All published articles of this journal are available on ScienceDirect.
Resistance Profile of Vibrio spp. Strains Collected from Lagoon Bays and Wastewater in the City of Abidjan, Côte d'Ivoire, from January to June 2017
Abstract
Background:
Lagoons and wastewater constitute aquatics environments which receive or accounts for most domestic discharges. These waters constitute an important ecosystem for the proliferation of microorganisms. The microorganisms that harbor these waters can provide information on the persistence of certain diseases in the human population, including gastrointestinal infections such as Cholera. The genus Vibrio contains pathogenic aquatic bacteria found in lagoon bays and wastewater.
Objective:
The main objectives of this work were to confirm the presence of Vibrio spp. in lagoon bays and sewage of the city of Abidjan over the entire interepidemic period, and to evaluate their sensitivity to commonly used antibiotics.
Methods:
The isolation and identification of the microorganisms were carried out using classical bacteriological techniques (biochemical test, API 20E gallery). When necessary, serotyping was carried out using agglutination tests on slides. Antibiotic susceptibility testing was carried out using the Kirby-Bauer disk diffusion (KBDD) method.
Results:
This study identified 12 bacterial strains, 9/12 (75%) of which were Vibrio sp. strains. Two Vibrio species, namely Vibrio parahaemolyticus 2/9 (22%) and 7/9 Vibrio cholerae (78%) were identified.
V. cholerae was isolated from both sewage and lagoon waters with dominance of serotype O1. The V. cholerae O1 and non-O1 strains showed a high level of resistance to sulfonamides, quinolones, fluoroquinolones, and moderate sensitivity to penicillins and tetracyclines. Resistant V. parahaemolyticus strains were also identified.
Conclusion:
The increased resistance of these bacteria could pose potential problems in the treatment of epidemics and other communicable diseases. The emergence of these multi-drug resistant strains of the genus Vibrio should prompt the Ivorian health authorities to maintain an epidemiological surveillance network for waterborne diseases throughout the country and to continue bacteriological sampling to monitor Vibrio's sensitivity to antibiotics.
1. INTRODUCTION
Discharges of untreated wastewater to the environment can be a source of infection by pathogenic microorganisms from a variety of sources [1]. According to the World Health Organization, each year nearly 1.8 million people, 90% of whom are children under five years of age, die from diarrheal diseases [2]. However, globally, the vast majority of diarrheal diseases are attributable to poor quality drinking water and inadequate wastewater sanitation [3]. While wastewater sanitation has become an imperative for modern societies, African countries face serious challenges in terms of both water supply and wastewater disposal and treatment [4]. It should be noted that 78% of African cities do not have any formal sewerage disposal and treatment services, and where they exist, most of them are outdated [5]. In these countries, the inadequacy of sanitation networks and the fact that the majority of the population (more than 30% in Abidjan) is not connected to the sewerage system mean that untreated wastewater is discharged into the natural environment [4].
In Côte d'Ivoire, the city of Abidjan was developed on the periphery of the Ebrié lagoon. The lack of a real sanitation system leads to the abusive discharge of wastewater [domestic and agro-industrial wastewater] into the natural environment without prior treatment [6]. The lagoon thus becomes the obligatory receptacle for all the wastes generated by human activities [7]. These discharges are responsible for eutrophication and microbial pollution [8], which are particularly accentuated by the confinement of the water in the city's bays. The aquatic environment then becomes the reservoir of many microorganisms potentially pathogenic to humans [9]. The aquatic environment constitutes the natural habitat for bacteria of the genus Vibrio [10-12]. This work reports the presence of pathogenic bacteria in the Ebrié lagoon, that contribute to the endemic nature of certain waterborne diseases such as cholera, gastroenteritis and septicemia [8, 10, 13-16]. Unlike many enteric pathogens, vibrios are well adapted to survive outside humans naturally in the aquatic environment [17]. The ubiquity of these bacteria in the aquatic environment and the potential severity of these infections are a cause for concern, given the increasing resistance to different families of antibiotics of certain vibrios that are emerging pathogens [18-23]. Thus, the general objective of this work was to study antibiotic resistance in environmental bacteria of the genus Vibrio isolated from Ebrié Lagoon bays and wastewater.
