All published articles of this journal are available on ScienceDirect.
Detection of OXA Beta Lactamases Among Clinical Isolates of Acinetobacter baumannii Isolated from Tehran Hospitals, Iran
Abstract
Background and Objective:
Acinetobacter baumannii is a non-motile Gram-negative bacterial pathogen with the history of vast resistant to antibiotics. The aim of this study was to determine the possibility of existence of OXAs genes among clinical isolates of A. baumannii obtained from Tehran hospitals.
Materials and Methods:
A total of 101 isolates were identified as A. baumannii by common biochemical and molecular tests. The susceptibility to different antibiotics was assessed with Kirby-Bauer disk diffusion method. Phenotypic Detection of MBLs was performed with CDT test and PCR assay was also performed for detection of blaOXA-23-like, blaOXA-24-like, blaOXA-40-like,blaOXA-51-like,blaOXA-58-like and blaOXA-143-like genes
Results:
All isolates of A. baumannii showed high-level of resistance to all antibiotics except for Polymyxin B. The blaOXA-51 like genes was found in all of the isolates and the prevalence of blaOXA-143like, blaOXA-23like,blaOXA-40like and blaOXA-24like were 56%, 45.45%, 33% and 11.8%, respectively.
Conclusion:
The blaOXA-51-like was the predominant mechanism of resistance to imipenem in A. baumannii and therefore, early recognition of carbapenem-resistant A. baumannii isolates is a useful tools to prevent their spreading within the hospital environment.
1. INTRODUCTION
Acinetobacter baumannii as a facultative, non-motile gram-negative coccobacillus, can spread from one patient to others, easily [1]. The organism is the third common pathogenic organism that isolates from hospitalized patients with pneumonia, urinary tract infections, bacteremia, surgical-site infection and Ventilator-Associated Pneumonia (VAP), significantly in intensive-care-unit (ICU) [2-6]. The low rates of the organism recovery from natural environments and its incidence in the community is opposed to the high rate of carriage among hospitalized patients in the hospital setting [7]. Its conditions for growth are simple and can survive in severe conditions even in the absence of humid. The persistence of A. baumannii in the hospital environment represents its main reservoirs, which might contribute to its long-term persistence [8].
A. baumannii strains are generally Multidrug Resistance (MDR), which leads in health worrying for their protection from the antibiotics like cephalosporins, aminoglycosides and fluoroquinolones [9]. Surveillance studies highlighted the gradual decrease of carbapenem-resistant isolates over the last ten years in Europe, North America, and Latin America [10, 11]. The high rate of carbapenem resistance among isolates has been considered recently so that carbapenem resistant A. baumannii has become to a challenge worldwide [12, 13]. On the other hand, the resistance of A. baumannii to carbapenems can be mediated by one of the known resistance mechanisms occur in bacteria, including enzymatic inactivation, active efflux of drugs, and modification of target sites. However, among the reportedly imipenem resistance mechanisms, the most common mechanism might contain the association between carbapenem-hydrolyzing-β-lactamases belonging to the metallo-β-lactamases (Ambler class B) and oxacillinases (Ambler class D). It should be noted that A. baumannii has the ability to hydrolyze oxacillin more efficiently than benzyl penicillin [14, 15]. Class B carbapenemase, such as IMP and VIM enzymes are metallo beta lactamases [16, 17]. Antimicrobial chemotherapy, particularly carbapenems, has contributed to the appearance of carbapenem-hydrolyzing class D β-lactamases (CHDLs). The application of this chemotherapy initiated from the first discovery of a CHDL-encoding gene in 1995 [18-20]. There are four major subgroups of acquired CHDLs in A. baumannii strains, including OXA-23, OXA-40, OXA-58, and OXA-143 β-lactamase groups in addition to the naturally-occurring OXA -51 β-lactamase [21]. The blaOXA-58 like has been identified worldwide, but frequently from France, England, Argentina, Spain, Turkey, Romania, Austria, Greece, Scotland, and Kuwait [22].
A significant contribution of these enzymes in the carbapenem resistance phenotypes has been emphasized, particularly when they are accompanied by ISAba1 and ISAba3 in the naturally occurring plasmid [23]. Therefore, the present study was aimed to determine the drug susceptibility patterns, MBLs detection with imipenem-EDTA Combined Disk Method (CDT) test and molecular epidemiology of blaOXA-23-like, blaOXA-24-like, blaOXA-40-like,blaOXA-51-like,blaOXA-58-like and blaOXA-143-like genes of A. baumannii isolates collected from Hospitals of Tehran, Iran.
