All published articles of this journal are available on ScienceDirect.
Biofilm Formation and Detection of icaD Gene in Staphylococcus aureus Isolated from Clinical Specimens
Abstract
Background:
S. aureus is found to be a major source of community as well as hospital acquired infections. The increase in antimicrobial resistance and emergence of multidrug resistance has become a big threat worldwide. The biofilm formation of S. aureus influenced the survival and persistence in both environment and host.
Aim:
The study was conducted with the aim to evaluate in-vitro biofilm formation and the presence of icaD gene in S. aureus from clinical isolates of S. aureus.
Methods:
A total of 570 wound/pus samples were processed by standard microbiological techniques. Colony morphology, Gram’s staining and biochemical tests were used for the identification of S. aureus. Antimicrobial susceptibility test was performed by Kirby-Bauer disc diffusion technique and methicillin-resistant S. aureus was detected by using cefoxitin antibiotics. The production of biofilm was screened by Congo Red Agar and finally, the presence of icaD gene was determined by PCR.
Results:
Out of 570 samples, a total 19.3% (110/570) samples showed the growth of S. aureus. Among which 59.1% (65/110) were multi-drug resistant. Similarly, 26.4% (29/110) isolates were methicillin-resistant S. aureus. Among MRSA isolates 93.1% (27/29) were MDR with more than 3 classes of antibiotics. Biofilm production was shown by 95.45% (105/110) and 77.3% (85/110) isolates on Congo Red Agar and presence of icaD gene respectively.
Conclusion:
In this study, the significant association was observed in phenotypic production of biofilm and the presence of icaD gene for the genotypic expression of biofilm. There were also increasing rates of MRSA and multidrug resistance S. aureus.
1. BACKGROUND
Staphylococcus aureus is one of the most potent human pathogens, which is commonly associated with nosocomial and community-acquired infections [1]. The severity of the infections ranges from mild to severe life-threatening infections like septicemia, meningitis, pneumonia and endocarditis [2, 3]. Infection imposes a high and increasing burden on health care resources as well as increased morbidity and mortality [4]. S. aureus produces many virulence factors, such as the production of biofilm, hemolysins, leukocidins, proteases, enterotoxins, exfoliative toxins, and immune-modulatory factors [5, 6]. Nosocomial infections that result in the formation of biofilms on the surfaces of biomedical implants are a leading cause of sepsis and are often associated with colonization of the implants by S. epidermidis [7]. Biofilm is an association of microorganisms in which cells stick to each other on a surface encased within the matrix of Extracellular Polymeric Substances (EPS) produced by bacteria themselves [8]. The ability of S. aureus to form biofilm helps the bacterium to resist host immune response and is considered responsible for chronicity of the disease and resistant towards antimicrobial agents [9, 10].
In Staphylococci, the main molecule responsible for intercellular adhesion is the Polysaccharide Intercellular Adhesion (PIA) protein. PIA biosynthesis is accomplished by the products of ica gene locus, which comprises an N-acetylglucosamine transferase (ica A and icaD), a PIA deacetylase (icaB), a putative PIA exporter (icaC) and a regulatory gene (icaR) [11]. Expression of ica gene locus is regulated by a variety of environmental factors and regulatory proteins. Among ica genes, icaA and icaD have been reported to play a significant role in biofilm formation [12]. The icaA gene encodes for N-acetylglucosamine transferase, the enzyme involved in the synthesis of N-acetlyglucosamine oligomers. Further, icaD has been reported to play a critical role in the maximal expression of N-acetylglucosamine transferase, leading to the phenotypic expression of capsular polysaccharide [13].
The antibiotic resistance among S. aureus is a major problem and has posed a serious therapeutic challenge for clinicians [4]. The core resistance phenotype mostly associated with S. aureus persistence to methicillin is due to its expression of penicillin-binding protein, PBP 2a [14]. Besides this, methicillin resistance S. aureus (MRSA) strains have been reported resistant to other antibiotics too such as macrolides, lincosamides, aminoglycosides etc [15].
Early detection and management of antibiotic-resistant and biofilms forming Staphylococci can be one of the essential steps to reduce staphylococcal infections. Thus, there is a need of simple reliable methods to detect biofilms producers. The study was designed with the objective to determine the biofilm producing ability as well as the presence of icaD gene in S. aureus isolated from clinical specimens along with its association with antimicrobial agents.