2. MATERIALS AND METHODS
2.1. Setting, Type and Duration of Study
This prospective experimental study was carried out in two stages:
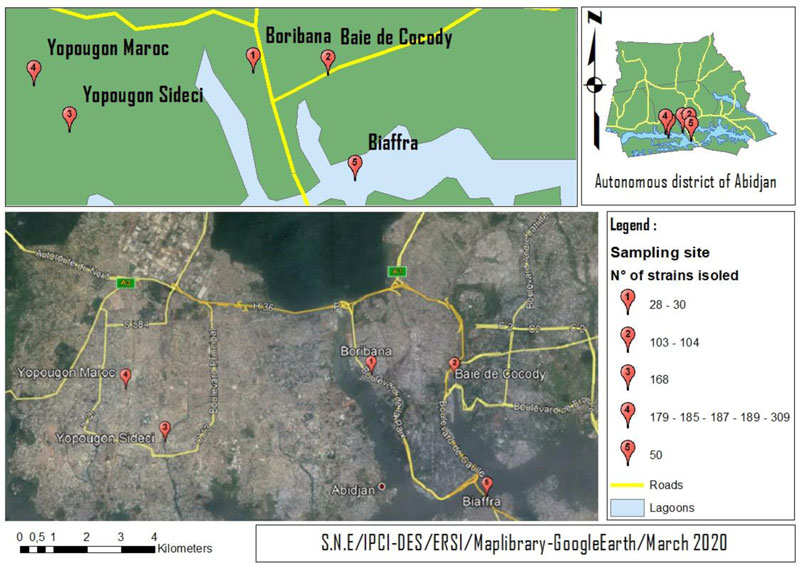
Water samples were taken from the lagoon bays: Boribana (Station 1: Lagoon Bay), Treichville Biafra (Station 2: Lagoon Bay), Cocody Bay (Station 3: Lagoon Bay) and the collector arrival points in Yopougon Maroc and Sideci (Station 4: Wastewater). Microbiological and biochemical analyses took place in the laboratories of the Pasteur Institute of Côte d'Ivoire (IPCI). During this study, which took place from January to July 2017, monthly sampling campaigns per sampling sites (5 sites) (Fig. 1) were carried out across the district of Abidjan. The sampling of wastewater and water from the lagoon bays was carried out at a rate of 1 litre (l) per sample. These samples were collected in sterile glass vials, stored at approximately 4°C in a cooler containing cold accumulators and transported to the laboratory within approximately 3 hours after sampling.
2.2. Microbiological Analysis
The bacteriological analysis was carried out in different steps of classical microbiology, namely enrichment, isolation, and identification [16, 24-26].
2.2.1. Enrichment
One (01) ml of water taken from the surface portion of the samples was inoculated in 9 mL of alkaline peptone water (EPA) and incubated at 37ºC for 24 hours.
2.2.2. Isolation
From the surface part of the inoculated alkaline peptone water, 10 µl was taken and then inoculated on TCBS (Thiosulfate Citrate Bile Sucrose) agar plates, which were—incubated at 37°C for 18-24 hours. The appearance of suspect colonies (large yellow -sucrose positive or green-sucrose negative colonies) on TCBS agar was considered. These were transferred to NAG (alkaline nutrient agar) and incubated for 24 hours at 37°C for re-isolation. Bluish colonies with a translucent appearance on GNA agar were considered.
2.2.3. Identification and Rapid Orientation Tests
Three rapid orientation tests were performed on the suspect colonies. These are: the study of mobility, Gram stain and cytochrome-oxidase test.
All suspect bacteria of the genus Vibrio on TCBS and GNA media, presenting as Gram-negative bacilli with monotrich polar ciliature mobility and possessing cytochrome oxidase C, were subjected to biochemical identification tests.
2.2.4. Biochemical Identification
The biochemical identification was carried out with the Le Minor reduced rack and with the API 20E gallery.
The bacteria supposedly belonging to the genus Vibrio are those that possessed an oxidase and had polar mobility. They produced indole, had LDC (lysine decarboxylase), but not urease and LDA (lysine deaminase). They did not produce gas or H2S (hydrogen sulphide) and did not use lactose.
