered among the participants.
2.3. Bacterial Analysis
A total of 140 fecal samples were obtained from diarrheal patients recruited for the study. Firstly, the diarrheal stool samples were directly streaked onto brilliance chromogenic agar plates (Merck, Modderfontein, Gauteng, South Africa) and incubated for 24 h at 37 °C. Two to three distinctly isolated steel blue colonies were selected as positive presumptive isolates for E. coli which was aseptically purified and sub-cultured into sterile Muller Hinton broth (Merck, Modderfontein, Gauteng, South Africa) for 30% glycerol stocks preparation and then stored at -80˚C freezer for additional analyses.
2.4. DNA Extraction, Molecular Confirmation and Pathotyping of Confirmed Bacterial Isolates by PCR Amplification.
For DNA extraction of the positive presumptive isolates of E. coli, the boiling method as described by Iweriebor et al. [19], was adapted. A loop full of the glycerol stock was resuscitated in a 10 mL Tryptone Soy Broth (TSB), incubated overnight and thereafter, 2mL of the broth culture was used for extraction. The 2 mL culture broth was transferred into sterile micro-centrifuge tubes and centrifuged at 10,000rpm for 10 min. The supernatant was discarded and 200 μL water of PCR grade was added to the pellet and re-suspended by vortexing. Thereafter, the bacterial suspension was boiled in a heating block to lyse the cells (Thermo Fisher Scientific, Leicestershire, UK) at 100 ºC for 10 min. The boiled cell lysate was then left to cool and then centrifuged for 5 min at 12,000 rpm. Thereafter, the supernatants were transferred into a new sterile 1.5 mL micro-centrifuge tube. The obtained supernatant containing the total crude DNA was used as a template for the confirmation of the presumptive isolates based on uidA house-keeping gene for E. coli that codes for beta-glucuronidase enzyme and delineation of the pathotypes of E.coli which are EPEC, EHEC, EAEC, ETEC, DAEC, and EIEC in a conventional PCR amplification. E. coli ATCC 25922 was used as positive quality control in the confirmation of the isolates.
The presumptive isolates were confirmed by PCR using E. coli uidA specific primers as described by Scot et al. [20] while the screening for E. coli pathotypes was performed according to the methods adopted by Titilawo et al. [21] as presented in Table 1. The annealing temperatures for the respective PCRs are shown in Table 1 and the reaction was run in a final volume of 25 µL. The amplicons were verified on 2% agarose gel electrophoresis which was stained with 5 μL of ethidium bromide and resolved at 100 V for 1 hour in a 0.5X TBE buffer and viewed with Alliance 4.7, UVITEC, UK transilluminator and 100bp DNA ladder was used as molecular weight marker.
2.5. Antimicrobial Susceptibility Testing (AST) of the E. coli Isolates
The antimicrobial susceptibility testing of the confirmed isolates was carried out by Kirby-Bauer disc diffusion method on Mueller Hinton Agar (MHA) following the standard guidelines of Clinical and Laboratory Standards Institute [28]. Briefly, the confirmed E. coli glycerol stocks were resuscitated in nutrient broth, after that they were sub-cultured onto nutrient agar plates and incubated for 18 hours at 37˚C. The colonies were picked and inoculated onto sterile normal saline (0.9% NaCl) to make up a suspension equivalent to 0.5 McFarland standards. The suspension was inoculated directly onto the Mueller-Hinton agar plates using a sterile swab, allowed to dry. Then, antibiotics discs were dispensed on the agar plates and incubated for 24h at 37 °C. Eighteen commercially obtained antibiotics impregnated discs were tested against the confirmed isolates and they include: ampicillin (10µg), amikacin (30 µg), cephalothin (30 µg), gentamicin (10 µg), nalidixic acid (30 µg), nitrofurantoin (300 units), meropenem (10 µg), imipenem (10 µg), doxycycline (30 µg), tetracycline (30 µg), chloramphenicol (30 µg), norfloxacin (10 µg), erythromycin (10 µg), trimethoprim (5 µg), polymyxin B (300 units), penicillin G (10 µg) cefotaxime (30µg), ceftazidime (30µg) and thereafter, the zones of inhibition were measured and were interpreted according to the guidelines of CLSI (2018). E. coli ATCC 25922 was used as negative control for the AMR testing as it is sensitive to all antibiotics.
