All published articles of this journal are available on ScienceDirect.
The Molecular Study of Antibiotic Resistance to Quinolones in Salmonella enterica Strains Isolated in Tehran, Iran
Abstract
Introduction:
Salmonella is known as one of the most important causes of gastrointestinal disease in the world. Quinolones and fluoroquinolones are used successfully in the treatment of salmonellosis particularly for infections that have become resistant to several antibiotics. But non-susceptible isolates to quinolones have been reported in several countries. The data are limited about the prevalence of quinolone-resistant isolates in our country. Therefore, this study investigated the plasmid-mediated quinolone resistance genes in Salmonella enterica isolated in Children's Medical Center in Tehran during 2014-2015.
Methods and Materials:
Salmonella isolates were isolated and identified using standard microbiological methods. Antibiotic susceptibility testing and screening of Salmonella strains resistant to quinolones were performed according to the CLSI guidelines. The molecular investigation was done using specific primers for detection of qnr genes including: qnrA, qnrB and qnrS, by polymerase chain reaction.
Results:
Overall, 92 (66.6%) strains were resistant to nalidixic acid. None of the strains showed resistance to ciprofloxacin. Out of the 92 nalidixic acid resistant strains, 52 (56.52%) harbored qnrS genes, 15 strains (16.30%) had both qnrA and qnrS genes. Two (1.1%) isolates were positive for qnrB gene. Twenty four (26.08%) nalidixic acid resistant isolates did not have any qnr qens.
Conclusion:
The results of this study show high prevalence of resistance to nalidixic and qnr genes in Salmonella isolates. Plasmid nature of this type of resistance poses an increased risk of dissemination of quinolone resistance between Salmonella and non-Salmonella isolates circulating in hospitals environments.
1. INTRODUCTION
The infections caused by enteric pathogens comprise second commonest medical problems after respiratory infectious disease [1]. Salmonella, is among the most prevalent and endemic food borne pathogens in Iran [2].
Salmonella is one of the most important genera of Enterobacteriaceae, causing a wide variety of different diseases in society. In Kaufmann-White classification, each serotype is considered a genus based on the antigenic lipopolysaccharide (LPS) structure of cell surface (AgO) and new proteins (AgH), accordingly, 2,600 genus was suggested for this organism [3-6].
Over 90% of serotypes causing intestinal illness in humans are related to Salmonella typhi, S. typhimurium, S. enteritidis and S. infantis [7]. Usually recovery from non typhoidal disease can be achieved without antibiotics. The antibiotics are used in patients suffering enteric fever, septicemia and also in high risk persons, including patients with a weak immune system, malnutrition and malignancy. Ampicillin, chloramphenicol and co-trimoxazole, have been considered for the treatment of salmonellosis; however due to increased resistance of these compounds, usage of fluoroquinolones such as ciprofloxacin and cephalosporin is increased in adults and children [7, 8]. Fluoroquinolones are broad-spectrum antimicrobial drugs which are used for the treatment of clinical salmonellosis infections in elderly or immunocompromised patients as well as patients suffering enteric fever, invasive disease or long-term carriers of Salmonella [8-10].
The increase of antibiotic resistance in developing countries’ patients has become an important concern [11]. Acquisition of antibiotic resistance occurs through several mechanisms [12] including chromosomal mutation, resistance transfer to other bacteria through the transposable elements such as plasmids and transposons, decrease in the cell wall permeability of antibiotics, antibiotics enzymatic inactivation, and efflux pump mechanisms for removing antibiotics [13].
Rapid detection and molecular epidemiology investigation of resistance genes in bacteria are extremely important [14-17].
We aimed to investigate the plasmid-mediated quinolone resistance genes in Salmonella enterica isolated from Children's Medical Center in Tehran during 2014-2015.
2. MATERIALS AND METHODS
The study was conducted on clinical specimens recovered from patients who were suspected to Salmonella infection admitted to Children's Medical Center in Tehran during the years 2014-2015. Bacterial isolation and identification were done according to the standard microbiological and biochemical methods.