2.3. Identification of Vibrio cholerae O1 strains
The identification of V. cholerae O1 strains was carried out using specific serum immunosera by slide agglutination. Immune serum O1 (antibodies to V. cholerae O1, Bio-Rad®F-92430 Mannes-la-coquette) was used.
2.4. Determination of Antibiotic Profiles
Antibiogram was used to determine the resistance profile of isolated Vibrio strains. It was performed using the agar disc diffusion method described by Kirby-Bauer [27].
The following antibiotics were tested: Azithromycin (AZM), Nitrofurantoine (FTN), Chloramphenicol (CHL), Tobramycin (TMN), Amikacin (AN), Norfloxacin (NOR), Ciproflaxin (CIP), Pefloxacin (PEF), Nalidixic Acid (NAL), Trimetoprim-sulfamethoxazole (SXT), Sulfonamide (SUL), Tigecyclin (TGC), Minocyclin (MNO), Doxicyclin (DOX), Tetracyclin (TET), Imipenem (IPM), Fosfomycin (FOS), Aztreonam (ATM), Cefotaxim (CTX), Ceftriaxon (CRO), Cefalotin (CF), Piperacillin (PIP), Ticarcillin (TIC), Amoxiccillin / Clavulanic Acid (AMC), Ampicillin (AMP), and Amoxiccillin (AMX). If all were tested on V.Cholerae strains, only one part was used for the V. parahaemolyticus.
The CLSI interpretative criteria for disk diffusion susceptibility testing of Vibrio spp. (CLSI, 2015) published in the M45 3rd edition were used for amoxicillin-clavulanic acid, cefotaxime, imipenem, chloramphenicol, ciprofloxacin, amikacin, piperacillin, cefoxitin (FOX), amikacin (AN), ciprofloxacin (CIP), trimethoprim-sulfamethoxazole association, and the CASFM, (2017)for Amoxicillin (AMX), Ceftriaxone (CRO) and Ticarcillin (TIC). For others, the interpretative criteria for Enterobacteriaceae edited by the CLSI were used (CLSI, 2016). The percentage of resistance was calculated for each antibiotic by dividing the frequency of resistant bacteria by the number of bacteria tested.
3. RESULTS
3.1. Isolated Bacteria and Identification of Vibrio spp
A total of 107 samples were collected, 60 of which were lagoon water samples and 47 wastewater samples. This study identified 12 bacterial strains, including nine (9) Vibrio spp (75%), one (1) Aeromonas hydrophilla(8%) and two (2) Plesiomonas shiggeloides (17%). Among the genus Vibrio, two Vibrio parahaemolyticus (2/9; 22%) and seven Vibrio cholerae (7/9, 78%) were identified. Only one V. cholerae strain was identified as serogroup O1 (Table 1).
3.2. Spatial Distribution of Vibrio spp
Vibrio cholerae strains were isolated in 2/60 (3%) and 5/47 (11%) samples taken from the lagoon water of Cocody Bay and sewage disposal channels in Yopougon, respectively. In contrast, the Vibrio parahaemolyticus strains were isolated from the samples taken in Boribana (Attécoubé).
3.3. Determination of Antibiotic Resistance Profiles of Vibrio Strains
The antimicrobial resistance determination was carried out on 2 V. parahaemolyticus and 7 V. cholerae strains using 27 antibiotics belonging to the different groups.
The 2 strains of Vibrio Parahaemolyticus were sensitive to all antibiotics except piperacillin with 100% resistance (Table 2).
The non-O1 strains of Vibrio cholerae isolated in this study were resistant to several families of antibiotics, including sulfonamides, fluoroquinolones, tetracyclines and beta lactamins.