2. MATERIALS AND METHODS
2.1. Bacterial Isolation and Identification
A total of 101 Acinetobacter baumannii. was obtained from patients with urinary tract infection from different hospitals of Tehran (Iran) during 2016–2017. The strains were identified as A. baumannii by conventional phenotypic tests (growth in 44˚C, motility, oxidative-fermentative, catalase and oxidase test) and their confirmation was completed by rpoB gene [24].
2.2. Antimicrobial Susceptibility Testing
All isolates were tested by the Kirby-Bauer disk diffusion method according to the Clinical and Laboratory Standards Institute (CLSI) guidelines (CLSI 2018) to assess their susceptibilities to Ciprofloxacin (5 μg), Trimethoprim-sulfamethoxazole (1.25-23.75 µg), Chloramphenicol (10 μg), Gentamicin (10 μg), Imipenem (10 μg), Meropenem (10 μg), Polymyxin B (300 unit), Clindamycin (5 μg), Nalidixic acid (75 μg), Tetracycline (30 μg) (Mast Co., Merseyside, UK). The Escherichia coli ATCC 25922 and Pseudomonas aeruginosa ATCC 27853 strains were used as quality control of antibiotics disc.
2.3. Phenotypic Detection of MBLs with CDT Test
A. baumannii isolates resistant to imipenem and meropenem were taken for confirmation of the MBLs enzymes CDT test. Each of isolates was subcultured on Mueller-Hinton agar plate and two 10 μg imipenem discs with and without of 0.5 M EDTA were placed on the culture plate. Isolates have shown ≥7 mm increase in inhibition zone imipenem-EDTA disc in comparison to the imipenem disc which is considered as MBLs producers [25].
2.4. Molecular Analysis of OXA Genes
DNA extraction of A. baumanni strains was performed by Alkaline lysis method from single colonies from purity plates [26]. Polymerase Chain Reaction (PCR) was done in 20 µl for detection of the carbapenemase-encoding genes (blaOXA-23-like, blaOXA-24-like, blaOXA-40-like,blaOXA-51-like,blaOXA-58-like and blaOXA-143-like). The primers and PCR reaction condition of OXA genes used in amplification reactions were previously described by Kuo et al. [27].
3. RESULTS
A total of 101 clinical isolates of A. baumannii were collected from two hospitals in Tehran (Iran). The complete resistance to Tetracycline (100%), Clindamycin (100%) and Gentamicin (100%) as well as the high resistance to other antibiotics i.e. Meropenem (97%), Nalidixic acid (95%), Ciprofloxacin (93%), Imipenem (94%) and Trimethoprim-sulfamethoxazole (94%) are shown in Table (1). The resistance rates for polymyxin B (86%) were relatively low. In the present study, 97 isolates were able to produce MBL IN CDT test that was significantly associated with imipenem and meropenem resistance resistance (χ2 p<0.05). Furthermore, all of the isolates were extensively drug-resistant phenotype (XDR), which have been defined for the strains showing resistance to some of the most effective anti-bacterial drugs.
| Antimicrobial Agent | Susceptibility (n=101) | ||
|---|---|---|---|
|
Susceptible no (%) |
Intermediate no (%) |
Resistant no (%) |
|
| Trimethoprim-sulfamethoxazole | 5 (4.9) | 1 (0.9) | 94 (93.0) |
| Tetracycline | 0 (0) | 1 (0.9) | 100 (99.0) |
| Clindamycin | 0 (0) | 1 (0.9) | 100 (99.0) |
| Polymyxin B | 6 (5.9) | 9 (8.9) | 86 (85.1) |
| Nalidixic acid | 4 (3.9) | 4 (3.9) | 95 (94.0) |
| Ciprofloxacin | 0 (0) | 6 (5.9) | 93 (92.0) |
| Gentamicin | 1 (0.9) | 0 (0) | 100 (99.0) |
| Imipenem | 0 (0) | 7 (6.9) | 94 (93.0) |
| Meropenem | 1 (0.9) | 3 (2.9) | 97 (96.0) |
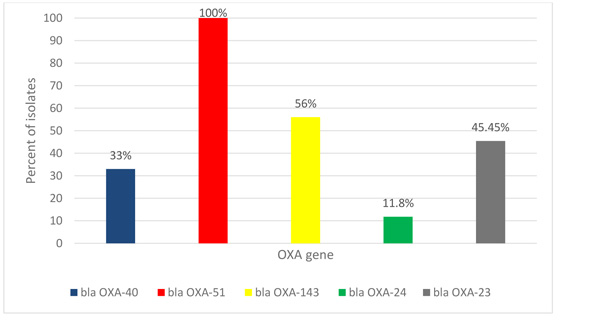
Using PCR assay, all of A. baumannii isolates were positive blaOXA-51 like genes but about (33%) of the isolates were positive for blaOXA-40 like. While 56%, 11.8%, and 45.45% of isolates were positive for blaOXA-143 like,blaOXA-24 like and blaOXA-23 like, respectively but none gave any amplicon for the blaOXA-58 like (Fig. 1).
