All published articles of this journal are available on ScienceDirect.
Statistical Optimization by Response Surface Methodology to Enhance Lipase Production by Aspergillus fumigatus
Abstract
Background:
Lipases have various commercial applications and microorganisms serve as a potential source of production.
Objective:
The aim of this paper was to study the effect of interactions among different production parameters on lipase yield of Aspergillus fumigatus.
Method:
Plackett Burman and Central Composite Design (CCD) were established by using Design Expert software 10.0.
Results:
In the present study, interactions were studied for six different variables such as inoculum size, pH, temperature, galactose concentration, peptone concentration and incubation time. In Plackett-Burman design, galactose concentration, peptone concentration, pH and incubation time were found to be important factors. Using the statistical approach, the optimum factors were found to be as: galactose concentration (1.5%), peptone concentration (1.8%), pH (10.0) and incubation time (72 h) at 45°C under response surface curves. Upon statistical analysis, the coefficient of determination (R2) obtained was 0.9318 which showed that the model was significant.
Conclusion:
The statistical tools used predicted the optimal conditions for the production of the lipase. The optimized parameters were galactose concentration 1.5%, peptone concentration 1.4%, temperature 45°C, pH 10.0 and incubation time of 72 h for obtaining a maximum lipase activity of 6.22 U/ml.
1. INTRODUCTION
Lipases (EC 3.1.1.3) are carboxylic ester hydrolases that catalyse the hydrolysis of triacylglycerol to glycerol and free fatty acids [1, 2]. Emulsified esters of glycerine and long-chain fatty acids such as triolein and tripalmitin were split by true lipases. Microbial lipases play an immense role in biotechnology because they are stable over a different range of pH, at high temperature and also in various organic solvents. Lipases are increasingly becoming popular for various industrial applications because of their diverse enzymatic properties and substrate specificities [3, 4]. Out of various sources fungi were potentially better for isolating different lipases with unique properties. Many fungal lipases producing microbes include Aspergillus, Geotrichum, Rhizopus, Mucor, Rhizomucor and Penicillium [5-7]. Lipases are versatile enzymes which are capable of valuable transformations in the different areas like organic chemistry, biofuels, oleo-chemistry and pharmaceutical industries [8, 9].
Filamentous fungi produce industrially important enzymes in batch and fed-batch cultures through submerged fermentation [10]. Submerged processes have some advantages such as higher homogeneity of the culture medium and can control temperature and pH as compared to solid-state processes [11].
The statistical optimization is more efficient in terms of cost and time consumption as compared to the method of changing one variable at a time [12], and also, the number of experiments can be less as well as the interaction among different variables can be studied simultaneously. Many researchers have carried out the study for the production of lipases by microorganisms by using these techniques [13-15]. Plackett–Burman design is an efficient method that can be used for the screening of different variables for further optimization [16]. The aim of this work was to find out the most significant parameters for the production of lipase by fermentation and to optimize parameters by using Response Surface Methodology.
| Run | Inoculum Size (spores/ml) |
Galactose (%,w/v) |
Peptone (%,w/v) | pH | Incubation Time (hrs) |
Temperature (°C) |
Response (U/ml) |
|---|---|---|---|---|---|---|---|
| 1 | 3 | 2 | 1.5 | 11 | 96 | 50 | 4.16 |
| 2 | 1.8 | 1 | 1.5 | 9 | 48 | 40 | 4.82 |
| 3 | 1.8 | 2 | 1.5 | 11 | 96 | 40 | 5.6 |
| 4 | 3 | 1 | 1.5 | 11 | 48 | 40 | 4.90 |
| 5 | 3 | 1 | 2.1 | 11 | 96 | 40 | 5.61 |
| 6 | 1.8 | 1 | 2.1 | 11 | 48 | 50 | 4.42 |
| 7 | 1.8 | 2 | 2.1 | 11 | 48 | 50 | 6.09 |
| 8 | 3 | 2 | 1.5 | 9 | 48 | 50 | 3.85 |
| 9 | 1.8 | 1 | 1.5 | 9 | 96 | 50 | 2.38 |
| 10 | 1.8 | 2 | 2.1 | 9 | 96 | 40 | 9.51 |
| 11 | 3 | 1 | 2.1 | 9 | 96 | 50 | 4.25 |
| 12 | 3 | 2 | 2.1 | 9 | 48 | 40 | 3.94 |
2. MATERIALS AND METHODS
2.1. Materials
The chemicals used were Potato Dextrose Agar (PDA), isopropanol, Tris-HCl buffer, p-nitrophenyl benzoate (p-NPB), p-nitrophenol (p-NP), galactose, NaCl, CaCl2, Bradford reagent, peptone, p-nitrophenol, and ammonium sulphate. All the chemicals used in the present investigation were procured from Hi-Media and Merck, Mumbai, India and from Sigma Aldrich (U.S.A.)