This study showed that the resistance of these strains to beta-lactams was 33% for molecules such as ampicillin (AMP), tircacillin (TIC), aztreonam (ATM) and fosfomycin (FOS), while it was 50% for amoxicillin (AMX), piperacillin (PIP), ceftriazone (CRO), cefoxitin (FOX) and cefotaxime (CTX).
| Strain Numbers | Origin of Isolation | Genus | Species | Antiserum O1 * |
|---|---|---|---|---|
| 28 | Lagoon | Vibrio | V. parahaemolyticus | N.E |
| 30 | Vibrio | V. parahaemolyticus | N.E | |
| 50 | Aeromonas | A. hydrophilla | N.E | |
| 103 | Vibrio | V. cholerae | - | |
| 104 | Vibrio | V. cholerae | - | |
| 168 | Sewage water | Vibrio | V. cholerae | - |
| 176 | Plesiomonas | P. shiggeloides | N.E | |
| 179 | Vibrio | V. cholerae | - | |
| 185 | Vibrio | V. cholerae | - | |
| 187 | Vibrio | V. cholerae | + | |
| 189 | Vibrio | V. cholerae | - | |
| 309 | Plesiomonas | P. shiggeloides | N.E |
| Antibiotics | Disk Content (µg) |
Antibiotics Interpretative Criteria (R ≤ I ≤ S) |
Non O1 V. cholerae (n=6) | V. parahaemolyticus (n=2) | ||
|---|---|---|---|---|---|---|
| Resistance (N (%)) | Sensitive (N (%)) | Resistance (N (%)) | Sensitive (N (%)) | |||
| Amoxiccillin (AMX) | 20 | 19 ≤ ≤ 19 | 3 (50%) | 3 (50%) | 0% | (2) 100% |
| Ampicillin (AMP) | 10 | 13 ≤ 14-16 ≤ 17 | 2 (33,33%) | 4 (66,66%) | ||
| Amoxiccillin / Clavulanic Acid (AMC) | 20 - 10 | 13 ≤ 14 – 17 ≤ 18 | 1 (16,66%) | 5 (83,33%) | 0% | (2) 100% |
| Ticarcillin (TIC) | 75 | 20 ≤ 22 - 21 ≤ 23 | 2 (33,33%) | 4 (66,66%) | 0% | (2) 100% |
| Piperacillin (PIP) | 100 | 17 ≤ 18 - 20 ≤ 21 | 3 (50%) | 3 (50%) | (2) 100% | 0% |
| Cefalotin (CF) | ≤ ≤ | 0 | 6 (100%) | |||
| Ceftriaxon (CRO) | 30 | 22 ≤ 24-23 ≤ 22 | 3 (50%) | 3 (50%) | 0% | (2) 100% |
| Cefotaxim (CTX) | 30 | 22 ≤ 23 - 25 ≤ 26 | 3 (50%) | 3 (50%) | 0% | (2) 100% |
| Cefoxitin (FOX) | 30 | 14 ≤ 15 – 17 ≤ 18 | 3 (50%) | 3 (50%) | 0% | (2) 100% |
| Aztreonam (ATM) | 30 | 17 ≤ 18 - 20 ≤ 21 | 2 (33,33%) | 4 (66,66%) | 0% | (2) 100% |
| Fosfomycin (FOS) | 200 | 12 ≤ 13 - 15 ≤ 16 | 2 (33,33%) | 4 (66,66%) | ||
| Imipenem (IPM) | 10 | 14 ≤ 14-15 ≤ 16 | 0 | 100% | 0% | (2) 100% |
| Tetracyclin (TET) | 30 | 11 ≤ 12-14 ≤ 15 | 3 (50%) | 3 (50%) | ||
| Doxicyclin (DOX) | 30 | 10 ≤ 11-13 ≤14 | 2 (33,33%) | 4 (66,66%) | ||
| Minocyclin (MNO) | 30 | 12 ≤ 13-15 ≤ 16 | 0 | 6 (100%) | ||
| Tigecyclin (TGC) | 15 | 15 ≤16 -17 ≤18 | 0 | 6 (100%) | 0% | (2) 100% |
| Sulfonamide (SUL) | 300 | 12 ≤ 13-16 ≤ 17 | 4 (66,66%) | 2 (33,33%) | ||
| Trimetoprim-sulfamethoxazole (SXT) | 1,25 – 23,75 | 10 ≤ 11 - 15 ≤ 16 | 2 (33,33%) | 4 (66,66%) | 0% | (2) 100% |
| Nalidixic Acid (NAL) | 30 | 13 ≤ 14 - 18 ≤ 19 | 16,66% | 83,33% | 0% | (2) 100% |
| Pefloxacin (PEF) | 5 | 23 ≤ ≤ 24 | 50,00% | 50,00% | ||
| Ciproflaxin (CIP) | 5 | 15 ≤ 16 - 20 ≤ 21 | 50,00% | 50,00% | 0% | (2) 100% |