2. METHODS
2.1. Study Design, Samples and Identification of S. aureus
The study was hospital based cross-sectional study conducted in Annapurna Neurological Institute and Allied Sciences (ANIAS) and Annapurna Research Center, Maitighar, Kathmandu from March to August 2017. The ethical approval was obtained from institutional review board of Nepal health research council (Reg. 27/2017). A total of 570 samples (345 pus aspirates and 225 wound swab) were taken for the study. The samples were collected in clean, sterile and leak proof containers using aseptic technique by experienced medical officers and taken immediately into the microbiology laboratory for further processing. All the samples were inoculated into Blood Agar (BA), Mac Conkey agar (MA) and Mannitol Salt Agar (MSA) and incubated at 37°C for 24 hours. The isolates were identified as S. aureus using standard microbiological techniques including biochemical, slide and tube coagulase tests [16, 17].
2.2. Antibiotic Susceptibility Pattern of S. aureus
Antibiotic susceptibility testing of all isolates of S. aureus was performed by modified Kirby-Bauer disc diffusion method following Clinical and Laboratory Standards Institute (CLSI) guidelines (CLSI 2015). Following antibiotic discs were tested: azithromycin (15 mcg), cefotaxime (30 mcg), cefoxitin (30 mcg), ciprofloxacin (5 mcg), clindamycin (2 mcg), cotri-moxazole (25 mcg), gentamicin (10 mcg) nitrofurantoin (300 mcg) and vancomycin (30mcg). Cefoxitin (30mcg) was used for the detection of MRSA isolates for which size of zone of inhibition is ≥22mm for sensitive and ≤21mm for resistant. Also, isolates which were resistant to 3 or more classes of antibiotics were detected as Multi-Drug Resistant (MDR).
2.3. Phenotypic Characterization of Biofilm Producers
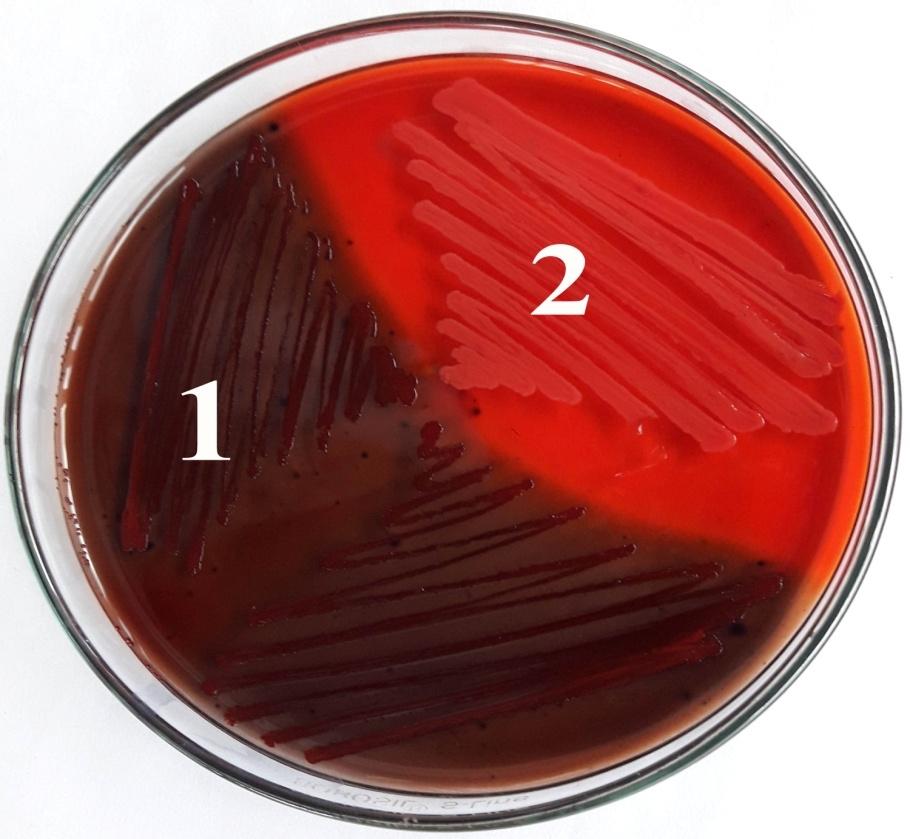
The screening of biofilm producing isolates was done using Congo Red Agar (CRA) method described by Freeman et al. (1989) [18]. The plates after the inoculation of S. aureus were incubated at 37°C for 24-48 hours. The plates were inspected for the color of the colonies. The positive result was indicated by black colonies whereas biofilm non-producing strains develop red colonies.