2.6. Multiple Antibiotic-Resistance Index (MARI) Interpretation
The MARI for each sample location was similarly performed using the Krumperman, expression which is given as: MARI = a / b, where ‘a’ is the sum of antibiotics which the isolate showed resistance to and ‘b’ as total sum of antibiotics against which an individual isolate was tested [29].
2.7. Screening for Antibiotic Resistance Determinants Among Resistant Isolates
The screening of the antimicrobial resistance determinants was done by PCR. Sulphonamide resistance gene (sulI), genes encoding β-lactamases (ampC and blaTEM), tetracycline resistance gene (tetA), and aminoglycoside resistance gene (aadA) were the only selected target genes profiled. The list of the primers sequences used is shown in Table 2 as previously described by Titilawo et al. [21]. The reaction mixture (25 µL) contained 1 µL of 10 pmol specific primer pairs, 12.5 µL of PCR master mix (New England Biolabs-NEB), 5.5 µL of nuclease free water and 5.0 µL of DNA template. E. coli ATCC 25922 was used as positive quality control while nuclease free water served as the negative control in all the PCR reactions. The cycling conditions were as follows as previously described by Maynard et al. [30] and Iweriebor et al. [19]. The amplicons were verified on 2% agarose gel electrophoresis which was stained with 5 μL of ethidium bromide and resolved at 100 V for 1 hour in a 0.5X TBE buffer and viewed in an Alliance 4.7, UVITEC, UK transilluminator and 100bp DNA ladder was used as molecular weight marker.
3. RESULTS
3.1. Confirmation of Presumptive E. coli Isolates by PCR
Of the 324 presumptive isolates, 265 were confirmed as E. coli based on the uidA gene by PCR. Based on the age categorization, 100 patients were below the age of 5 years, 25 were within 5-10 years, while 15 were above 10 years of age.
3.2. Pathotype Characterization by PCR
The confirmed E. coli isolates were further delineated into pathotypes targeting the existence of specific genes (Table 1). The Enterotoxigenic E. coli was the most prevalent strain with 40% positive for the lt gene, followed by Enteroaggregative E. coli with 18% where 71% (34/48) had Eagg gene and 33% (16/48) positive for AafII gene; Diffusely adherent E. coli (14%) for the daaE gene and Enteropathogenic E. coli (11%) for eae gene. Enteroinvasive E. coli and Enterohaemorrhagic E. coli were not detected among the isolates, as shown in Table 3. The occurrence of different pathotypes of DAE was more in the age group less than 5 years followed by those between 5 to 10 years and those above 10 years of age as can be seen in Table 4.
| Target strains | Target genes | Primer sequence (5->3) | Amplicon size | References |