2.2. Microorganism and Enzyme Production
Samples were collected from different oil contaminated soil such as those from railway track, Himachal Road Transport Corporation (HRTC) workshop, dairy plant and garden soil in and around Shimla district, Himachal Pradesh. The production medium containing (w/v) Peptone 1.5%, NaCl 0.5%, CaCl2 0.1% and Tween 80 1% [17] was autoclaved at 15 lb/inch2 pressure in 121°C for 15 min and cooled at room temperature. The sterile production medium (50 ml in 250 ml Erlenmeyer flask) was inoculated with 2.4 ×105 spores and incubated for 3 days at 37°C. The culture broth was checked for enzyme activity.
2.3. Lipase Assay
Assay method given by Winkler and Stuckmann (1979) was used for lipase activity [18]. One unit of lipase activity was defined as the amount of enzyme required to release one micromole of p-nitrophenol from the substrate per minute under standard assay conditions.
2.4. Identification of Lipase-Producing Fungi
The isolated strain with high lipase activity was identified by 18S r RNA sequencing. Then, the evolutionary status and species of the strains were determined through sequence similarity analysis (BLAST, CLUSTALW) and by construction of phylogenetic tree (Neighbor-Joining Method, Jukes Cantor Method and MEGA5).
2.5. Experimental Design
Six independent production variables such as inoculum size, galactose concentration, peptone concentration, pH, incubation time and temperature were investigated by RSM for combined interactions for lipase production from Aspergillus fumigatus. Plackett-Burman and Central Composite Designs (CCD) of Design expert software (version 10.0.3) were used to carry out RSM.
2.6. Plackett-Burman Analysis
The production parameters were analysed for six variables at two different levels, minimum and maximum. Plackett-Burman design and regression analysis were conducted by Design Expert (Version 10.0). The effect of each factor was calculated by using the following formula:
E = (ΣM+−ΣM−)/N,
where E is the effect of the tested variable and M+ and M- are responses (enzyme activity) of trials at which the parameter was at its higher and lower level, respectively, and N is the number of experiments carried out [19].
2.7. Central Composite Design (CCD)
The concentration of various independent parameters which showed a significant effect (galactose concentration, peptone concentration, pH and incubation time) on lipase production was optimized by central composite design (CCD) [20].
3. RESULTS AND DISCUSSION
3.1. Identification of Strain
Fungal isolate A6 was characterized by partial sequencing of 18s rRNA gene from Exceleris Genomics, Ahmedabad and identified as Aspergillus fumigatus KU891253.1. Sequence analysis of the obtained sequence revealed a homology of more than 95% with Aspergillus fumigatus isolate NRRL (EF66 9931.1), Aspergillus fumigatus (KR023997.1), Aspergillus fumigatus strain LCF 20 and Aspergillus fumigatus (FJ8 67935.1) with 18s rRNA sequences.
3.2. Screening of Important Production Parameters using Plackett-Burman Design
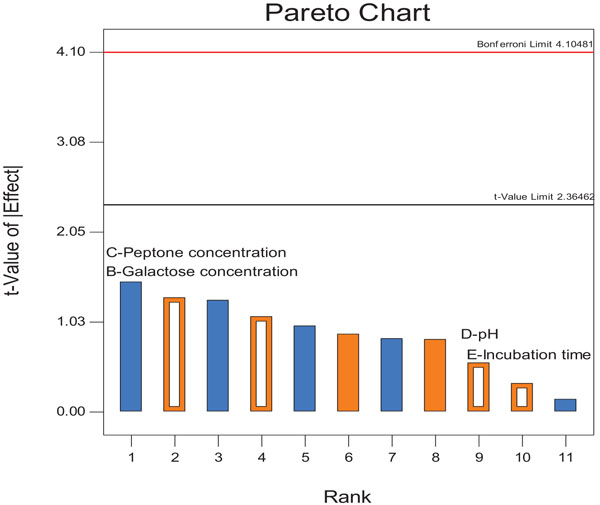
Table 1 shows the value of six parameters and their responses to lipase production. Experiments showed variations in activity from 2.38 to 9.51 U/ml and run 10 was found to be optimum which exhibited the maximum activity of 9.51 U/ml. Pareto chart was formed by results obtained from Plackett-Burman with six variables (Fig. 1) to find out the significant variables. In an earlier study, sunflower oil, glucose, peptone, agitation rate, and incubation temperature were the most significant variables for the production of lipase from Aspergillus carneus [12]. The optimization of lipase production by using the stepwise experimental design of fungus Aspergillus flavus showed that the 100 g/L wheat bran was present in the culture medium and 20 g/L soybean oil as the inducer [10].