After performing biochemical tests for Salmonella confirmation, serotyping was performed to determine the O and H antigens with specific antiserum.
Antibiotic susceptibility testing for nalidixic acid (NA 30μg) and ciprofloxacin (CP 5μg) was done by modified Kirby-Bauer method. Escherichia coli ATCC 25922 were used as a control in the experiment.
| Amplicon Size (bp) | Length |
Tm (C°) |
Sequences | Primers | Gene |
|---|---|---|---|---|---|
| 516 | 20 | 53.9 | F: 5′-ATT TCT CAC GCC AGG ATT TG-3′ | qnrA | qnrA |
| 20 | 53.9 | R: 5′-GAT CGG CAA AGG TTA GGT CA-3′ | |||
| 526 | 22 | 55.6 | F: 5′-GTT GGC GAA AAA ATT GAC AGA A-3′ | qnrB | qnrB |
| 20 | 53.6 | R: 5′-ACT CCG AAT TGG TCA GAT CG-3′ | |||
| 417 | 20 | 53.7 | F: 5′-ACG ACA TTC GTC AAC TGC AA-3′ | qnrS | qnrS |
| 20 | 54 | R: 5′-TTA ATT GGC ACC CTG TAG GC-3′ |
DNA was extracted by boiling method. The primers used in this study are listed in Table (1) [18]. PCR was performed for the amplification of qnrA, qnrB, qnrS genes in thermocycler machine under the conditions shown in (Table 2).
| PCR Steps | Temperature C ° | Time (min) | Number of Cycles |
|---|---|---|---|
| Initial Denaturation | 94 | 10:00 | 1 |
| Denaturation | 94 | 1:00 | 30 |
| Annealing | 52 | 1:00 | 30 |
| Elongation | 72 | 1:00 | 30 |
| Final Extension | 72 | 7:00 | 1 |
3. RESULTS
Antibiotic susceptibility testing results showed that 92(66.6%) strains of Salmonella enterica were resistant to nalidixic acid. None of the investigated strains showed resistance to ciprofloxacin.
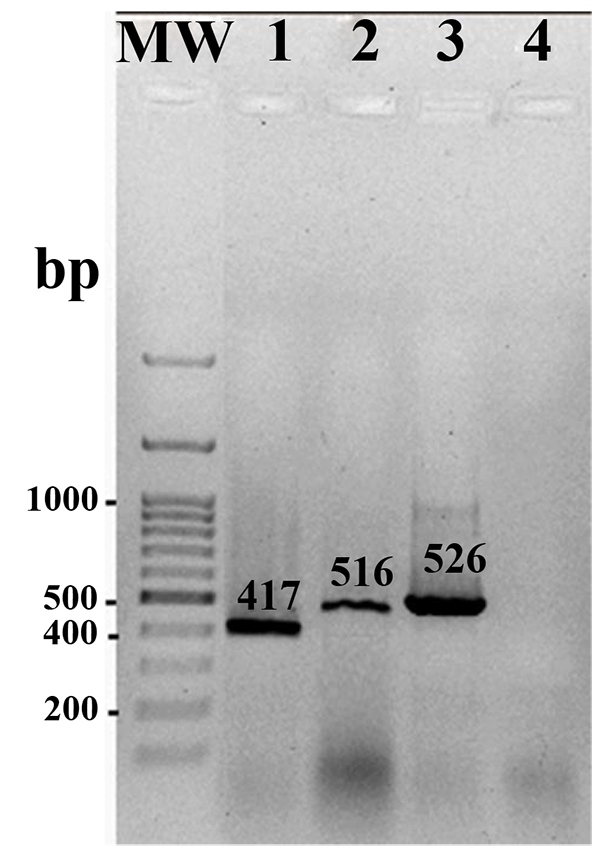
PCR amplified the expected amplicons with sizes of 417bp, 516bp, and 526bp for qnrS, qnrA and qnrB genes, respectively. Fig. (1) shows amplified target genes from representatives Salmonella isolates harboring different qnr genes. Overall, out of 92 nalidixic acid resistant Salmonella strains, 52 (56.5%) had qnrS gene. The genes qnrA and qnr B were observed in 28 and 2 strains respectively. Among nalidixic acid-resistant strains, 15 (16.30%) strains harbored qnrS gene together. Only one strain contained qnr B gene.