|---|---|---|---|---|
| E. coli | uidA | F-GACCTCGGTTAGTTCACAGA R-CACACGCTGACGCTGACCA |
147 | [22] |
| EPEC | Eae | F-TCAATGCAGTTCCGTTATCAGTT R-GTAAAGTC GTTACCCCAACCTG |
482 | [23] |
| ETEC | Lt | F-GGCGACAGATTATACCGTGC R-GCGGTCTCTATATTCCCTGTT |
450 | [24] |
| EAEC | Eagg | F-AGACTCTGGCGAAAGACTGTATC R-ATGGCTGTCTGTAATAGATGAGAAC |
194 | [25] |
| EIEC | ipaH | F-CTCGGCACGTTTTAATAGTCTGG R-GTGGAGAGCTGAAGTTTCTCTGC |
933 | [26] |
| DAEC | daaE | F-GAACGTTGGTTAATGTGGGGTAA R-TATTCACCGGTCGGTTATCAGT |
542 | [26] |
| EHEC | stx1 stx2 |
F-CAG TTAATGTGGTGG CGAAGG R-CACCAGACAATGTAACCGCTG F-ATCCTATTCCCGGGAGTTTACG R -GCG TCATCGTATACACAGGAGC |
384 584 |
[27] [27] |
| Antibiotics class | Primer | Primer sequence (5ʹ–3ʹ) | Amplicon size (bp) | Reference |
|---|---|---|---|---|
| Sulphonamides | sul1 | F:TTCGGCATTCTGAATCTCAC R:ATGATCTAACCCTCGGTCTC |
625 | [30] |
| Beta-lactams |
ampC blaTEM |
F:TTCTATCAAACTGGCACC R:CCTTTTATGTACCCAGA F:TTTCGTGTCGCCCTTATTCC R:CCGGCTCCAGATTTATCAGC |
202 690 |
[31] [32] |
| Tetracycline | tetA | F:GCTACATCCTGCTTGCCTTC R:CATAGATCGCCGTGAAGAGG |
201 | [33] |
| E.coli Pathotypes | ETEC | EAEC | DAEC | EPEC | EIEC | EHEC |
|---|---|---|---|---|---|---|
| Targeted gene(s) | Lt | Eagg AafII | daaE | Eae | ipaH |
stx1 stx2 |
| % of positive isolates | 40% | 18% | 14% | 11% | 0 | 0 |
| E.coli pathotype | Age groups and prevalence of pathotype per age | ||
|---|---|---|---|
| <5 years | >5-10 years | >10 years | |
| ETEC (102) | 80 | 20 | 2 |
| EAEC (48) | 32 | 10 | 6 |
| DAEC (37) | 20 | 14 | 3 |
| EPEC (13) | 6 | 5 | 2 |
3.3. Antibiotic Susceptibility Profile of the Confirmed E. coli Isolates
All the 265 confirmed isolates were subjected to testing of susceptibility against a panel of eighteen empiric antibiotics. Results were interpreted according to CLSI [28] standards. All intermediates resistances were taken as resistant. The results showed that resistance against penicillin G was 99% while 88% were resistant to ampicillin, trimethoprim (84%), tetracycline (83%), doxycycline (82%), and erythromycin (71%). High susceptibility of the isolates was observed for gentamicin (95%), amikacin (91%), nitrofurantoin (91%), meropenem (90%), chloramphenicol (91%) norfloxacin (84%) and imipenem (83%) as shown in Table 5.
3.4. Multiple Antibiotic Resistance Index (MARI)
The MARI was calculated and it revealed that all the isolates had MARI values that exceeded the threshold of 0.2. The average MARI index was 0.8 which is four times more than the threshold limit as shown in Table 6. This suggests that the E. coli isolates recovered from this study may have originated from high risk-sources of contaminated drinking water or food.
| Antimicrobial agents | Code | Concentration (µg) | Isolates n=265 | ||