3.3. Central Composite Design
A central composite design (Table 2) was prepared for variables having a positive effect as shown in Pareto chart for determining the optimum level and to analyze the combined effect of variables (concentration of peptone, galactose, pH and incubation time). There were 30 experimental combinations for four significant variables obtained by CCD. In another study, the three factors (sunflower oil, glucose, and agitation rate) influenced the lipase production as obtained by response surface methodology [12]. The model was analysed and used to generate the ANOVA (Table 3), showing that the model was highly reliable. The cubic process was found to be good and the model was significant (Table 4).
| Standard | Run |
Galactose Concentration (%,w/v) |
Peptone Concentration (%,w/v) |
pH |
Incubation Time (hrs) |
Response U/ml |
|---|---|---|---|---|---|---|
| 5 | 1 | 1.4 | 1.6 | 11 | 48 | 5.63 |
| 27 | 2 | 1.5 | 1.8 | 10 | 72 | 5.31 |
| 20 | 3 | 1.5 | 2.2 | 10 | 72 | 5.86 |
| 4 | 4 | 1.6 | 2 | 9 | 48 | 5.2 |
| 26 | 5 | 1.5 | 1.8 | 10 | 72 | 5.79 |
| 8 | 6 | 1.6 | 2 | 11 | 48 | 4.79 |
| 7 | 7 | 1.4 | 2 | 11 | 48 | 4.58 |
| 9 | 8 | 1.4 | 1.6 | 9 | 96 | 5.38 |
| 22 | 9 | 1.5 | 1.8 | 12 | 72 | 5.26 |
| 2 | 10 | 1.6 | 1.6 | 9 | 48 | 5.02 |
| 6 | 11 | 1.6 | 1.6 | 11 | 48 | 5 |
| 12 | 12 | 1.6 | 2 | 9 | 96 | 5.31 |
| 16 | 13 | 1.6 | 2 | 11 | 96 | 4.8 |
| 10 | 14 | 1.6 | 1.6 | 9 | 96 | 4.71 |
| 15 | 15 | 1.4 | 2 | 11 | 96 | 4.68 |
| 3 | 16 | 1.4 | 2 | 9 | 48 | 4.95 |
| 11 | 17 | 1.4 | 2 | 9 | 96 | 5.11 |
| 1 | 18 | 1.4 | 1.6 | 9 | 48 | 5.61 |
| 29 | 19 | 1.5 | 1.8 | 10 | 72 | 5.79 |
| 13 | 20 | 1.4 | 1.6 | 11 | 96 | 5.28 |
| 30 | 21 | 1.5 | 1.8 | 10 | 72 | 5.79 |
| 23 | 22 | 1.5 | 1.8 | 10 | 24 | 4.58 |
| 19 | 23 | 1.5 | 1.4 | 10 | 72 | 6.22 |
| 17 | 24 | 1.3 | 1.8 | 10 | 72 | 5.24 |
| 28 | 25 | 1.5 | 1.8 | 10 | 72 | 5.79 |
| 25 | 26 | 1.5 | 1.8 | 10 | 72 | 5.79 |
| 24 | 27 | 1.5 | 1.8 | 10 | 120 | 3.3 |
| 18 | 28 | 1.7 | 1.8 | 10 | 72 | 4.98 |
| 21 | 29 | 1.5 | 1.8 | 8 | 72 | 5.74 |
| 14 | 30 | 1.6 | 1.6 | 11 | 96 | 4.71 |
| Source | Sum of Squares | df | Mean Square | F Value |
P Value Prob>F |
Remark |
|---|---|---|---|---|---|---|
| Model | 8.92 | 14 | 0.64 | 14.63 | – | Significant |
| A-Galactose concentration | 0.20 | 1 | 0.20 | 4.63 | – | – |
| B-Peptone concentration | 0.29 | 1 | 0.29 | 6.67 | – | – |
| C-pH | 0.32 | 1 | 0.32 | 7.39 | – | – |
| D-Incubation time | 0.47 | 1 | 0.47 | 10.80 | – | – |
| AB | 0.66 | 1 | 0.66 | 15.06 | – | – |
| AC | 2.250E-004 | 1 | 2.250E-004 | 5.165E-003 | – | – |