2.4. DNA Extraction of S. aureus
The genomic DNA template for PCR was extracted from overnight growth of bacterial isolates in Luria-Bertani broth. The DNA was extracted using Cetyltrimethyl Ammonium Bromide (CTAB) method by some modification [19]. About 1 ml of bacterial suspension was centrifuged and supernatant was discarded. The pellet was suspended in Tris-Ethylene Diamine Tetra Acetic acid (EDTA) buffer (TE buffer). Then the lysis of cell wall and proteins was done by adding 10% Sodium dodecyl sulphate (SDS) and 20 mg/ml proteinase K respectively followed by incubation at 37°C for up to 1 hour. After incubation 5M sodium chloride (NaCl) solution followed by CTAB/NaCl (2% CTAB/0.7M NaCl) were added and incubated at 37°C for 10 minutes to remove proteins and polysaccharide complexes. Again equal volume of chloroform: isoamylpropanol (C: I) in the ratio 24: 1 was added to the suspension to separate DNA from proteins and other cellular components. Then, upper aqueous solution containing DNA was transferred to new eppendorf tube after centrifugation. Further pure form of DNA was obtained by treating the solution with isopropanol followed by 70% ethanol. Thus obtained DNA was air dried and re-suspended in 50µl of TE buffer.
2.5. Polymerase Chain Reaction (PCR) of icaD Gene
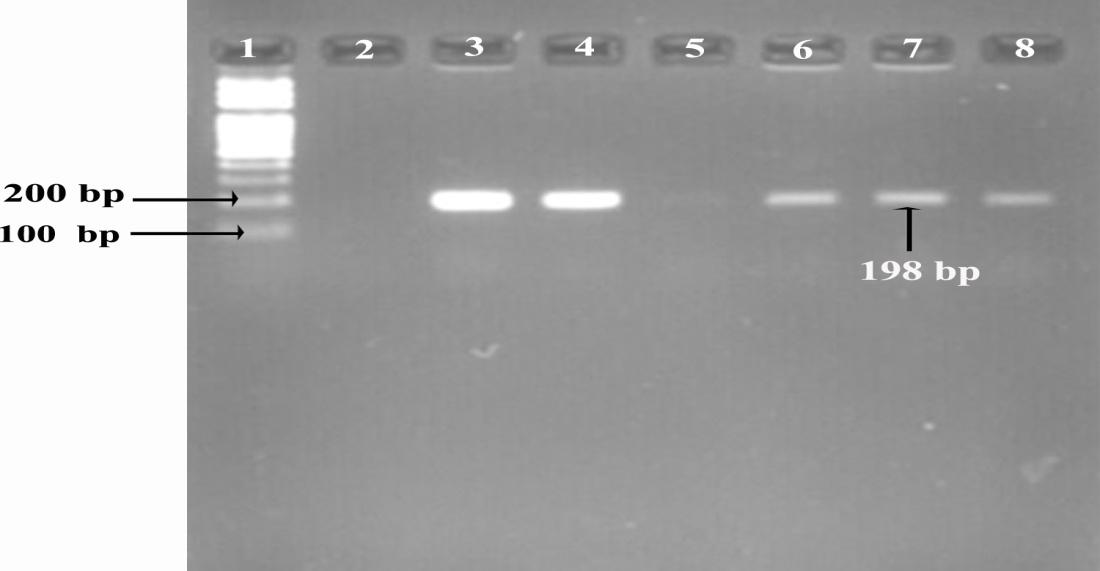
The presence of icaD gene was detected by polymerase chain reaction (PCR) using forward and reverse primers 5ˈ-ATGGTCAAGCCCAGACAGAG-3ˈ and 5ˈ-CGTGTTTTC-AACATTTAATGCAA-3ˈ of 198 base pair respectively. The PCR reaction mixture (25µl) contained 12.5µl Master mix (MgCl2, dNTPs, PCR buffer and Taq polymerase), 2µl of DNA sample, 0.5µl of each primer and 9.5µl sterile deionized distilled water. PCR conditions were initial denaturation (94°C for 5 minutes), followed by 30 cycles of denaturation (94°C for 30 seconds), annealing (55°C for 30 seconds) and extension (72°C for 45 seconds), followed at the end of cycling with final extension (72°C for 7 minutes) [20].