|---|---|---|---|---|---|
| Susceptible | Intermediate | Resistant | |||
| Ampicillin | AP | 10 | 23 (9%) | 10 (3%) | 232(88%) |
| Amikacin | AK | 30 | 251 (95%) | 5(2%) | 9 (3%) |
| Cephalothin | KF | 30 | 110 (41%) | 79 (30%) | 76 (29%) |
| Gentamicin | G | 10 | 252 (95%) | 9(3%) | 4 (2%) |
| Norfloxacin | NOR | 10 | 224 (85%) | 7 (3%) | 34 (13%) |
| Doxycycline | DXT | 30 | 28 (11%) | 20 (7%) | 217(82%) |
| Nalidixic Acid | NA | 30 | 153 (58%) | 52 (20%) | 60 (22%) |
| Erythromycin | E | 15 | 3 (1%) | 73 (28%) | 189 (71%) |
| Trimethoprim | TM | 5 | 56 (21%) | 2 (1%) | 207 (78%) |
| Nitrofurantoin | NI | 300 | 243 (92%) | 12(5%) | 10 (3%) |
| Tetracycline | T | 30 | 37 (14%) | 8 (3%) | 220 (83%) |
| Polymyxin B | PB | 300 | 218 (82%) | 0 (0%) | 47 (18%) |
| Penicillin G | PG | 10 | 2 (1%) | 0 (0%) | 263 (99%) |
| Imipenem | IMI | 10 | 220 (83%) | 43 (16%) | 2 (1%) |
| Meropenem | MEM | 10 | 238 (90%) | 27 (10%) | 0 (0%) |
| Cefotaxime | CTX | 30 | 190 (72%) | 13 (5%) | 62 (23%) |
| Ceftazidime | CAZ | 30 | 164 (62%) | 21 (8%) | 80 (30%) |
| Chloramphenicol | C | 30 | 240 (91%) | 3 (1%) | 22 (8%) |
| Antimicrobial agents | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sampling site | AP | AK | G | KF | NOR | DXT | NA | E | TM | NI | T | PB | PG | IMI | MEM | CTX | CAZ | C | TOTAL | MARI |
| H1 | 46 | 0 | 0 | 0 | 1 | 40 | 4 | 47 | 35 | 2 | 45 | 4 | 51 | 0 | 0 | 9 | 20 | 3 | 307 | 0.72 |
| H2 | 28 | 1 | 0 | 7 | 1 | 23 | 7 | 1 | 27 | 2 | 25 | 3 | 31 | 0 | 0 | 7 | 3 | 0 | 166 | 0.77 |
| H3 | 40 | 2 | 2 | 17 | 12 | 34 | 11 | 31 | 29 | 0 | 36 | 10 | 46 | 0 | 0 | 7 | 20 | 5 | 302 | 0.83 |
| H4 | 38 | 4 | 2 | 26 | 0 | 43 | 19 | 45 | 38 | 6 | 35 | 11 | 45 | 0 | 0 | 7 | 2 | 2 | 317 | 0.83 |
| H5 | 11 | 0 | 0 | 0 | 0 | 10 | 5 | 0 | 6 | 0 | 12 | 3 | 20 | 1 | 0 | 5 | 3 | 6 | 82 | 0.61 |
| H6 | 69 | 2 | 0 | 26 | 20 | 67 | 14 | 65 | 72 | 0 | 67 | 17 | 70 | 1 | 1 | 27 | 32 | 6 | 556 | 0.88 |
| Total | 232 | 9 | 4 | 76 | 34 | 217 | 60 | 189 | 207 | 10 | 220 | 47 | 263 | 2 | 1 | 62 | 80 | 22 | 1, 730 | - |
3.5. Screening for Antimicrobial Resistance Determinants
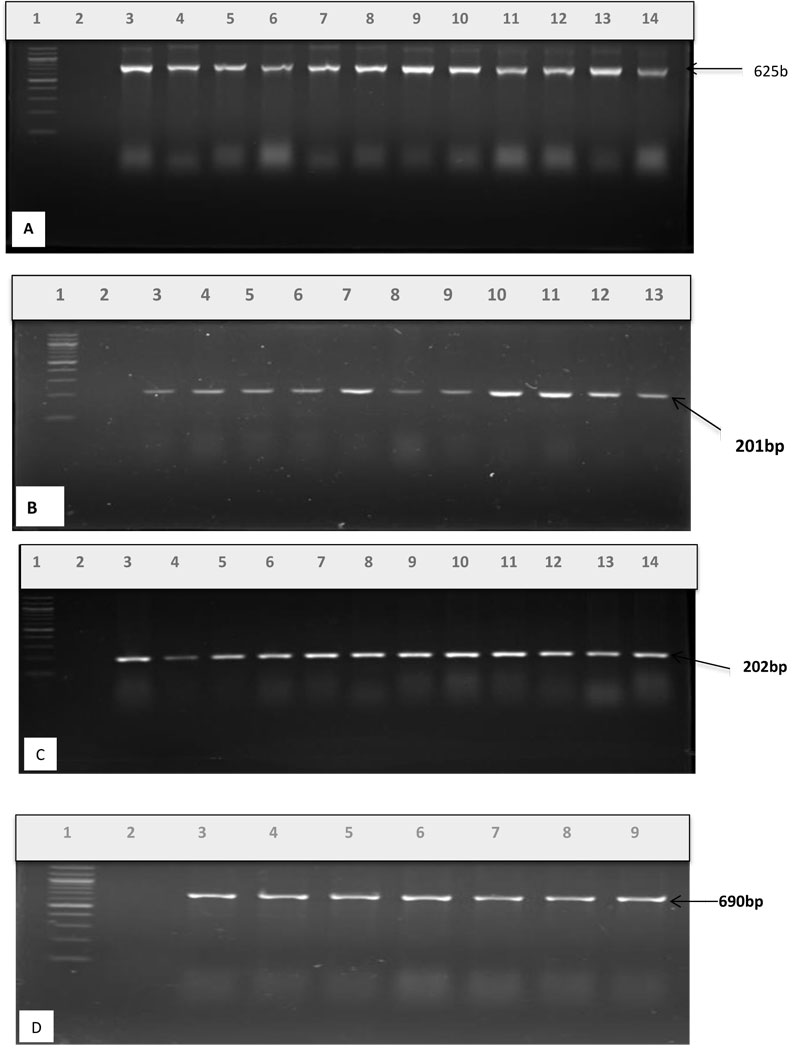
PCR was used to screen for resistance genes mediating resistance to penicillin G, ampicillin, trimethoprim, tetracycline, doxycycline and erythromycin. The results showed that 12% of 207 sulphonamide-resistant isolates possessed the sulII gene, whereas sulI gene was not harbored at all. For 220 E. coli isolates resistant to tetracyclines, only 25% harbored tetA resistance gene to tetracycline. Among 263 penicillin G and 232 ampicillin resistant isolates, the ampC gene was 28% and blaTEM gene was 47%. Fig. (1A-D) show the PCR products of the resistance genes profiled.