| AD | 1.600E-003 | 1 | 1.600E-003 | 0.037 | – | – |
| BC | 0.16 | 1 | 0.16 | 3.77 | – | – |
| BD | 0.15 | 1 | 0.15 | 3.49 | – | – |
| CD | 4.225E-003 | 1 | 4.225E-003 | 0.097 | – | – |
| A2 | 0.76 | 1 | 0.76 | 17.49 | – | – |
| B2 | 0.12 | 1 | 0.12 | 2.73 | – | – |
| C2 | 0.13 | 1 | 0.13 | 3.01 | – | – |
| D2 | 5.78 | 1 | 5.78 | 132.74 | – | – |
| Residual | 0.65 | 15 | 0.044 | – | – | – |
| Lack of fit | 0.46 | 10 | 0.046 | 1.20 | 0.4447 | Non significant |
| Pure Error | 0.19 | 5 | 0.038 | – | – | – |
| Cor Total | 9.58 | 29 | – | – | – | – |
| – | Sequential | Lack of Fit | Adjusted | Predicted | Remark |
|---|---|---|---|---|---|
| Source | p-value | p-value | R-Squarred | R-Squarred | – |
| Linear | 0.4424 | 0.0080 | -0.0044 | -0.2649 | – |
| 2FI | 0.8542 | 0.0050 | -0.1657 | -0.3316 | – |
| Quadratic | <0.0001 | 0.4447 | 0.8681 | 0.6936 | Suggested |
| Cubic | 0.3231 | 0.5308 | 0.8930 | 0.1387 | Aliased |
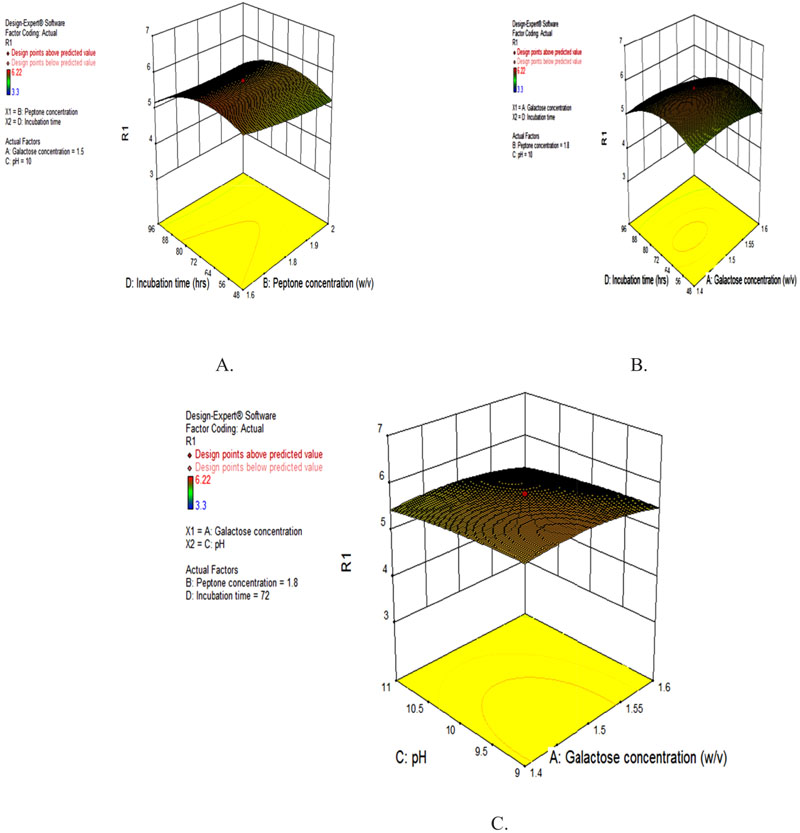
The 3D response graphs were obtained by regression analysis of CCD design data using a different combination of four significant factors for lipase production Fig. (2 A-C). The response surface graphs were more or less hill shaped and showed interactions among variables with optimum activity around concentration of galactose 1.5% (w/v), peptone 1.8% (w/v), pH 10.0 and incubation time 72 h. There was moderate interaction between peptone concentration and incubation time (Fig. 2A). Neither high nor low concentration of peptone and incubation time gave higher enzyme yield. Peptone at a concentration of 1.8% and incubation time of 72 h gave an optimum yield of the enzyme but incubation time had a distinct impact on enzyme production in comparison to peptone concentration. In a recent study, lipase form tolerant Rhizopus sp showed maximum lipase activity in the presence of peptone (2%, w/v) [21]. In other studies, 72 h was reported as the optimal fermentation time for Aspergillus niger lipase grown in solid-state fermentation, with a subsequent decrease in activity seen at 96 h and 120 h [22, 23]. There was a moderate interaction between galactose concentration and incubation time (Fig. 2B). Galactose at a concentration of 1.5% and incubation time of 72 h gave an optimum yield of the enzyme but incubation time had a distinct impact on enzyme production in comparison to galactose concentration for the production of lipase. Neither high nor low concentration of galactose and pH gave higher enzyme yield. Galactose at a concentration of 1.5% and a pH of 10.0 gave an optimum yield of enzyme lipase (Fig. 2C).