| Norfloxacin (NOR) | 10 | 12 ≤ 13-16 ≤ 17 | 50,00% | 50,00% | ||
| Amikacin (AN) | 10 | 14 ≤ 15 – 17 ≤ 17 | 0 | 100% | 0% | (2) 100% |
| Tobramycin (TMN) | 10 | 12 ≤ 13 - 14 ≤ 15 | 0 | 100% | 0% | (2) 100% |
| Chloramphenicol (CHL) | 30 | 12 ≤ 13 - 17 ≤ 18 | 0 | 6 (100%) | 0% | (2) 100% |
| Nitrofurantoine (FTN) | ≤ ≤ | 0 | 6 (100%) | |||
| Azithromycin (AZM) | 15 | 12 ≤ ≤ 13 | 0 | 6 (100%) | ||
However, less than 20% of strains of Vibrio cholerae non-O1 were observed to be resistant to the amoxicillin-clavulanic acid (AMC) combination when they were all sensitive to Imipenem (IPM).
Regarding fluoroquinolones, nearly 50% of the bacteria studied were resistant to the molecules tested.
The resistance of non-O1 Vibrio cholerae strains was of the order of 50% for the 1st generation cyclins (tetracycline (TET) but the other generations (minocycline (MNO), Tigecycline (TGC)) remained fully effective against these bacteria.
The strains were all sensitive to aminoglycosides (Amikacin (AN), Tobramycin (TMN)) and Phenicols (Chloramphenicol (CHL)) but presented a high rate of resistance to sulfonamides (67% of strains resistant to sulfonamides (SUL) and 33% resistance to the combination Trimetoprime-Sulfamethoxazole (SXT) (Table 2).
The Vibrio cholerae O1 strain isolated from wastewater was resistant to penicillins (ampicillin (AMP), piperacillin (PIP), quinolones (nalidixic acid (NAL), fluoroquinolones (pefloxacin (PEF), norfloxacin (NOR), ciprofloxacin (CIP)) and sulfonamides (sulphonamides (SUL) and cotrimozaxol (SXT)). It is very sensitive to the action of aminoglycosides (Amikacin (AN), Tobramycin (TMN), tigecycline (TGC)), cephalosporins (cefalotin (CF), cefotaxime (CTX, cefoxitin (FOX), ceftriaxone (CRO)) and tetracyclines (tetracyclines (TET), doxycycline (DOX), minocycline (MNO)).
4. DISCUSSION
The presence of V. cholerae in the waters of lagoon bays and sewage suggests faecal contamination caused by poor excreta management and open defecation at the edge of lagoons or in vacant lots at the edge of neighborhoods [28]. The presence of V. cholerae and V. parahaemolyticus in lagoon bays and sewage could pose serious problems that would be even more serious if these bacteria are resistant. The use of antibiotics, both in human and veterinary medicine, exerts selection pressure over commensal and pathogenic bacteria from humans or animals that can reach the natural environment [continental waters, coastal waters, or wastewater]. This could lead to the emergence and dissemination of antibiotic resistance genes in environmental microbial communities.
The existence of links between different ecological compartments leads to interest in the pathways of resistant bacteria and resistance genes in the environment. The resistance of these strains is independent of their origin [29].