4. DISCUSSION
DEC strains are mainly associated with gastrointestinal infections, such as diarrhea [9, 34]. Though diarrhea in South Africa is the main cause of morbidity and death in children, the true impact of childhood diarrhea is not precisely known [35]. In this study, we confirmed the presumptive E. coli isolates from diarrheagenic stool samples of patients in medical facilities within the Buffalo City Metropolitan Municipality, evaluated the prevalence of DEC pathogenic strains, and determined their antibiotic resistance patterns and their resistance determinants.
In the present study, 218 (82%) isolates were identified as DEC strains by PCR method. The findings are incongruent with the previous study done in South Africa by Makhari [36], who reported high prevalence of DEC from diarrhea samples collected in under five DEC in South Africa. However, it is in contrast with studies conducted by Kabir et al. [37] and Omran et al. [38] as they reported the prevalence of 18% and 4% respectively of DEC among their cohorts. The predominance of DEC differs across regions and even among and within countries. In this study, four out of six DEC screened for were detected with enterotoxigenic E. coli being the highest prevalent pathotype identified with (40%) of the lt gene, followed by enteroaggregative E. coli with 18% [where 71% (34/48) possessed Eagg gene and 33% (16/48) had AafII gene]; diffusely adherent E. coli (14%) with the daaE gene and enteropathogenic E. coli (11%) had eae gene. EIEC and EHEC strains were not detected among the isolates, which signifies that their role in diarrhea is limited in Buffalo City. These findings are almost in agreement with those of the study conducted in Iran by Salmani et al. [10] in which they also reported ETEC as the most commonly isolated strain (ST, 62% and LT, 25%) followed by EAEC (24%), DAEC (13%), EIEC (7%) and EPEC (5%).
ETEC is a progressively recognized cause of childhood diarrhea among all E. coli pathotypes, especially in developing countries [39, 40]. The predominance of ETEC among diarrheal infections in this study is much similar to a recent report in a study by Salmani et al. [10] where ETEC was the most prevalent pathotype with an incidence of 62% though higher than the finding in this study. Ifeanyi et al. [41] reported ETEC possessing lt gene as responsible for 37.5% diarrhea infections in Nigeria, which agrees with the results of this present study. The prevalence of EAEC in this study was lower than other investigative studies conducted in developing countries such as the one reported by Dutta et al. [42] where EAEC was 48.2%. EAEC is generally regarded as the causative agent of diarrheal infection in infants in poor settings. The findings from this study are in line with a previous study conducted also in South Africa in 2007 by Samie et al. [43] who indicated that 16.5% of study patients with diarrhea were infected with EAEC strains. The high percentages of EAEC strain observed from other studies could probably be attributed to the daily living conditions in study sites or to the epidemiological characteristics of EAEC which could be linked to possible sources of infection and routes of transmission [44]. The prevalence of DAEC in this study was 14% of the total confirmed isolates and this result is similar to the findings of the study done by Lozer et al. [45] in which they reported DAEC with a prevalence of 11.6% among their study samples. The prevalence of EPEC was lower in this study, with 11% of positive isolates harbouring eae gene than in a study conducted by Moshtagian et al. [46] where it was the most predominant strain associated with diarrhea. However, this finding correlates with the report from a study done by Ali et al. [47], where EIEC and EHEC strains were not detected as it was the case in this study. Similar studies conducted by Pourakbari et al. [48],) and Al-Dulaimi [49], did not isolate EIEC strains in diarrhea samples they investigated as it is with our study. Worthy of note is that the prevalence of the different pathotypes of E.coli was highest in age group below 5 years of age. This could be attributed to the fact that kids under this age bracket are used to picking up things and are less hygienic when compared with older age groups hence the prevalence of diarrhea is always higher among them.