Co-existence of blaOXA-40 and blaOXA-51 like was observed among 33% of the isolates. The co-relations between carbapen-emase, blaOXA-51 like with blaOXA-143 like and blaOXA-51 like with blaOXA-23 like were statistically significant (χ2 p<0.05).
4. DISCUSSION
Carbapenem resistance has been emerged as a recent common challenge among A. baumannii isolates in Iran and it is difficult to treat of various infections caused by A. baumannii Clinical isolate. This study focused to evaluate the susceptibility rates of A. baumannii isolates toward most of the clinically available antimicrobial agents and molecular epidemiology of blaOXA-23-like, blaOXA-24-like, blaOXA-40-like,blaOXA-51-like,blaOXA-58-like and blaOXA-143-like genes in Tehran. Our results showed that there was a 100% resistance to Tetracycline, Clindamycin, Gentamicin antibiotics, whereas relatively low resistance to Polymyxin B. This antibiotic can be helpful in treating A. baumannii-related infections in hospitals. The global incidence of meropenem resistance in A. baumannii was approximately 6% in 1998 but it has dramatically increased to approximately 29% in 2005 [28].
This rapid increase has also been observed in the Tehran hospitals, where resistance rates to imipenem and meropenem were 94% and 97%, respectively. On the other hand, A. baumannii carbapenem resistance is a multiple drug resistance which is needed in combination therapy. In addition, A. baumannii isolates from other countries were also highly resistant to carbapenems [29-31].
All the clinical A. baumannii isolates obtained in 2006–2007 from Malaysia, exhibited high resistance to all the examined antimicrobial agents except for polymyxin B (different mechanisms are involved in the A. baumannii resistance to imipenem. β-Lactamase enzymes is an important factor to carbapenem-resistance [32].
The acquisition of carbapenem resistance in A. baumannii is mainly because of the production of two types of β-lactamases; Metallo- β-lactamases (MBLs) and carbapenem-hydrolyzing class D β-lactamases (CHDLs).
Amongst carbapenemase enzymes, MBLs show the most important role in creating resistance to carbapenems but CHDLs are the most prevalence carbapenemases detected in A. baumannii. The CHDLs such as blaOXA-23, blaOXA-24, blaOXA-51 appear to be more prevalent and important to carbapenem resistance in this bacterium in some countries, including Bahrain, UAE and Kuwait. In our study, the most detected genes were OXA-51, OXA-143, OXA-23 and OXA-40 which weresimilar to another study from Iran and Bahrain, UAE and Kuwait [33-35]
Despite the weak capability blaOXA-51 and blaOXA-23 to hydrolyze carbapenem antibiotics but, sometimes because of the insertion of ISAba1 element upstream of the blaOXA-51/23-like gene, it may be the decreased level of susceptibility to carbapenems in A. baumannii isolates.
CONCLUSION
In conclusion, blaOXA-51-like has the predominant mechanisms of resistance to imipenem in A. baumannii. However, for a global epidemiologic analysis, further studies on large scale and at different places should be conducted.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
The study was approved by Ethical committe in Baqiyatallah University of Medical Sciences, Tehran, Iran.
HUMAN AND ANIMAL RIGHTS
No animals/humans were used for studies that are the basis of this research.
CONSENT FOR PUBLICATION
Informed consent was obtained from all the participants.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We would like to thank from the “Clinical Research Development Center of Baqiyatallah hospital” for their kindly cooperation. This study was supported in part by a grant from “Clinical Research Development Center of Baqiyatallah hospital”.