The resistance of Vibrio spp documented so far to Beta-lactam is mediated by plasmid coding at the same time for many other antibiotics [30, 31]. Plasmid-mediated penicillinases have often been found in most gram-negative bacilli, the best known of which are the narrow spectrum penicillinases TEM1 and SHV1 [32]. A study carried out by Aucher in 1998 reports that ampicillin is generally inactive against the majority of vibrios. In this study, more than 30% of vibrio cholerae strains were resistant to Ampicillin. According to Guévard [22], this resistance is due to multiple mechanisms, including transferable plasmid and chromosomal resistance. The resistance of V. cholerae to 3rd generation cephalosporins (ceftriaxone, cefoxitin, and cefotaxime)is linked to the production of a cephalosporinase or Broad Spectrum Beta-Lactamase (ESBL). This enzyme-like resistance mechanism is developed in most Gram-negative bacilli [33]. Fluoroquinolones are generally active on V. cholerae, but this study showed an increase in the resistance of Vibrio to fluoroquinolone to about 50%. In general, vibrios are usually sensitive to beta-lactam antibiotics, aminosides, quinolones, tetracyclines, chloramphenicol, sulfonamides, trimethoprim, furans, nalidixic acid and fluoroquinolones. This study showed an increase in the resistance of Vibrio Cholerae strains to antibiotics. Indeed, this work observed a decrease in the sensitivity of most antibiotics to vibrio, unlike previous observations, in particular, that of Tiécoura in 2010 [34]. Multi resistance acquired through transferable genetic elements (plasmids, conjugative transposons, genomic islands) within Vibrionaceae has become very common [35, 36]. This ability of Vibrio cholerae non O1, non O139 to acquire genetic material that could modify its virulence and its resistance has also been demonstrated by Arteaga et al. [37].
This work has also shown that although it has been an accepted fact that the resistance of vibrio to antibiotics has increased in the endemic areas of Cholera [38], the resistance of these bacteria to Fluoroquinolones was a new observation in Côte d'Ivory to our knowledge.
Horizontal transfer of antibiotic resistance in V. cholerae is commonly associated with plasmids [39]. Other conjugative elements allow the acquisition of this resistance. For instance, a conjugative transposon called SXT confers resistance to several antibiotics [Chloramphenicol, Streptomycin, Ampicillin, Neomycin, Tetracycline, Gentamicin, Sulfonamide and Furazolidone] [40-42]. Unlike plasmids, the genome of the SXT transposon does not contain the genes necessary for autonomous replication and segregation. Nevertheless, this structure has all the genes necessary for its excision, transmission and integration into the chromosome. The presence of V. cholerae in wastewater supports the hypothesis that wastewater discharges contribute to an increase in bacterial contamination of lagoon bays. The distribution of bacteria in Ebrié lagoon bays and wastewater varies from one sampling site to another. The detection of V. parahaemolyticus in the lagoon bays (Boribana Attenuated) could be explained by the hydrodynamics and geographical location. This station is located in bays, where confinement reduces the flow velocity and water renewal rate [43]. Boribana's remoteness from the sea makes the water there less salty and more eutrophic. Given the halophilic nature of V. parahaemolyticus, it is not surprising that this organism is found in aquatic environments around the world, particularly in waters with a salinity intermediate between fresh and seawater. The combination of these factors creates a favorable environment for the survival and development of bacteria of the genus Vibrio.
CONCLUSION
The results of this study showed the presence of 2 Vibrio species among the strains isolated from the wastewater: Vibrio cholerae (V cholerae O1 and V cholerae non O1) and Vibrio parahaemolyticus, with a predominance of Vibrio cholerae non O1. The isolated strains showed resistance to at least one usually active antibiotic, indicating a relatively high circulation of resistant strains. Since the effective management of gastroenteritis is based primarily on the replacement of hydroelectrolytic loss and antibiotic therapy, the circulation of multi-resistant Vibrio spp may compromise the effectiveness of probabilistic treatments and explain treatment failures.
In view of this bacterial pollution in wastewater and lagoon bays, we can propose possible solutions to try to reduce the contamination: 1) - build a good sanitation system for the collection of domestic wastewaters; 2)build treatment centers for domestic wastewater and industrial effluents; and 3) implement actions to raise awareness of the community and prescribers on the rational use of antimicrobials in order to contribute to limiting the spread of resistant strains.
In an environmental context of preserving the quality of groundwater, lagoons and marine waters where wastewater is discharged without prior treatment, a study of the impact of emerging pollutants on the growth and resistance of these bacteria must be carried out, bearing in mind that there is a risk of proliferation of these strains in groundwater and on the sea surface by the coastal current.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Not applicable.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
Declared none.