2.6. Gel Electrophoresis for Characterization of icaD Gene
For the separation of PCR products, the amplified PCR product along with positive control, negative control and ladder was run through 2% agarose gel stained with ethidium bromide (EtBr). The gel was visualized under gel documentation system (UVTEC Cambridge, UK) and image was captured.
3. RESULTS
Out of 570 samples, 19.3% (110/570) were positive for S. aureus; in which 70.9% (78/110) and 29.1% (32/110) from pus aspirates and wound swab respectively. Among 110 isolates, 53.6% (59/110) were from outdoor patients while 46.4% (51/110) were from indoor patients. Similarly, 42.7% (47/110) and 57.3% (63/110) were from male and female respectively. In this study, the highest number of the isolates were from the age group 20-30 years and lowest number of isolates were from age group more than 60 years which were 32.7% (36/110) and 10% (11/110) respectively.
Among the antibiotics profile, vancomycin was the most effective drug to which all the isolates of S. aureus were sensitive followed by clindamycin (83.6%), cefoxitin (73.7%) and nitrofurantoin (72.7%). While 59.1% (65/110) isolates were multidrug resistant. Similarly, 26.4% (29/110) isolates were MRSA among which 93.1% (27/29) were MDR and there were significant association (p=0.001) (Table 1).
Out of 110 isolates, 86.3% (95/110) were detected as biofilm producer by the CRA method among which 57.2% (63/110) were strong, 22.7% (28/110) were moderate and 12.7% (14/110) were weak biofilm producers while 4.5% (5/110) isolates were biofilm non producers (Fig. 1). According to genomic analysis, 77.2% (85/110) isolates were positive for the presence of icaD gene (Fig. 2). Among isolates with icaD gene, 97.6% (83/85) isolates showed a positive result for biofilm formation by CRA method and there was significant association (p=0.042). Also 83.1% (55/65) MDR and 75.8% (22/29) MRSA were biofilm producers with the presence of icaD gene (Table 2).
4. DISCUSSION
S. aureus is a common pathogen associated with pyogenic infections and remains a versatile and potent pathogen in human [21]. In this study; 19.3% (110/570) samples showed growth of S. aureus which is almost similar to that of Belbase et al. (2017) which showed 20.37% and 20.9% growth of S. aureus respectively [15]. However, in other studies showed 51.52% and 30.40% of S. aureus respectively, which were comparatively higher rates than that of the present study which may be because of large sample size and long study period [4, 22]. In this study, the rates of infection in indoor and outdoor patients were 46.36% and 53.64% respectively which were comparable to the study of Belbase et al. (2017) as inpatients (43.4%) and outpatients (56.6%) [15]. The infection was higher within the age group of 20-30 years (32.7%) and lower at 10-20 years (14.5%) and ≥60 years (10%). But in the other studies, the rates of S. aureus were higher in the age group ≤10 years which were 24% and 49.1% respectively [4, 23].
| MRSA | MDR | Total | ||
|---|---|---|---|---|
| Positive | Negative | P-value | ||
| Positive | 27 (24.5%) | 2 (1.8%) | 29 (26.3%) | 0.001 |
| Negative | 38 (34.5%) | 43 (39.0%) | 81 (73.6%) | |
| Biofilm | IcaD gene | P-value | ||
|---|---|---|---|---|
| Positive | Negative | Total | ||
| Positive | 83 (75.4%) | 22 (20.0%) | 105 (95.4%) | 0.042 |
| Negative | 2 (1.8%) | 3 (2.7%) | 5 (4.6%) | |
| MRSA | ||||
| Positive | 22 (20.0%) | 7 (6.4%) | 29 (26.4%) | 0.833 |
| Negative | 63 (57.3%) | 18 (16.36%) | 81 (73.6%) | |
| MDR | ||||
| Positive | 54 (49.0%) | 11 (10.0%) | 65 (59.1%) | 0.081 |
| Negative | 31 (28.2%) | 14 (12.7%) | 45 (40.9% | |
Antimicrobial resistance is a global threat and MRSA has emerged as an important human pathogen with a wide range of antibiotic resistance. In recent years, there has been an alarming increase in the prevalence of resistance to methicillin and reduced susceptibility to vancomycin in the S. aureus strains [24]. In this study, the prevalence rate of MRSA was found to be 26.4% which is similar to the other various studies [25-27]. The low rates of prevalence of MRSA were shown in the study of Sudedi and Brahmadathan (2005) and Bhatta et al (2014) which were 15.4% and 19% respectively [28, 29] . On contrary, some of the studies showed high prevalence of MRSA as 56.1% to 69.1% [21, 30-32]. These high rates of MRSA may be due to indiscriminate use of antibiotics and its accessibility in these, lack of awareness and failure to observe simple yet effective infection control precautions like strict patient isolation and frequent hand washing by health care personnel, population and area studied etc. [33].