Antibiotic resistance in E. coli and bacteria, in general, has become a foremost health concern globally. The present study tested all the positive study isolates for antibiotic susceptibility against a panel of eighteen antibiotics that are commonly used for the empiric treatment of diarrhea. The isolates exhibited high levels of resistance to penicillin G (99%), ampicillin (88%), trimethoprim (84%), tetracycline (83%), doxycycline (82%), and erythromycin (71%). These findings are in accordance with the study done in Chellapandi et al. [50] in northern India where high resistances were observed against the above mentioned antibiotics in this study. Therefore, these findings suggest that these antibiotics should not be considered as empiric treatment of DEC infections as it could further fuel the spread of antibiotic resistance in the community. The most important factors that contribute to the spread of antibiotic resistance could be abuse and poor stewardship in the prescription and usage of the drug in developing countries for infections that may not require antibiotics as well as administration of inadequate regimen of the drug [22].
Multiple antibiotic resistance index was calculated in order to determine the health risks that are associated with the high level and development of antibiotics resistance in this study area. In interpreting the MARI, when the MARI is ≥0.2 it implies that the bacteria might have originated from high risk-sources of the environment in which antibiotics are frequently used or abused [51]. This present study has shown that MARI value for all the isolates was more than the expected threshold range of 0.2. The average MARI was 0.8 which is four times above the 0.2 limit thus suggesting that the E. coli isolates may have originated from high risk-sources of contamination that could be linked to either contaminated drinking water or food which possibly could result in multidrug resistance increase.
The screening of the antimicrobial resistance determinants was performed based on the observed phenotypic resistances. The resistance determinants profiled were sulphonamide resistance gene (sulI), genes encoding β-lactamases (ampC and blaTEM) and tetracycline resistance gene (tetA) amongst the genes encoding resistance factors to the antibiotics that the isolates showed resistance. The results showed that 12% of 207 sulphonamide-resistant isolates possessed the sulII gene, whereas sulI gene was not harboured, thus indicating that other genetic factors might be responsible for the observed phenotypic resistance against sulphonamide. For 220 E. coli isolates that showed resistance to tetracycline, only 25% harboured tetA resistance gene and this is because there are many variants of tet genes that encode for tetracycline resistance. Among 263 penicillin G and 232 ampicillin resistant isolates, the ampC gene was 28% and blaTEM gene was 47%. These results show that the observed multidrug resistance of the isolates to the antibiotics could probably be attributed to many resistance genes harboured by the organism order than the ones profiled. The spread of these antimicrobial resistance genes might be via horizontal gene transfer mechanisms [19, 21].
CONCLUSION
In conclusion, our findings have shown a high proportion of DEC among study patients. This suggests that DEC probably has an important role in the aetiology of diarrhea in the study sites. This should be considered in the surveillance programme of diarrheal infections. ETEC was the most predominant pathotype detected from diarrheal stool samples in Buffalo City Metropolitan Municipality. The increased levels of multidrug resistance profiles observed among E.coli isolates to penicillin G, ampicillin, trimethoprim, tetracycline and doxycycline could be a major health problem as they could initiate the spread of antimicrobial resistance in bacteria within the community. These findings require high surveillance for risk assessment and strategies that are essential for control and protection of public health. The results of this study could add more data to the epidemiological database of the country and help medical personnel in the diagnostic and treatment of gastroenteritis in the Province.
AUTHORS’ CONTRIBTIONS
BCI conceived, designed, collected data, analysed, interpreted results and prepared the article for publication, supervised and revised article for publication. NAM collected the samples, performed the experiment, and gathered the data, collated the data, analysed, interpreted results, LCO revised the manuscript.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study has been approved by the University of Fort Hare Research ethics board, South Africa (Protocol number IWE001).
HUMAN AND ANIMAL RIGHTS
No animals were used in this research. All human research procedures were followed in accordance with the ethical standards of the committee responsible for human experimentation (institutional and national), and with the Helsinki Declaration of 1975, as revised in 2013.
CONSENT FOR PUBLICATION
Informed consent was obtained from all participants.
AVAILABILITY OF DATA AND MATERIALS
The datasets used and/or analysed during the current study are available within the manuscript.
FUNDING
This study was funded by a grant from South African Medical Research Council.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The authors would like to express their gratitude to the South African Medical Research Council, the study participants and the Nurses and Midwives who helped with sample collection. Without them, the study would not have been successful.