The model was very significant as suggested by ANOVA which was evident from the coefficient of variation (CV%=4.01) and low-pressure value (2.93) Table 5. The coefficient of determination (R2) (0.9264) suggested that the model was significant with the obtained values, which was better than R2 value of the 0.81 and 0.91 for lipase production from Burkholderia sp. C20 and Candida sp. 99-125 [24, 25].
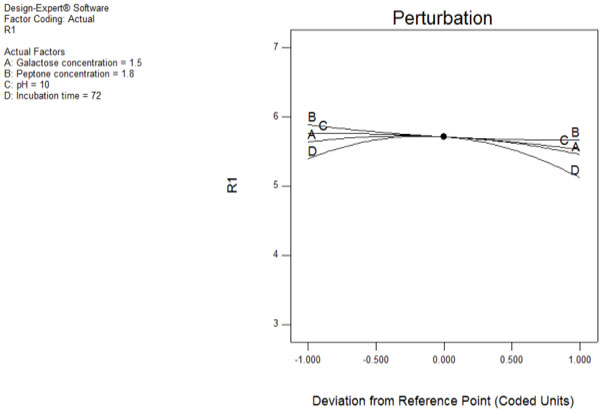
3.4. Perturbation Plot
A perturbation plot shows the optimum values of variables for lipase production from Aspergillus fumigatus (Fig. 3). Perturbation plot helps to compare the effects of all the significant variables at a particular point in the RSM design space and response is plotted by changing only one variable over its range, while holding all other factors constant.
| Std. Dev. | 0.21 | R-Squared | 0.9318 |
| Mean | 5.21 | Adjusted R-Squared | 0.8681 |
| C.V.% | 4.01 | Predicted R-Squared | 0.6936 |
| PRESS | 2.93 | Adequate Precision | 17.616 |
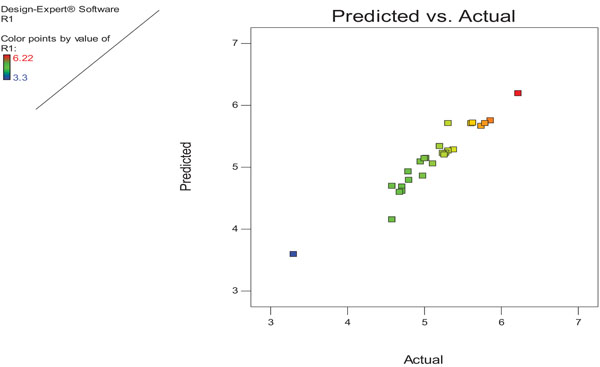
3.5. Validation of the Model
The RSM adequacy was shown by comparing the experimental data and the predicted values, by generating a fitted line plot for the obtained results, showing its closeness or deviation from the fitted line. The experimental and predicted values are shown in Fig. (4). As calculated by ANOVA, the activity was obtained with an experimental value of 6.22 U/ml, which is nearly related to the predicted value of 6.20 U/ml. Perturbation plot shows the optimum value for variables such as: galactose concentration 1.5% (w/v), peptone concentration 1.8% (w/v), pH 10.0 and incubation time 72 h. There was a 1.4-fold increment in lipase production by performing statistical analysis.
CONCLUSION
Lipase has been found to be a potential candidate for different industrial sectors. In this work, RSM was performed for the optimization of production parameters for lipase from Aspergillus fumigatus. A quadratic polynomial obtained by the CCD was very important for determining the diffrented optimal parameters that had positive effects on lipase production. The optimum production parameters consisted of 1.5% galactose, 1.8% peptone, pH 10.0 and incubation time 72 h. After optimization, lipase activity from Aspergillus fumigatus increased by 1.4-fold.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used in the study that are the basis of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
The financial support from the Department of Biotechnology, Ministry of Science and Technology, Govt. of India, Department of Biotechnology, Himachal Pradesh University, Shimla (India), is thankfully acknowledged. The financial assistance from the Department of Environment, Science and Technology, Himachal Pradesh, in the form of a Minor Research Project is also acknowledged.