In the present study, the rate of biofilm formation by S. aureus was 95.45% phenotypical which is comparatively very high rate than in the studies done by Bekir et al. (2012) and Yousefi et al. (2016) which were 46.1%, and 69.2% respectively [24, 34]. The genotypic rate in this study was 77.27% which is similar with the findings of Eftekhar and Dadaei (2010) that showed the genotypic rate of 75% [35]. This indicates that a study of biofilm is required in all the clinical samples in order to know about the status of infection and to get appropriate treatment. In this study, all the isolates which were biofilm producer phenotypically did not show the presence of icaD gene which may be due to the point mutation or may be due to the fact that ica expression is subjected to environmental condition.
Due to the protective nature of biofilm, the bacteria growing in it are intrinsically resistant to many antibiotics [36]. In this study, 59.1% of the total isolates were MDR which is similar to other study 59.2% MDR [15]. In the present study, higher rates of MDR and MRSA were found among biofilm producing strains in comparison to biofilm non-producing strains. In this study, icaD gene were present in 63.5% of the isolates and were MDR which proves that biofilm forming organisms are more resistant than those which do not form biofilm. The antibiotic resistance among strains of bacteria residing in biofilm may increase up to 1000 times [36]. The reason for this may be difficulty in the penetration of biofilm layer by antibiotics, slow growth rate of bacteria and the presence of antibiotic degradation mechanisms. Furthermore, biofilm formation gives a platform for horizontal gene transfer among bacteria, causing easy spread of drug resistance markers and other virulence factors. Also, 27 out of the 29 MRSA isolates were MDR which indicated that almost all the MRSA strains are MDR.
In this study, 24.13% of MRSA isolates showed the absence of icaD gene in them. Clinically MSSA strains predominantly form biofilm dependent in icaADBC operon and PIA synthesis whereas MRSA forms biofilm independent of PIA. There are important roles for surface proteins, the major autolysin and extracellular-DNA (eDNA) during MRSA biofilm formation in absence of PIA. Also, acquisition of MRSA appears to repress polysaccharide type biofilm [37].
S. aureus has the capacity to adhere to indwelling medical devices and form biofilm. In addition, slime producing strains are more virulent and responsible for severe post-surgical infections [34]. The detection of ica locus in clinical S. aureus isolates may improve the clinical decision for treatment and prevention option and could support the development of strategies to interact with the bacterial capacity to colonize and invade in-dwelling medical devices.
CONCLUSION
S. aureus is one of the major infectious agents of wound infection. Only vancomycin was effective antibiotics towards the isolates of S. aureus followed by clindamycin, cefoxitin, and nitrofurantoin while resistant to the rest of the antibiotics used in the study. The study also showed that there was a significant association between MDR and MRSA. Further, the study showed a significant association in between the pheno-typic production of biofilm and the presence of icaD gene for genetic expression of biofilm in S. aureus.
LIST OF ABBREVIATIONS
| CTAB | = Cetyltrimethyl Ammonium Bromide |
| CRA | = Congo Red Agar |
| PCR | = Polymerase Chain Reaction |
AUTHORS’ CONTRIBUTIONS
SK: Data Collection, Funding acquisition, methodology, Manuscript Writing. AKS and SK: Conceptualization, Data analysis, Methodology, Manuscript Writing/Editing. JL, AM, LS and SP: Investigation, Data analysis; RR and SA: Supervision, Investigation. All authors critically revised the manuscript for intellectual content. All authors read and approved the final manuscript.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Ethical approval was obtained from the Institutional Review Board of Nepal Health Research Council (NHRC) (Ref. No. 27/2017).
HUMAN AND ANIMAL RIGHTS
Not applicable.
CONSENT FOR PUBLICATION
Written informed consent was obtained from all patients.
AVAILABILITY OF DATA AND MATERIALS
All data generated or analyzed during this study are included in this article.
FUNDING
None.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We would like to thank Prof. Dr. Bijaya Pant and Prof. Dr. Basant Pant of Annapurna Research Center for providing such a great platform for research. We would also like to thank Mr. Subodh Kant Singh and Ms. Anjana Shrestha for helping us during the laboratory work.