4. DISCUSSION
In the past, Ampicillin, chloramphenicol and co-trimoxazole, have been used for the treatment of salmonellosis, but today, fluoroquinolones such as ciprofloxacin and cephalosporin are used for adults and children respectively [8]. The strains resistant to nalidixic acid have a reduced sensitivity to fluoroquinolones [18, 19].
In recent years, several cases of treatment failure with fluoroquinolones have been reported in patients infected by Salmonella strains resistant to nalidixic acid and reduced sensitivity to ciprofloxacin. Unfortunately, their number is increasing [20, 21].
In Nigeria, Akinyemi et al. showed that 42%, 43%, 42%, 47%, 51%, 57%, 56% and 59% of Salmonella strains recovered in the years 1997-2004 were resistant to nalidixic acid respectively [22].
Similar results were reported by Ghaderi et al. in which 67.2% Salmonella strains were resistant to nalidixic acid [4].
In a study conducted in Spain by Prats et al. showed 10% increase in resistance of nalidixic acid from 0.1% to 11% over seven years [23].
Compared to previous studies, our results show the increasing rates of Salmonella strains resistant to nalidixic acid in Tehran. The rate of resistance to nalidixic acid can be important, because fluoroquinolones have been used as a chosen antibiotic in the treatment of infections caused by Salmonella. Antibiotic therapy using fluoroquinolones, may not be successful in people who were infected by nalidixic acid resistant Salmonella strains [20, 24].
Ferrari et al. showed plasmid- depended quinolone resistance, by gene qnr and gene aac (6 ') Ib-cr in 126 Salmonella strains isolated from poultry and humans food sources during the years 1999 to 2007 in the state of Paraná (Brazil). Overall, 88% and 23% of strains showed resistant to Nalidixic acid and reduced susceptibility to ciprofloxacin respectively. QnrA1 gene was shown in a S. enteritidis strain [25].
Ghaima et al. in 2016, accomplished a study to determine the presence of qnr genes in 40 strains of Salmonella isolated from stool samples of patients in Iraq. They found 30 isolates (75%) had multidrug-resistant (MDR). Based on the results of PCR, 15 (37.5%), 22 (55%), and 11 (27.5%) isolates had qnrB, qnrS and qnrA respectively. 10 cases (25%) contain qnrA, qnrB and qnrS [26].
Lunn et al. determined the mechanism of plasmid- depended quinolone resistance in 41 collected Salmonella strains in 2010. None of the isolates were resistant to ciprofloxacin, while 41.5% of the strains showed resistance to nalidixic acid by mutation in gyrA. Although qepA, qnrB, qnrS1, were found in 4 isolates, but the expression of this gene was not accompanied by reduced susceptibility to quinolones [27].
CONCLUSION
Compared to previous studies, the rate of resistance to nalidixic acid is increasing among clinical strains of Salmonella enterica that poses a warning alarm for antibiotic therapy of infections caused by this organism in Iran. The molecular method used in this study was able to show the genetic nature of resistance to quinolones among Salmonella strains under the study. Plasmid nature of this type of resistance poses an increased risk of dissemination of quinolone resistance between Salmonella and non- Salmonella isolates circulating in hospitals environments in our country.
ETHICS APPROVAL AND CONSENT TO PARTICIPATE
Not applicable.
HUMAN AND ANIMAL RIGHTS
No Animals/Humans were used for studies that are base of this research.
CONSENT FOR PUBLICATION
Not applicable.
CONFLICT OF INTEREST
The authors declare no conflict of interest, financial or otherwise.
ACKNOWLEDGEMENTS
We would like to thank from the "Clinical Research Developement Center of Baqiyatallah hospital" for their kindly cooperation. This study was supported in part by a grant from "Clinical Research Developement Center of Baqiyatallah hospital".


